Udaranga Wickramasinghe
Neural Annotation Refinement: Development of a New 3D Dataset for Adrenal Gland Analysis
Jul 08, 2022



Abstract:The human annotations are imperfect, especially when produced by junior practitioners. Multi-expert consensus is usually regarded as golden standard, while this annotation protocol is too expensive to implement in many real-world projects. In this study, we propose a method to refine human annotation, named Neural Annotation Refinement (NeAR). It is based on a learnable implicit function, which decodes a latent vector into represented shape. By integrating the appearance as an input of implicit functions, the appearance-aware NeAR fixes the annotation artefacts. Our method is demonstrated on the application of adrenal gland analysis. We first show that the NeAR can repair distorted golden standards on a public adrenal gland segmentation dataset. Besides, we develop a new Adrenal gLand ANalysis (ALAN) dataset with the proposed NeAR, where each case consists of a 3D shape of adrenal gland and its diagnosis label (normal vs. abnormal) assigned by experts. We show that models trained on the shapes repaired by the NeAR can diagnose adrenal glands better than the original ones. The ALAN dataset will be open-source, with 1,584 shapes for adrenal gland diagnosis, which serves as a new benchmark for medical shape analysis. Code and dataset are available at https://github.com/M3DV/NeAR.
Deep Active Latent Surfaces for Medical Geometries
Jun 21, 2022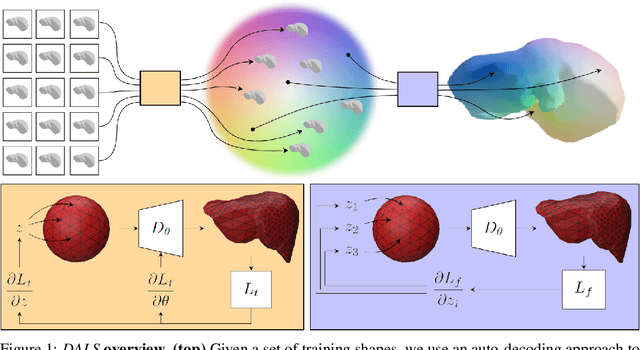
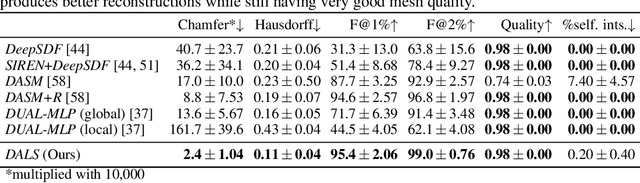

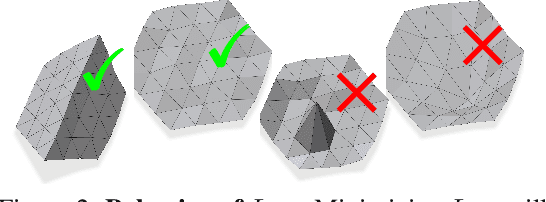
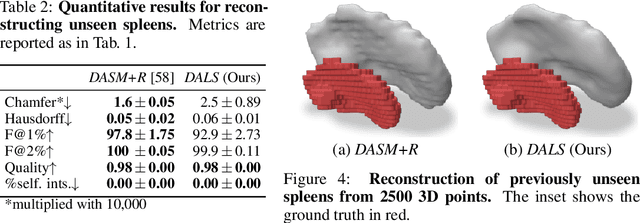
Abstract:Shape priors have long been known to be effective when reconstructing 3D shapes from noisy or incomplete data. When using a deep-learning based shape representation, this often involves learning a latent representation, which can be either in the form of a single global vector or of multiple local ones. The latter allows more flexibility but is prone to overfitting. In this paper, we advocate a hybrid approach representing shapes in terms of 3D meshes with a separate latent vector at each vertex. During training the latent vectors are constrained to have the same value, which avoids overfitting. For inference, the latent vectors are updated independently while imposing spatial regularization constraints. We show that this gives us both flexibility and generalization capabilities, which we demonstrate on several medical image processing tasks.
Weakly Supervised Volumetric Image Segmentation with Deformed Templates
Jun 07, 2021



Abstract:There are many approaches that use weak-supervision to train networks to segment 2D images. By contrast, existing 3D approaches rely on full-supervision of a subset of 2D slices of the 3D image volume. In this paper, we propose an approach that is truly weakly-supervised in the sense that we only need to provide a sparse set of 3D point on the surface of target objects, an easy task that can be quickly done. We use the 3D points to deform a 3D template so that it roughly matches the target object outlines and we introduce an architecture that exploits the supervision provided by coarse template to train a network to find accurate boundaries. We evaluate the performance of our approach on Computed Tomography (CT), Magnetic Resonance Imagery (MRI) and Electron Microscopy (EM) image datasets. We will show that it outperforms a more traditional approach to weak-supervision in 3D at a reduced supervision cost.
Deep Active Surface Models
Nov 20, 2020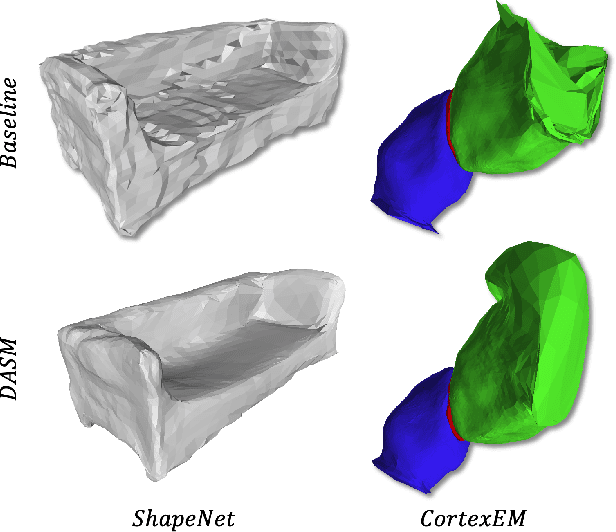
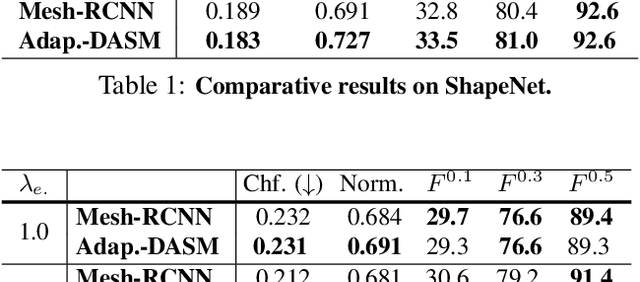
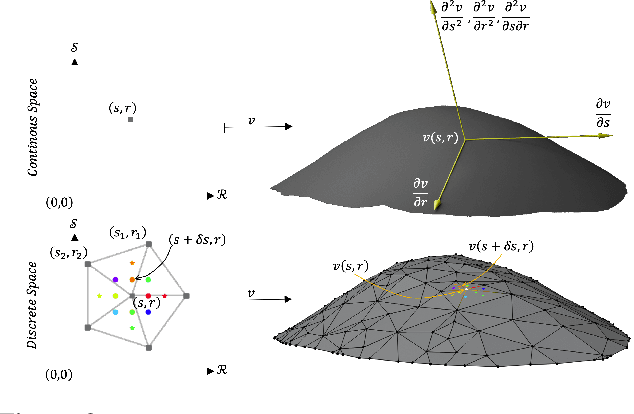
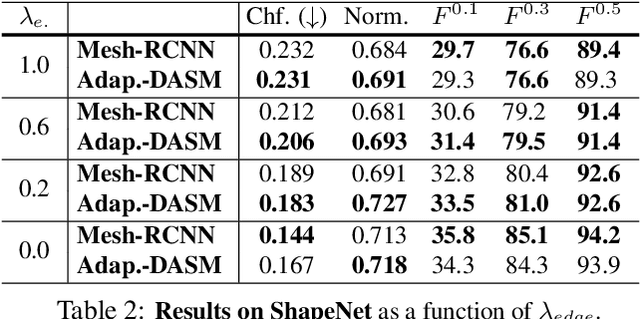
Abstract:Active Surface Models have a long history of being useful to model complex 3D surfaces but only Active Contours have been used in conjunction with deep networks, and then only to produce the data term as well as meta-parameter maps controlling them. In this paper, we advocate a much tighter integration. We introduce layers that implement them that can be integrated seamlessly into Graph Convolutional Networks to enforce sophisticated smoothness priors at an acceptable computational cost. We will show that the resulting Deep Active Surface Models outperform equivalent architectures that use traditional regularization loss terms to impose smoothness priors for 3D surface reconstruction from 2D images and for 3D volume segmentation.
VM-Net: Mesh Modeling to Assist Segmentation in Volumetric Data
Dec 08, 2019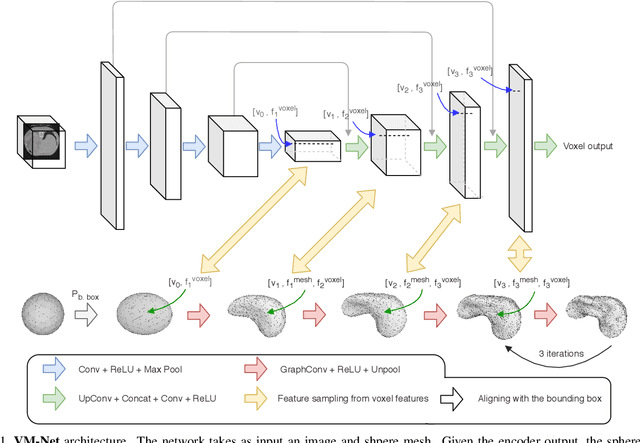
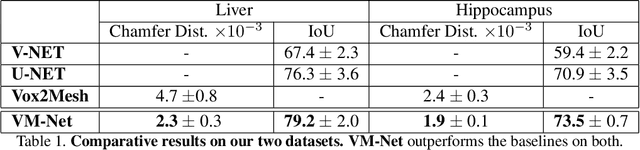
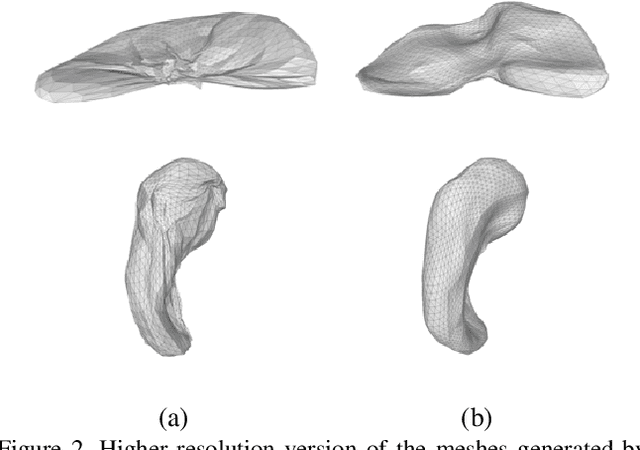
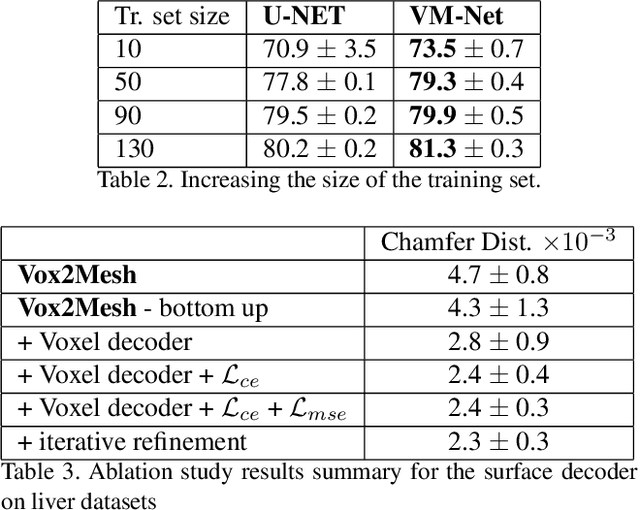
Abstract:CNN-based volumetric methods that label individual voxels now dominate the field of biomedical segmentation. In this paper, we show that simultaneously performing the segmentation and recovering a 3D mesh that models the surface can boost performance. To this end, we propose an end-to-end trainable two-stream encoder/decoder architecture. It comprises a single encoder and two decoders, one that labels voxels and the other outputs the mesh. The key to success is that the two decoders communicate with each other and help each other learn. This goes beyond the well-known fact that training a deep network to perform two different tasks improves its performance. We will demonstrate substantial performance increases on two very different and challenging datasets.
Probabilistic Atlases to Enforce Topological Constraints
Sep 18, 2019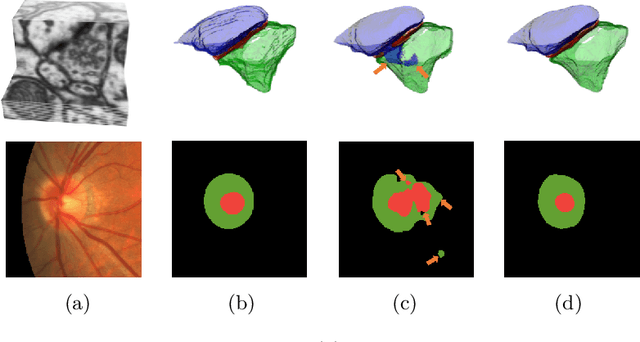
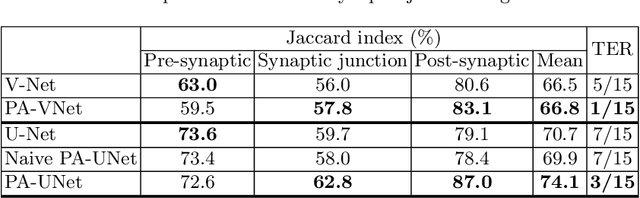
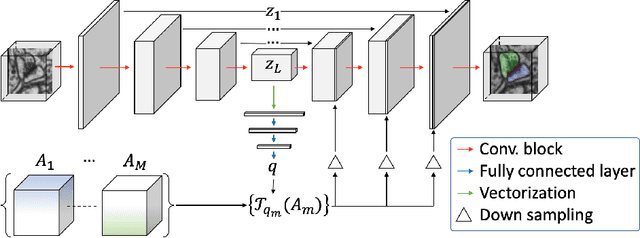
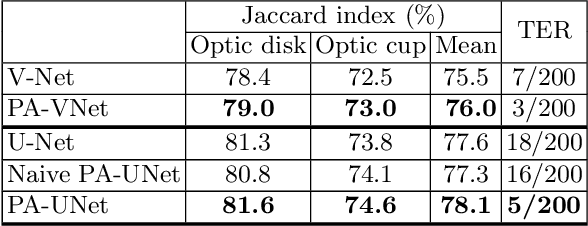
Abstract:Probabilistic atlases (PAs) have long been used in standard segmentation approaches and, more recently, in conjunction with Convolutional Neural Networks (CNNs). However, their use has been restricted to relatively standardized structures such as the brain or heart which have limited or predictable range of deformations. Here we propose an encoding-decoding CNN architecture that can exploit rough atlases that encode only the topology of the target structures that can appear in any pose and have arbitrarily complex shapes to improve the segmentation results. It relies on the output of the encoder to compute both the pose parameters used to deform the atlas and the segmentation mask itself, which makes it effective and end-to-end trainable.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge