Ursula Schmidt-Erfurth
on behalf of the PINNACLE consortium
Projective Skip-Connections for Segmentation Along a Subset of Dimensions in Retinal OCT
Aug 02, 2021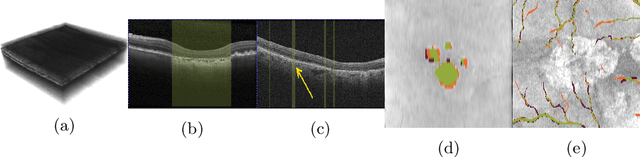
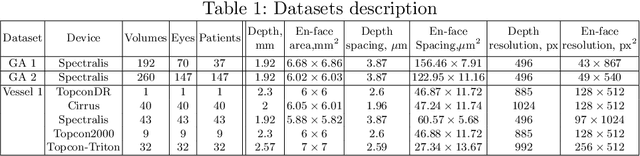
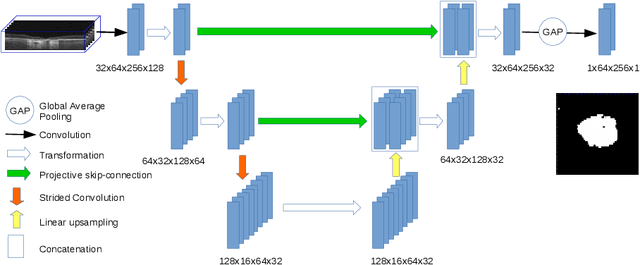
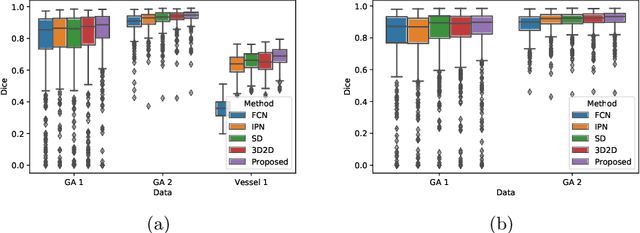
Abstract:In medical imaging, there are clinically relevant segmentation tasks where the output mask is a projection to a subset of input image dimensions. In this work, we propose a novel convolutional neural network architecture that can effectively learn to produce a lower-dimensional segmentation mask than the input image. The network restores encoded representation only in a subset of input spatial dimensions and keeps the representation unchanged in the others. The newly proposed projective skip-connections allow linking the encoder and decoder in a UNet-like structure. We evaluated the proposed method on two clinically relevant tasks in retinal Optical Coherence Tomography (OCT): geographic atrophy and retinal blood vessel segmentation. The proposed method outperformed the current state-of-the-art approaches on all the OCT datasets used, consisting of 3D volumes and corresponding 2D en-face masks. The proposed architecture fills the methodological gap between image classification and ND image segmentation.
U-Net with spatial pyramid pooling for drusen segmentation in optical coherence tomography
Dec 11, 2019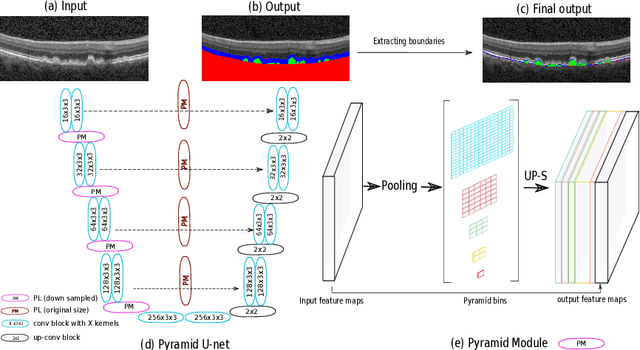
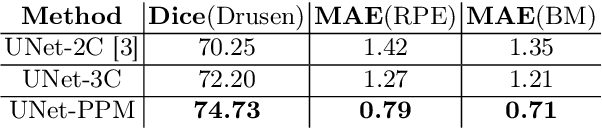

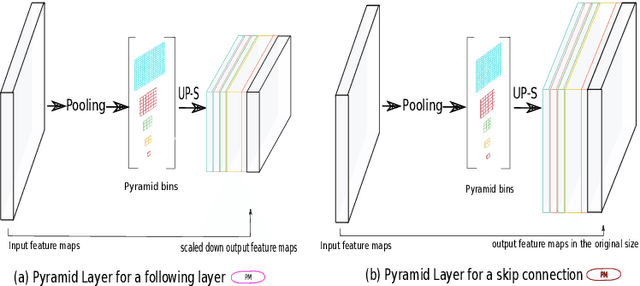
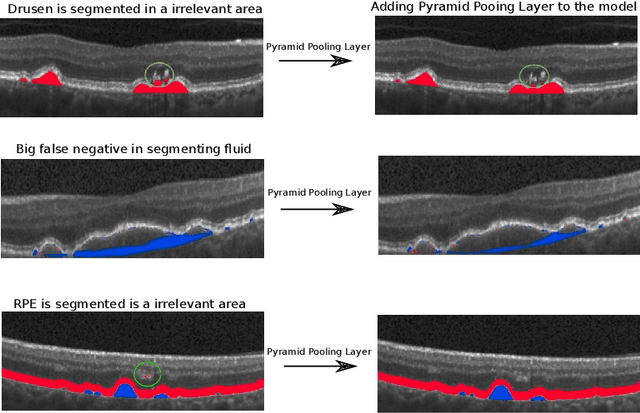
Abstract:The presence of drusen is the main hallmark of early/intermediate age-related macular degeneration (AMD). Therefore, automated drusen segmentation is an important step in image-guided management of AMD. There are two common approaches to drusen segmentation. In the first, the drusen are segmented directly as a binary classification task. In the second approach, the surrounding retinal layers (outer boundary retinal pigment epithelium (OBRPE) and Bruch's membrane (BM)) are segmented and the remaining space between these two layers is extracted as drusen. In this work, we extend the standard U-Net architecture with spatial pyramid pooling components to introduce global feature context. We apply the model to the task of segmenting drusen together with BM and OBRPE. The proposed network was trained and evaluated on a longitudinal OCT dataset of 425 scans from 38 patients with early/intermediate AMD. This preliminary study showed that the proposed network consistently outperformed the standard U-net model.
Modeling Disease Progression In Retinal OCTs With Longitudinal Self-Supervised Learning
Oct 24, 2019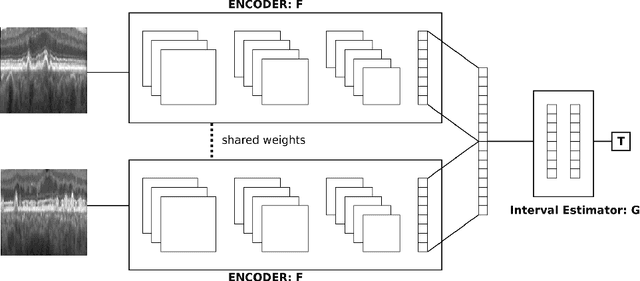
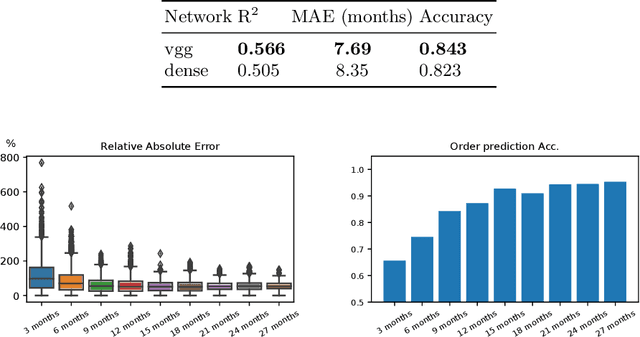
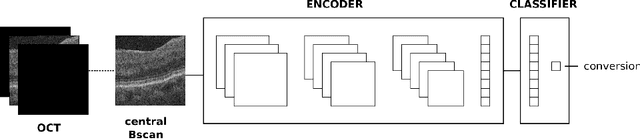
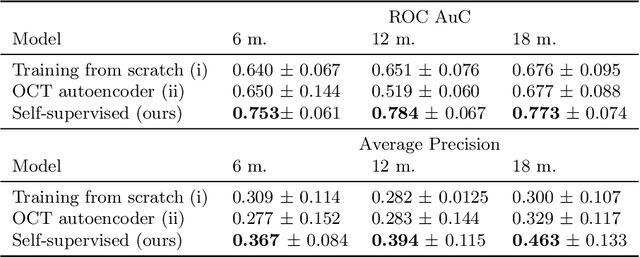
Abstract:Longitudinal imaging is capable of capturing the static ana\-to\-mi\-cal structures and the dynamic changes of the morphology resulting from aging or disease progression. Self-supervised learning allows to learn new representation from available large unlabelled data without any expert knowledge. We propose a deep learning self-supervised approach to model disease progression from longitudinal retinal optical coherence tomography (OCT). Our self-supervised model takes benefit from a generic time-related task, by learning to estimate the time interval between pairs of scans acquired from the same patient. This task is (i) easy to implement, (ii) allows to use irregularly sampled data, (iii) is tolerant to poor registration, and (iv) does not rely on additional annotations. This novel method learns a representation that focuses on progression specific information only, which can be transferred to other types of longitudinal problems. We transfer the learnt representation to a clinically highly relevant task of predicting the onset of an advanced stage of age-related macular degeneration within a given time interval based on a single OCT scan. The boost in prediction accuracy, in comparison to a network learned from scratch or transferred from traditional tasks, demonstrates that our pretrained self-supervised representation learns a clinically meaningful information.
An amplified-target loss approach for photoreceptor layer segmentation in pathological OCT scans
Aug 02, 2019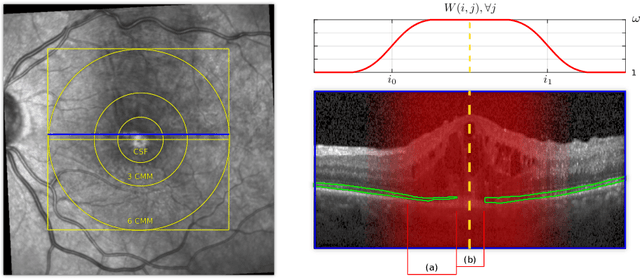
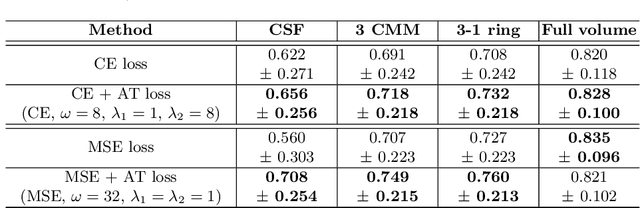
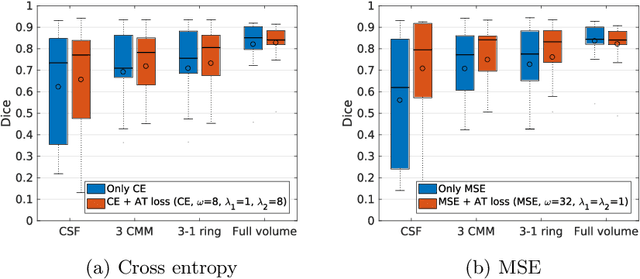
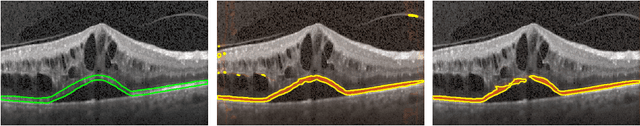
Abstract:Segmenting anatomical structures such as the photoreceptor layer in retinal optical coherence tomography (OCT) scans is challenging in pathological scenarios. Supervised deep learning models trained with standard loss functions are usually able to characterize only the most common disease appeareance from a training set, resulting in suboptimal performance and poor generalization when dealing with unseen lesions. In this paper we propose to overcome this limitation by means of an augmented target loss function framework. We introduce a novel amplified-target loss that explicitly penalizes errors within the central area of the input images, based on the observation that most of the challenging disease appeareance is usually located in this area. We experimentally validated our approach using a data set with OCT scans of patients with macular diseases. We observe increased performance compared to the models that use only the standard losses. Our proposed loss function strongly supports the segmentation model to better distinguish photoreceptors in highly pathological scenarios.
Multiclass segmentation as multitask learning for drusen segmentation in retinal optical coherence tomography
Jul 24, 2019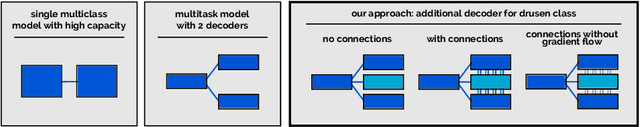
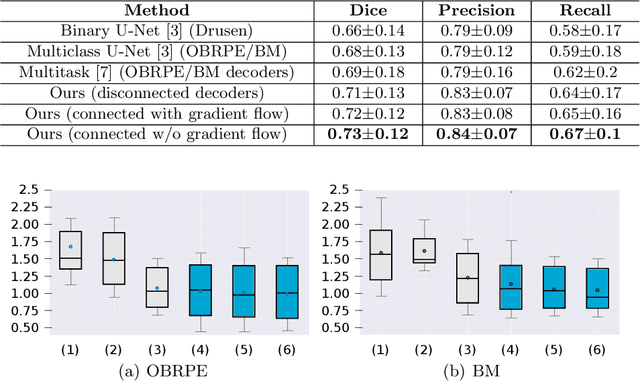
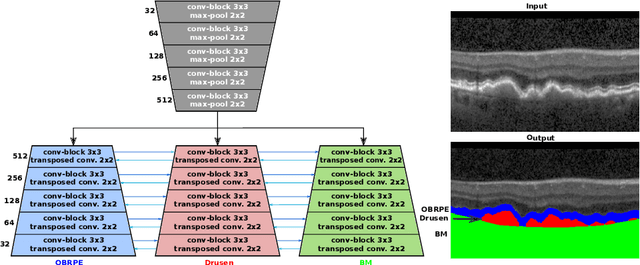
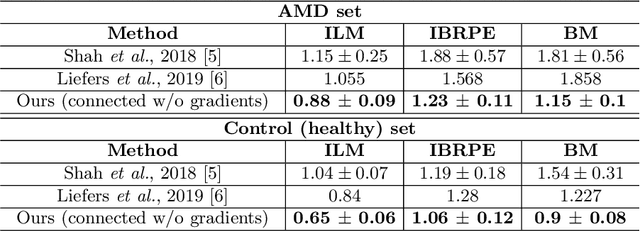
Abstract:Automated drusen segmentation in retinal optical coherence tomography (OCT) scans is relevant for understanding age-related macular degeneration (AMD) risk and progression. This task is usually performed by segmenting the top/bottom anatomical interfaces that define drusen, the outer boundary of the retinal pigment epithelium (OBRPE) and the Bruch's membrane (BM), respectively. In this paper we propose a novel multi-decoder architecture that tackles drusen segmentation as a multitask problem. Instead of training a multiclass model for OBRPE/BM segmentation, we use one decoder per target class and an extra one aiming for the area between the layers. We also introduce connections between each class-specific branch and the additional decoder to increase the regularization effect of this surrogate task. We validated our approach on private/public data sets with 166 early/intermediate AMD Spectralis, and 200 AMD and control Bioptigen OCT volumes, respectively. Our method consistently outperformed several baselines in both layer and drusen segmentation evaluations.
Exploiting Epistemic Uncertainty of Anatomy Segmentation for Anomaly Detection in Retinal OCT
May 29, 2019
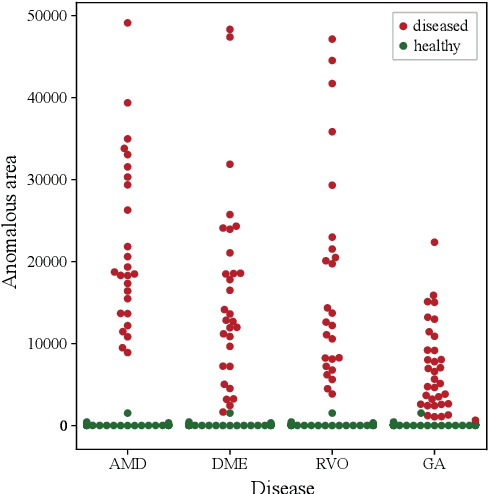
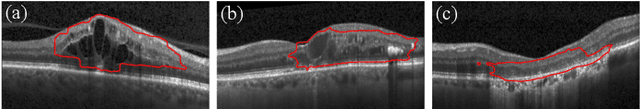
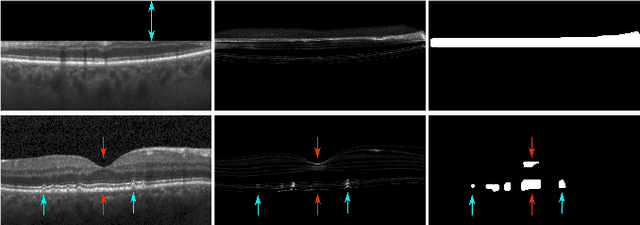
Abstract:Diagnosis and treatment guidance are aided by detecting relevant biomarkers in medical images. Although supervised deep learning can perform accurate segmentation of pathological areas, it is limited by requiring a-priori definitions of these regions, large-scale annotations, and a representative patient cohort in the training set. In contrast, anomaly detection is not limited to specific definitions of pathologies and allows for training on healthy samples without annotation. Anomalous regions can then serve as candidates for biomarker discovery. Knowledge about normal anatomical structure brings implicit information for detecting anomalies. We propose to take advantage of this property using bayesian deep learning, based on the assumption that epistemic uncertainties will correlate with anatomical deviations from a normal training set. A Bayesian U-Net is trained on a well-defined healthy environment using weak labels of healthy anatomy produced by existing methods. At test time, we capture epistemic uncertainty estimates of our model using Monte Carlo dropout. A novel post-processing technique is then applied to exploit these estimates and transfer their layered appearance to smooth blob-shaped segmentations of the anomalies. We experimentally validated this approach in retinal optical coherence tomography (OCT) images, using weak labels of retinal layers. Our method achieved a Dice index of 0.789 in an independent anomaly test set of age-related macular degeneration (AMD) cases. The resulting segmentations allowed very high accuracy for separating healthy and diseased cases with late wet AMD, dry geographic atrophy (GA), diabetic macular edema (DME) and retinal vein occlusion (RVO). Finally, we qualitatively observed that our approach can also detect other deviations in normal scans such as cut edge artifacts.
Using CycleGANs for effectively reducing image variability across OCT devices and improving retinal fluid segmentation
Jan 25, 2019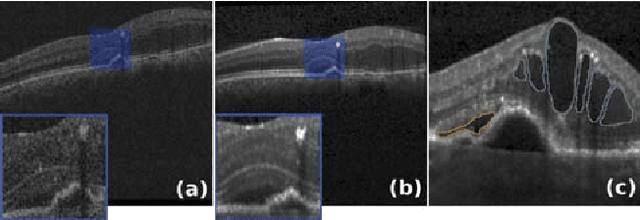
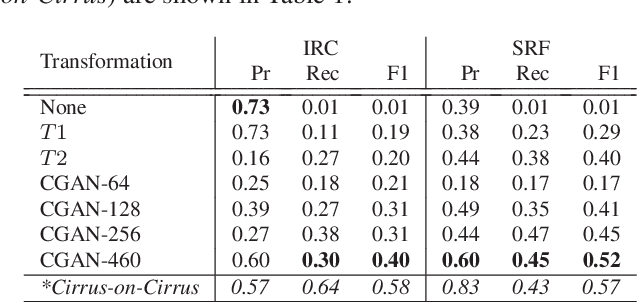
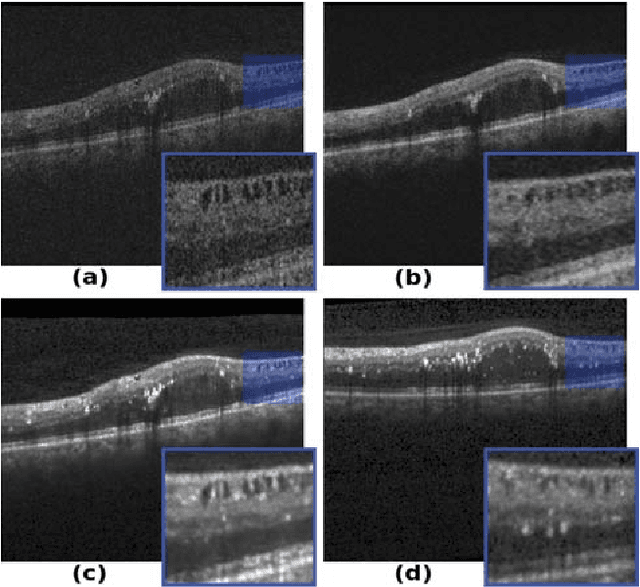
Abstract:Optical coherence tomography (OCT) has become the most important imaging modality in ophthalmology. A substantial amount of research has recently been devoted to the development of machine learning (ML) models for the identification and quantification of pathological features in OCT images. Among the several sources of variability the ML models have to deal with, a major factor is the acquisition device, which can limit the ML model's generalizability. In this paper, we propose to reduce the image variability across different OCT devices (Spectralis and Cirrus) by using CycleGAN, an unsupervised unpaired image transformation algorithm. The usefulness of this approach is evaluated in the setting of retinal fluid segmentation, namely intraretinal cystoid fluid (IRC) and subretinal fluid (SRF). First, we train a segmentation model on images acquired with a source OCT device. Then we evaluate the model on (1) source, (2) target and (3) transformed versions of the target OCT images. The presented transformation strategy shows an F1 score of 0.4 (0.51) for IRC (SRF) segmentations. Compared with traditional transformation approaches, this means an F1 score gain of 0.2 (0.12).
U2-Net: A Bayesian U-Net model with epistemic uncertainty feedback for photoreceptor layer segmentation in pathological OCT scans
Jan 23, 2019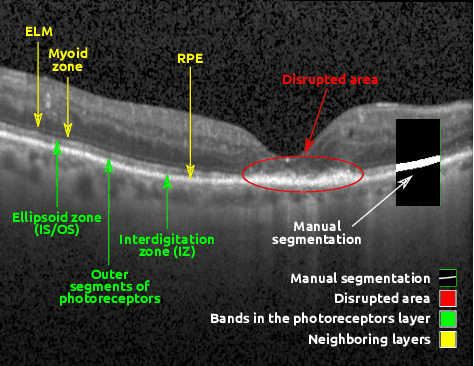
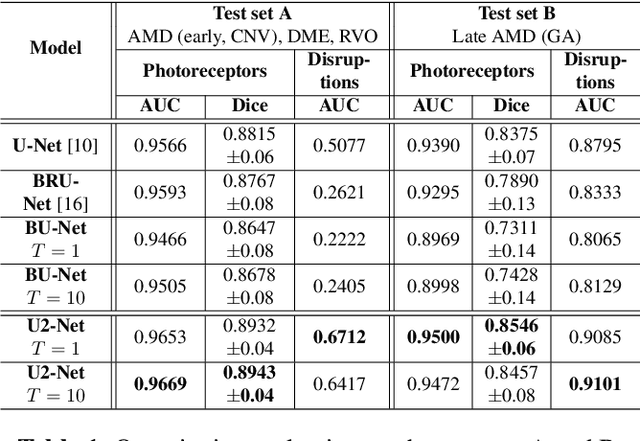
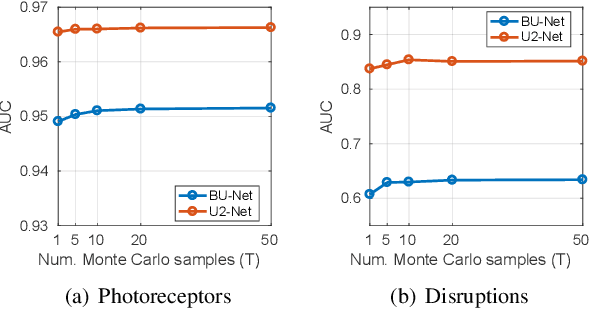
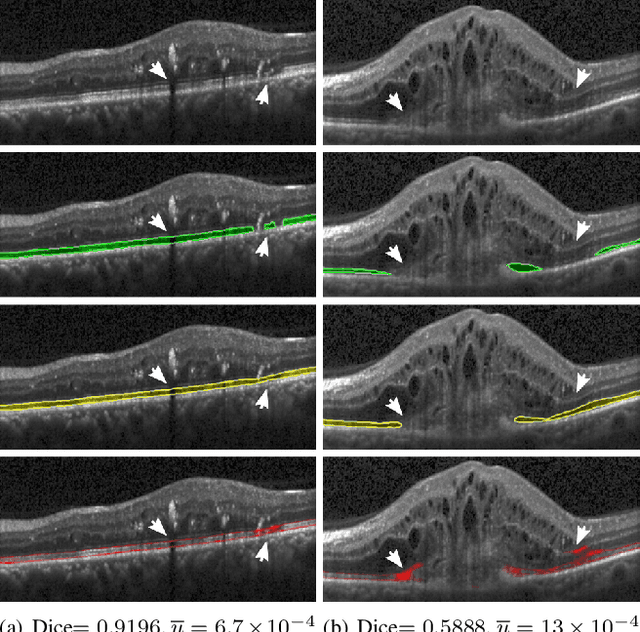
Abstract:In this paper, we introduce a Bayesian deep learning based model for segmenting the photoreceptor layer in pathological OCT scans. Our architecture provides accurate segmentations of the photoreceptor layer and produces pixel-wise epistemic uncertainty maps that highlight potential areas of pathologies or segmentation errors. We empirically evaluated this approach in two sets of pathological OCT scans of patients with age-related macular degeneration, retinal vein oclussion and diabetic macular edema, improving the performance of the baseline U-Net both in terms of the Dice index and the area under the precision/recall curve. We also observed that the uncertainty estimates were inversely correlated with the model performance, underlying its utility for highlighting areas where manual inspection/correction might be needed.
* Accepted for publication at IEEE International Symposium on Biomedical Imaging (ISBI) 2019
On orthogonal projections for dimension reduction and applications in variational loss functions for learning problems
Jan 22, 2019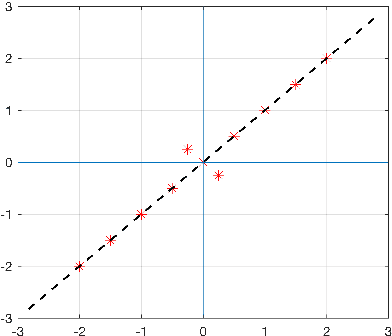
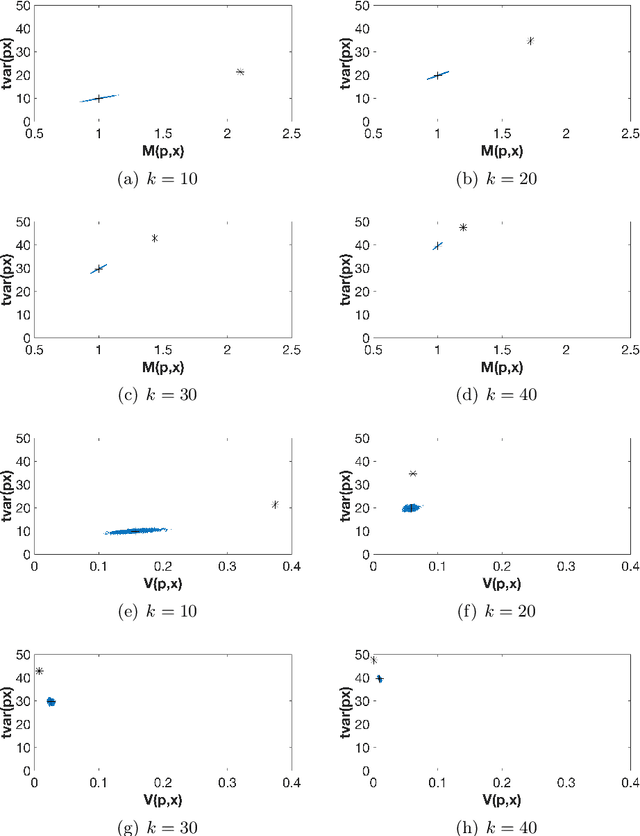
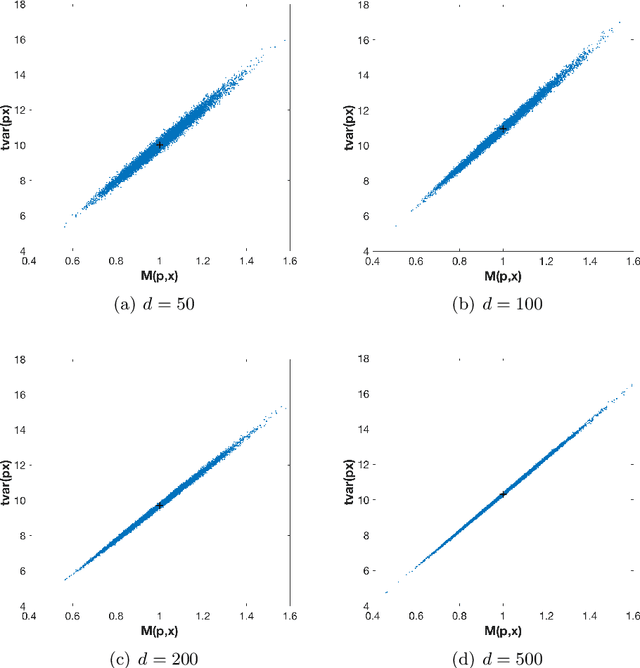
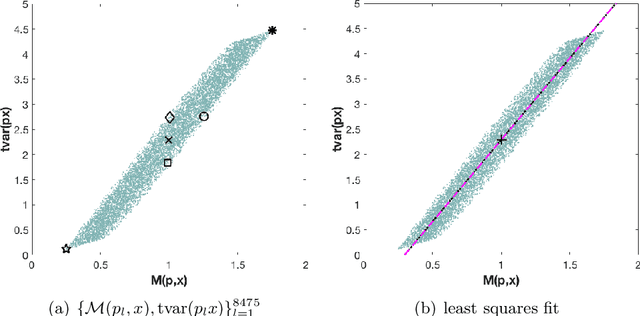
Abstract:The use of orthogonal projections on high-dimensional input and target data in learning frameworks is studied. First, we investigate the relations between two standard objectives in dimension reduction, maximizing variance and preservation of pairwise relative distances. The derivation of their asymptotic correlation and numerical experiments tell that a projection usually cannot satisfy both objectives. In a standard classification problem we determine projections on the input data that balance them and compare subsequent results. Next, we extend our application of orthogonal projections to deep learning frameworks. We introduce new variational loss functions that enable integration of additional information via transformations and projections of the target data. In two supervised learning problems, clinical image segmentation and music information classification, the application of the proposed loss functions increase the accuracy.
Unsupervised Identification of Disease Marker Candidates in Retinal OCT Imaging Data
Oct 31, 2018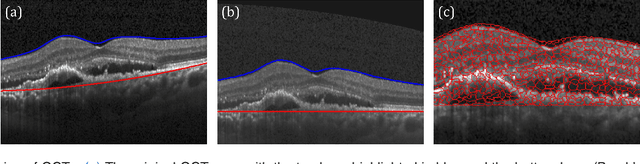
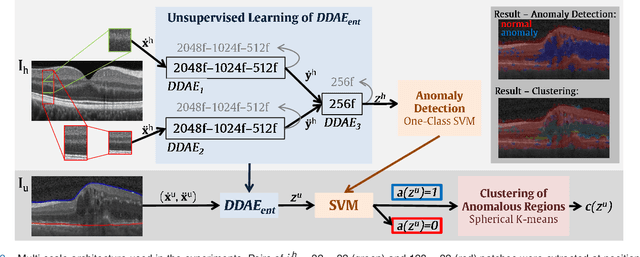
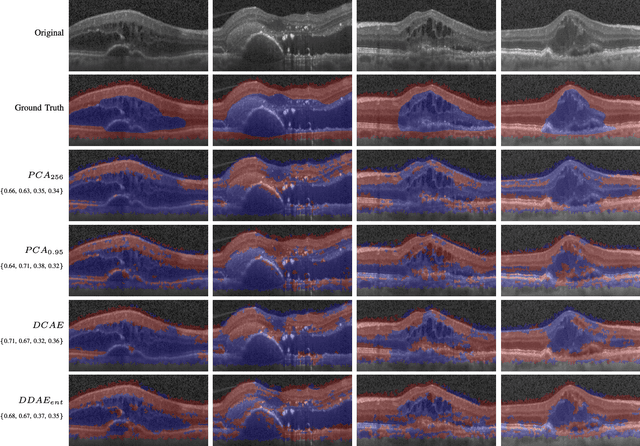
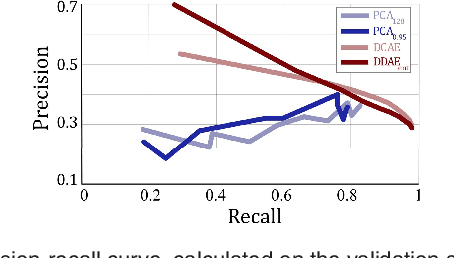
Abstract:The identification and quantification of markers in medical images is critical for diagnosis, prognosis, and disease management. Supervised machine learning enables the detection and exploitation of findings that are known a priori after annotation of training examples by experts. However, supervision does not scale well, due to the amount of necessary training examples, and the limitation of the marker vocabulary to known entities. In this proof-of-concept study, we propose unsupervised identification of anomalies as candidates for markers in retinal Optical Coherence Tomography (OCT) imaging data without a constraint to a priori definitions. We identify and categorize marker candidates occurring frequently in the data, and demonstrate that these markers show predictive value in the task of detecting disease. A careful qualitative analysis of the identified data driven markers reveals how their quantifiable occurrence aligns with our current understanding of disease course, in early- and late age-related macular degeneration (AMD) patients. A multi-scale deep denoising autoencoder is trained on healthy images, and a one-class support vector machine identifies anomalies in new data. Clustering in the anomalies identifies stable categories. Using these markers to classify healthy-, early AMD- and late AMD cases yields an accuracy of 81.40%. In a second binary classification experiment on a publicly available data set (healthy vs. intermediate AMD) the model achieves an area under the ROC curve of 0.944.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge