Umar Marikkar
Domain Adaptation Without the Compute Burden for Efficient Whole Slide Image Analysis
Mar 16, 2026Abstract:Computational methods on analyzing Whole Slide Images (WSIs) enable early diagnosis and treatments by supporting pathologists in detection and classification of tumors. However, the extremely high resolution of WSIs makes end-to-end training impractical compared to typical image analysis tasks. To address this, most approaches use pre-trained feature extractors to obtain fixed representations of whole slides, which are then combined with Multiple Instance Learning (MIL) for downstream tasks. These feature extractors are typically pre-trained on natural image datasets such as ImageNet, which fail to capture domain-specific characteristics. Although domain-specific pre-training on histopathology data yields more relevant feature representations, it remains computationally expensive and fail to capture task-specific characteristics within the domain. To address the computational cost and lack of task-specificity in domain-specific pre-training, we propose EfficientWSI (eWSI), a careful integration of Parameter-Efficient-Fine-Tuning (PEFT) and Multiple Instance Learning (MIL) that enables end-to-end training on WSI tasks. We evaluate eWSI on seven WSI-level tasks over Camelyon16, TCGA and BRACS datasets. Our results show that eWSI when applied with ImageNet feature extractors yields strong classification performance, matching or outperforming MILs with in-domain feature extractors, alleviating the need for extensive in-domain pre-training. Furthermore, when eWSI is applied with in-domain feature extractors, it further improves classification performance in most cases, demonstrating its ability to capture task-specific information where beneficial. Our findings suggest that eWSI provides a task-targeted, computationally efficient path for WSI tasks, offering a promising direction for task-specific learning in computational pathology.
DC-ViT: Modulating Spatial and Channel Interactions for Multi-Channel Images
Mar 15, 2026Abstract:Training and evaluation in multi-channel imaging (MCI) remains challenging due to heterogeneous channel configurations arising from varying staining protocols, sensor types, and acquisition settings. This heterogeneity limits the applicability of fixed-channel encoders commonly used in general computer vision. Recent Multi-Channel Vision Transformers (MC-ViTs) address this by enabling flexible channel inputs, typically by jointly encoding patch tokens from all channels within a unified attention space. However, unrestricted token interactions across channels can lead to feature dilution, reducing the ability to preserve channel-specific semantics that are critical in MCI data. To address this, we propose Decoupled Vision Transformer (DC-ViT), which explicitly regulates information sharing using Decoupled Self-Attention (DSA), which decomposes token updates into two complementary pathways: spatial updates that model intra-channel structure, and channel-wise updates that adaptively integrate cross-channel information. This decoupling mitigates informational collapse while allowing selective inter-channel interaction. To further exploit these enhanced channel-specific representations, we introduce Decoupled Aggregation (DAG), which allows the model to learn task-specific channel importances. Extensive experiments across three MCI benchmarks demonstrate consistent improvements over existing MC-ViT approaches.
Channel-Aware Probing for Multi-Channel Imaging
Feb 13, 2026Abstract:Training and evaluating vision encoders on Multi-Channel Imaging (MCI) data remains challenging as channel configurations vary across datasets, preventing fixed-channel training and limiting reuse of pre-trained encoders on new channel settings. Prior work trains MCI encoders but typically evaluates them via full fine-tuning, leaving probing with frozen pre-trained encoders comparatively underexplored. Existing studies that perform probing largely focus on improving representations, rather than how to best leverage fixed representations for downstream tasks. Although the latter problem has been studied in other domains, directly transferring those strategies to MCI yields weak results, even worse than training from scratch. We therefore propose Channel-Aware Probing (CAP), which exploits the intrinsic inter-channel diversity in MCI datasets by controlling feature flow at both the encoder and probe levels. CAP uses Independent Feature Encoding (IFE) to encode each channel separately, and Decoupled Pooling (DCP) to pool within channels before aggregating across channels. Across three MCI benchmarks, CAP consistently improves probing performance over the default probing protocol, matches fine-tuning from scratch, and largely reduces the gap to full fine-tuning from the same MCI pre-trained checkpoints. Code can be found in https://github.com/umarikkar/CAP.
C3R: Channel Conditioned Cell Representations for unified evaluation in microscopy imaging
May 24, 2025Abstract:Immunohistochemical (IHC) images reveal detailed information about structures and functions at the subcellular level. However, unlike natural images, IHC datasets pose challenges for deep learning models due to their inconsistencies in channel count and configuration, stemming from varying staining protocols across laboratories and studies. Existing approaches build channel-adaptive models, which unfortunately fail to support out-of-distribution (OOD) evaluation across IHC datasets and cannot be applied in a true zero-shot setting with mismatched channel counts. To address this, we introduce a structured view of cellular image channels by grouping them into either context or concept, where we treat the context channels as a reference to the concept channels in the image. We leverage this context-concept principle to develop Channel Conditioned Cell Representations (C3R), a framework designed for unified evaluation on in-distribution (ID) and OOD datasets. C3R is a two-fold framework comprising a channel-adaptive encoder architecture and a masked knowledge distillation training strategy, both built around the context-concept principle. We find that C3R outperforms existing benchmarks on both ID and OOD tasks, while a trivial implementation of our core idea also outperforms the channel-adaptive methods reported on the CHAMMI benchmark. Our method opens a new pathway for cross-dataset generalization between IHC datasets, without requiring dataset-specific adaptation or retraining.
Review of multimodal machine learning approaches in healthcare
Feb 12, 2024Abstract:Machine learning methods in healthcare have traditionally focused on using data from a single modality, limiting their ability to effectively replicate the clinical practice of integrating multiple sources of information for improved decision making. Clinicians typically rely on a variety of data sources including patients' demographic information, laboratory data, vital signs and various imaging data modalities to make informed decisions and contextualise their findings. Recent advances in machine learning have facilitated the more efficient incorporation of multimodal data, resulting in applications that better represent the clinician's approach. Here, we provide a review of multimodal machine learning approaches in healthcare, offering a comprehensive overview of recent literature. We discuss the various data modalities used in clinical diagnosis, with a particular emphasis on imaging data. We evaluate fusion techniques, explore existing multimodal datasets and examine common training strategies.
LT-ViT: A Vision Transformer for multi-label Chest X-ray classification
Nov 13, 2023Abstract:Vision Transformers (ViTs) are widely adopted in medical imaging tasks, and some existing efforts have been directed towards vision-language training for Chest X-rays (CXRs). However, we envision that there still exists a potential for improvement in vision-only training for CXRs using ViTs, by aggregating information from multiple scales, which has been proven beneficial for non-transformer networks. Hence, we have developed LT-ViT, a transformer that utilizes combined attention between image tokens and randomly initialized auxiliary tokens that represent labels. Our experiments demonstrate that LT-ViT (1) surpasses the state-of-the-art performance using pure ViTs on two publicly available CXR datasets, (2) is generalizable to other pre-training methods and therefore is agnostic to model initialization, and (3) enables model interpretability without grad-cam and its variants.
A Sensitivity Matrix Approach Using Two-Stage Optimization for Voltage Regulation of LV Networks with High PV Penetration
Aug 23, 2021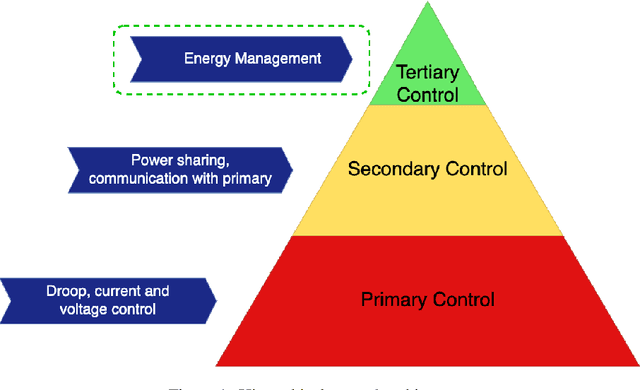
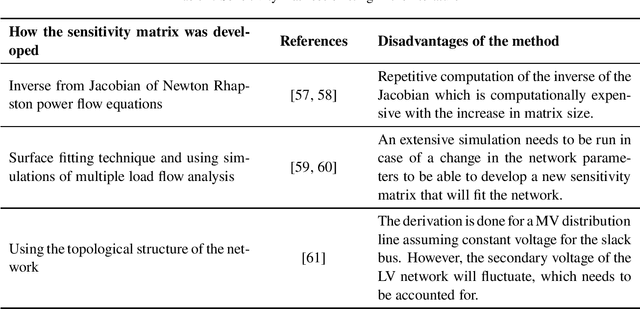
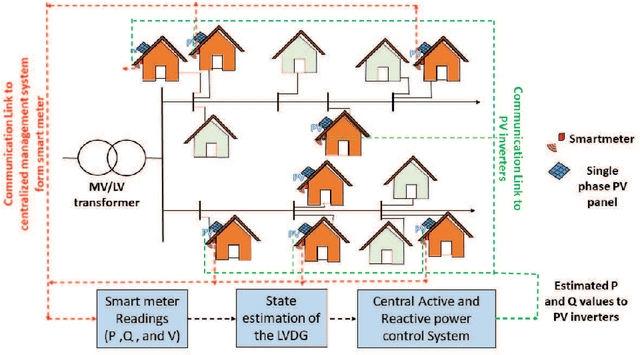

Abstract:The occurrence of voltage violations are a major deterrent for absorbing more roof-top solar power to smart Low Voltage Distribution Grids (LVDG). Recent studies have focused on decentralized control methods to solve this problem due to the high computational time in performing load flows in centralized control techniques. To address this issue a novel sensitivity matrix is developed to estimate voltages of the network by replacing load flow simulations. In this paper, a Centralized Active, Reactive Power Management System (CARPMS) is proposed to optimally utilize the reactive power capability of smart photo-voltaic inverters with minimal active power curtailment to mitigate the voltage violation problem. The developed sensitivity matrix is able to reduce the time consumed by 48% compared to load flow simulations, enabling near real-time control optimization. Given the large solution space of power systems, a novel two-stage optimization is proposed, where the solution space is narrowed down by a Feasible Region Search (FRS) step, followed by Particle Swarm Optimization (PSO). The performance of the proposed methodology is analyzed in comparison to the load flow method to demonstrate the accuracy and the capability of the optimization algorithm to mitigate voltage violations in near real-time. The deviation of mean voltages of the proposed methodology from load flow method was; 6.5*10^-3 p.u for reactive power control using Q-injection, 1.02*10^-2 p.u for reactive power control using Q-absorption, and 0 p.u for active power curtailment case.
A generalized forecasting solution to enable future insights of COVID-19 at sub-national level resolutions
Aug 21, 2021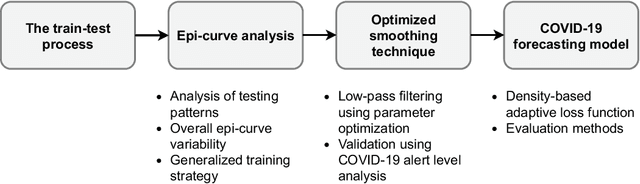
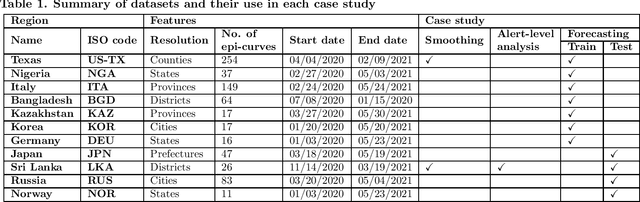
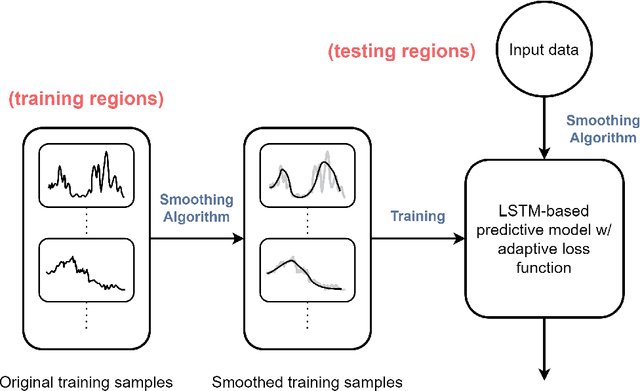
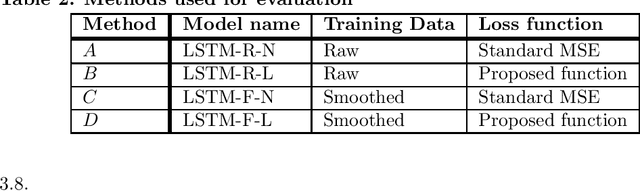
Abstract:COVID-19 continues to cause a significant impact on public health. To minimize this impact, policy makers undertake containment measures that however, when carried out disproportionately to the actual threat, as a result if errorneous threat assessment, cause undesirable long-term socio-economic complications. In addition, macro-level or national level decision making fails to consider the localized sensitivities in small regions. Hence, the need arises for region-wise threat assessments that provide insights on the behaviour of COVID-19 through time, enabled through accurate forecasts. In this study, a forecasting solution is proposed, to predict daily new cases of COVID-19 in regions small enough where containment measures could be locally implemented, by targeting three main shortcomings that exist in literature; the unreliability of existing data caused by inconsistent testing patterns in smaller regions, weak deploy-ability of forecasting models towards predicting cases in previously unseen regions, and model training biases caused by the imbalanced nature of data in COVID-19 epi-curves. Hence, the contributions of this study are three-fold; an optimized smoothing technique to smoothen less deterministic epi-curves based on epidemiological dynamics of that region, a Long-Short-Term-Memory (LSTM) based forecasting model trained using data from select regions to create a representative and diverse training set that maximizes deploy-ability in regions with lack of historical data, and an adaptive loss function whilst training to mitigate the data imbalances seen in epi-curves. The proposed smoothing technique, the generalized training strategy and the adaptive loss function largely increased the overall accuracy of the forecast, which enables efficient containment measures at a more localized micro-level.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge