Tommi Jaakkola
MIT
Efficiently Controlling Multiple Risks with Pareto Testing
Oct 14, 2022
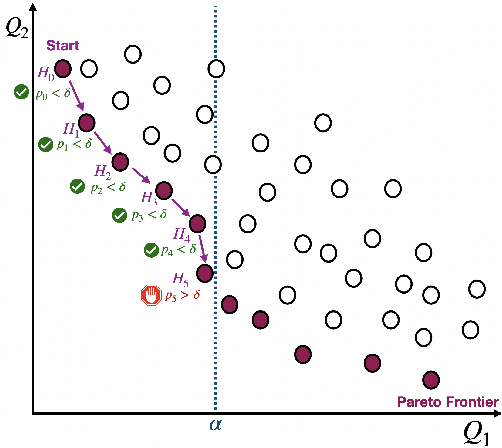
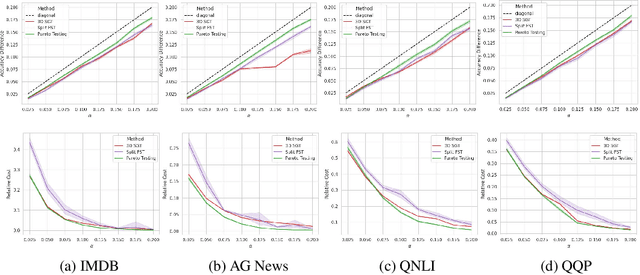
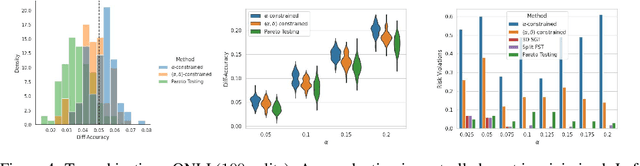
Abstract:Machine learning applications frequently come with multiple diverse objectives and constraints that can change over time. Accordingly, trained models can be tuned with sets of hyper-parameters that affect their predictive behavior (e.g., their run-time efficiency versus error rate). As the number of constraints and hyper-parameter dimensions grow, naively selected settings may lead to sub-optimal and/or unreliable results. We develop an efficient method for calibrating models such that their predictions provably satisfy multiple explicit and simultaneous statistical guarantees (e.g., upper-bounded error rates), while also optimizing any number of additional, unconstrained objectives (e.g., total run-time cost). Building on recent results in distribution-free, finite-sample risk control for general losses, we propose Pareto Testing: a two-stage process which combines multi-objective optimization with multiple hypothesis testing. The optimization stage constructs a set of promising combinations on the Pareto frontier. We then apply statistical testing to this frontier only to identify configurations that have (i) high utility with respect to our objectives, and (ii) guaranteed risk levels with respect to our constraints, with specifiable high probability. We demonstrate the effectiveness of our approach to reliably accelerate the execution of large-scale Transformer models in natural language processing (NLP) applications. In particular, we show how Pareto Testing can be used to dynamically configure multiple inter-dependent model attributes -- including the number of layers computed before exiting, number of attention heads pruned, or number of text tokens considered -- to simultaneously control and optimize various accuracy and cost metrics.
Forces are not Enough: Benchmark and Critical Evaluation for Machine Learning Force Fields with Molecular Simulations
Oct 13, 2022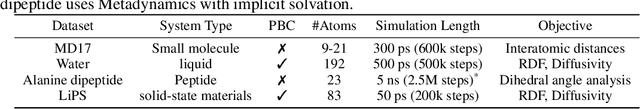
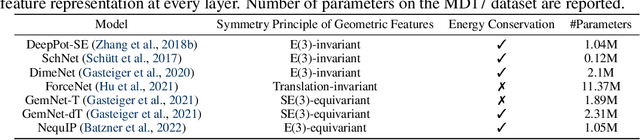
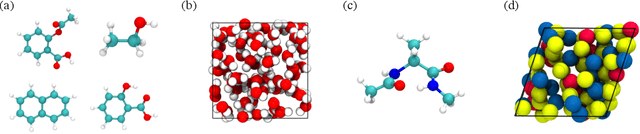

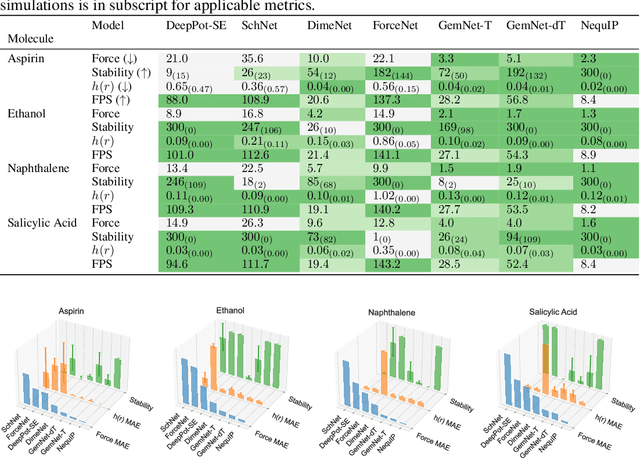
Abstract:Molecular dynamics (MD) simulation techniques are widely used for various natural science applications. Increasingly, machine learning (ML) force field (FF) models begin to replace ab-initio simulations by predicting forces directly from atomic structures. Despite significant progress in this area, such techniques are primarily benchmarked by their force/energy prediction errors, even though the practical use case would be to produce realistic MD trajectories. We aim to fill this gap by introducing a novel benchmark suite for ML MD simulation. We curate representative MD systems, including water, organic molecules, peptide, and materials, and design evaluation metrics corresponding to the scientific objectives of respective systems. We benchmark a collection of state-of-the-art (SOTA) ML FF models and illustrate, in particular, how the commonly benchmarked force accuracy is not well aligned with relevant simulation metrics. We demonstrate when and how selected SOTA methods fail, along with offering directions for further improvement. Specifically, we identify stability as a key metric for ML models to improve. Our benchmark suite comes with a comprehensive open-source codebase for training and simulation with ML FFs to facilitate further work.
DiffDock: Diffusion Steps, Twists, and Turns for Molecular Docking
Oct 04, 2022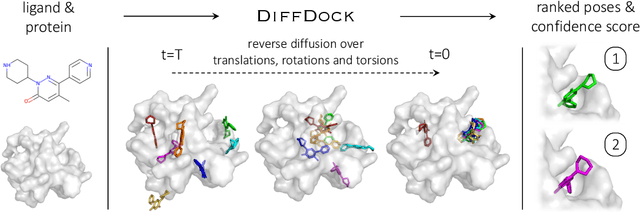
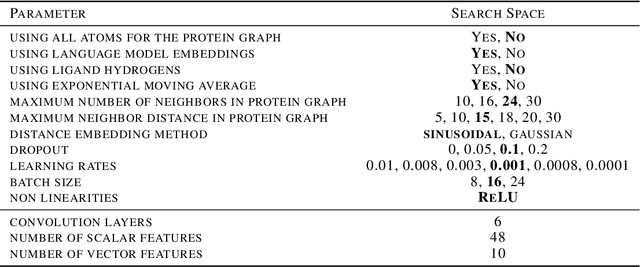
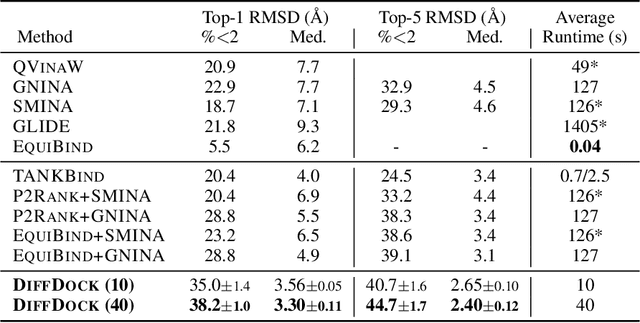
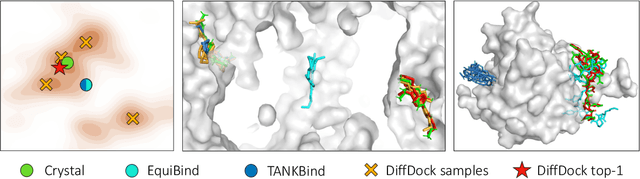

Abstract:Predicting the binding structure of a small molecule ligand to a protein -- a task known as molecular docking -- is critical to drug design. Recent deep learning methods that treat docking as a regression problem have decreased runtime compared to traditional search-based methods but have yet to offer substantial improvements in accuracy. We instead frame molecular docking as a generative modeling problem and develop DiffDock, a diffusion generative model over the non-Euclidean manifold of ligand poses. To do so, we map this manifold to the product space of the degrees of freedom (translational, rotational, and torsional) involved in docking and develop an efficient diffusion process on this space. Empirically, DiffDock obtains a 38% top-1 success rate (RMSD<2A) on PDBBind, significantly outperforming the previous state-of-the-art of traditional docking (23%) and deep learning (20%) methods. Moreover, DiffDock has fast inference times and provides confidence estimates with high selective accuracy.
Poisson Flow Generative Models
Sep 22, 2022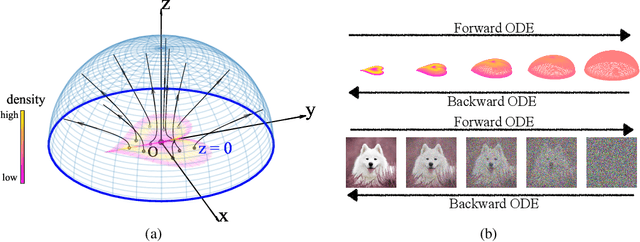
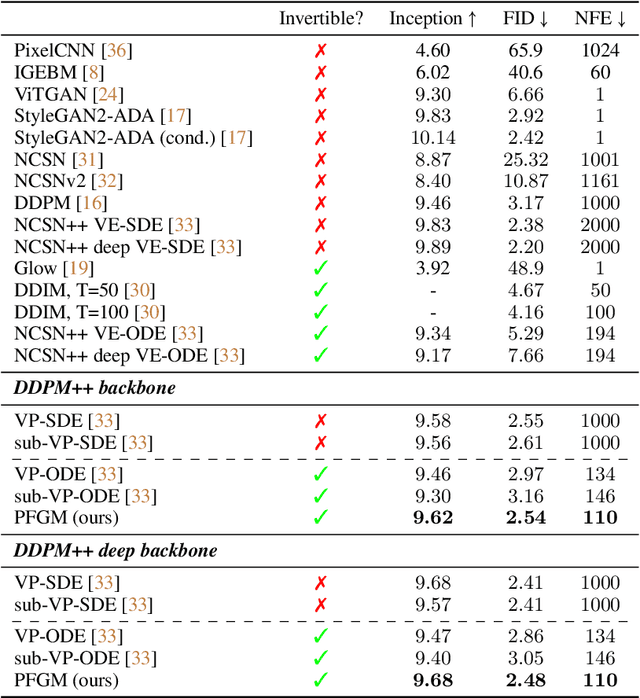
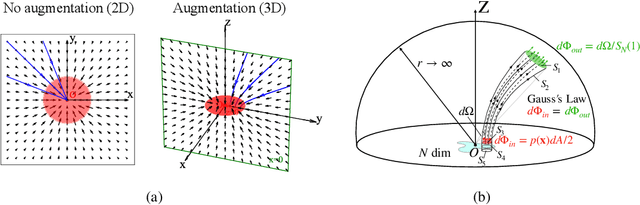

Abstract:We propose a new "Poisson flow" generative model (PFGM) that maps a uniform distribution on a high-dimensional hemisphere into any data distribution. We interpret the data points as electrical charges on the $z=0$ hyperplane in a space augmented with an additional dimension $z$, generating a high-dimensional electric field (the gradient of the solution to Poisson equation). We prove that if these charges flow upward along electric field lines, their initial distribution in the $z=0$ plane transforms into a distribution on the hemisphere of radius $r$ that becomes uniform in the $r \to\infty$ limit. To learn the bijective transformation, we estimate the normalized field in the augmented space. For sampling, we devise a backward ODE that is anchored by the physically meaningful additional dimension: the samples hit the unaugmented data manifold when the $z$ reaches zero. Experimentally, PFGM achieves current state-of-the-art performance among the normalizing flow models on CIFAR-10, with an Inception score of $9.68$ and a FID score of $2.48$. It also performs on par with the state-of-the-art SDE approaches while offering $10\times $ to $20 \times$ acceleration on image generation tasks. Additionally, PFGM appears more tolerant of estimation errors on a weaker network architecture and robust to the step size in the Euler method. The code is available at https://github.com/Newbeeer/poisson_flow .
Calibrated Selective Classification
Aug 25, 2022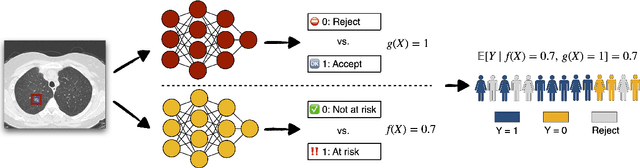
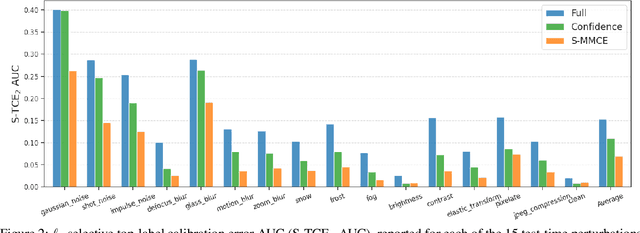
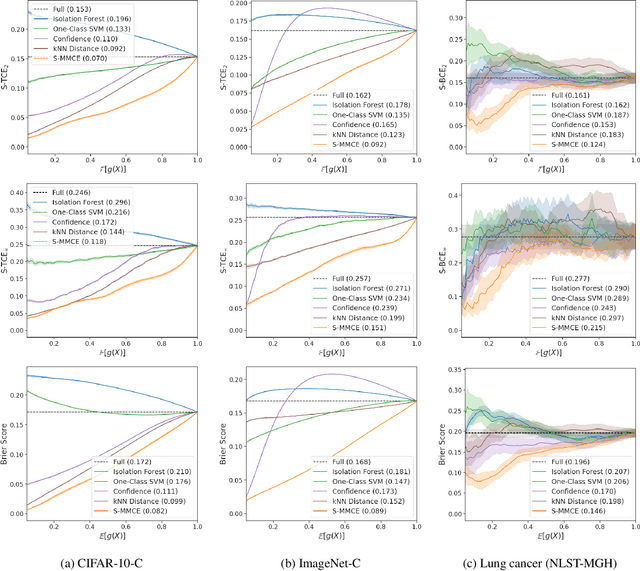
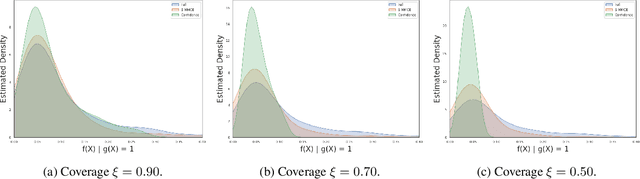
Abstract:Selective classification allows models to abstain from making predictions (e.g., say "I don't know") when in doubt in order to obtain better effective accuracy. While typical selective models can be effective at producing more accurate predictions on average, they may still allow for wrong predictions that have high confidence, or skip correct predictions that have low confidence. Providing calibrated uncertainty estimates alongside predictions -- probabilities that correspond to true frequencies -- can be as important as having predictions that are simply accurate on average. However, uncertainty estimates can be unreliable for certain inputs. In this paper, we develop a new approach to selective classification in which we propose a method for rejecting examples with "uncertain" uncertainties. By doing so, we aim to make predictions with {well-calibrated} uncertainty estimates over the distribution of accepted examples, a property we call selective calibration. We present a framework for learning selectively calibrated models, where a separate selector network is trained to improve the selective calibration error of a given base model. In particular, our work focuses on achieving robust calibration, where the model is intentionally designed to be tested on out-of-domain data. We achieve this through a training strategy inspired by distributionally robust optimization, in which we apply simulated input perturbations to the known, in-domain training data. We demonstrate the empirical effectiveness of our approach on multiple image classification and lung cancer risk assessment tasks.
Antibody-Antigen Docking and Design via Hierarchical Equivariant Refinement
Jul 14, 2022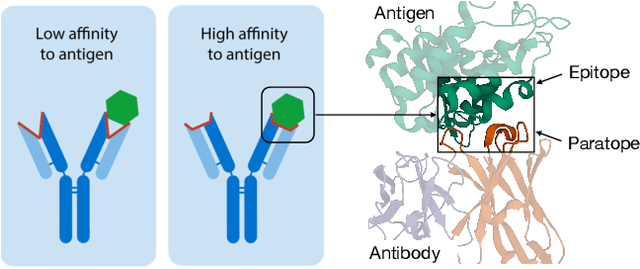
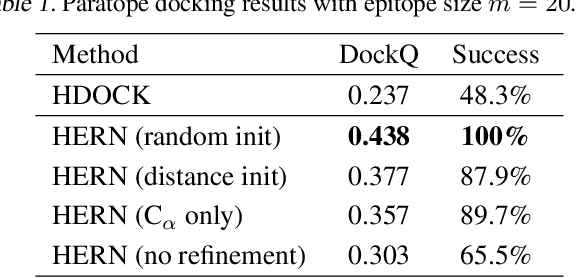
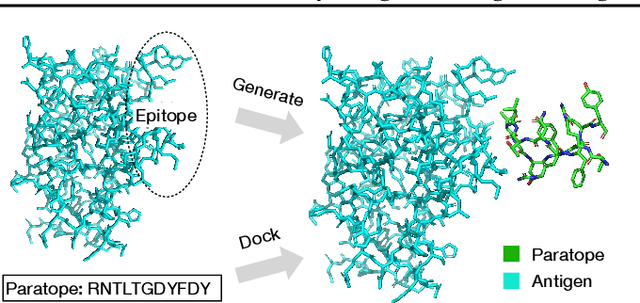
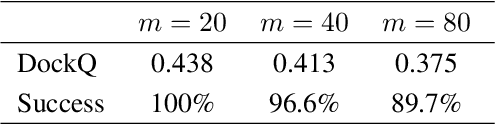
Abstract:Computational antibody design seeks to automatically create an antibody that binds to an antigen. The binding affinity is governed by the 3D binding interface where antibody residues (paratope) closely interact with antigen residues (epitope). Thus, predicting 3D paratope-epitope complex (docking) is the key to finding the best paratope. In this paper, we propose a new model called Hierarchical Equivariant Refinement Network (HERN) for paratope docking and design. During docking, HERN employs a hierarchical message passing network to predict atomic forces and use them to refine a binding complex in an iterative, equivariant manner. During generation, its autoregressive decoder progressively docks generated paratopes and builds a geometric representation of the binding interface to guide the next residue choice. Our results show that HERN significantly outperforms prior state-of-the-art on paratope docking and design benchmarks.
Diffusion probabilistic modeling of protein backbones in 3D for the motif-scaffolding problem
Jun 08, 2022



Abstract:Construction of a scaffold structure that supports a desired motif, conferring protein function, shows promise for the design of vaccines and enzymes. But a general solution to this motif-scaffolding problem remains open. Current machine-learning techniques for scaffold design are either limited to unrealistically small scaffolds (up to length 20) or struggle to produce multiple diverse scaffolds. We propose to learn a distribution over diverse and longer protein backbone structures via an E(3)-equivariant graph neural network. We develop SMCDiff to efficiently sample scaffolds from this distribution conditioned on a given motif; our algorithm is the first to theoretically guarantee conditional samples from a diffusion model in the large-compute limit. We evaluate our designed backbones by how well they align with AlphaFold2-predicted structures. We show that our method can (1) sample scaffolds up to 80 residues and (2) achieve structurally diverse scaffolds for a fixed motif.
Torsional Diffusion for Molecular Conformer Generation
Jun 01, 2022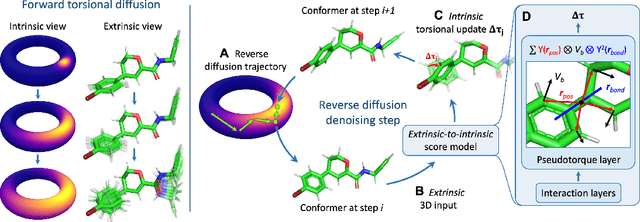

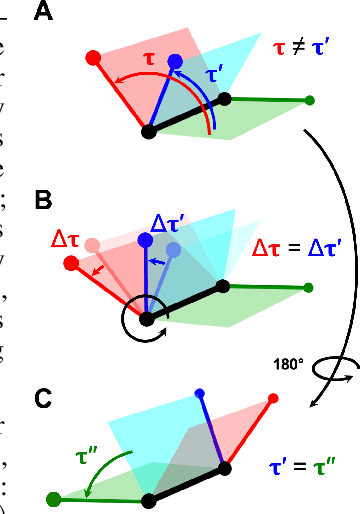
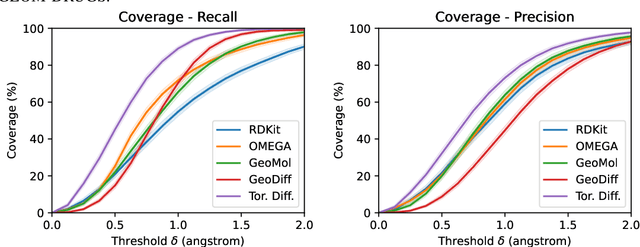
Abstract:Molecular conformer generation is a fundamental task in computational chemistry. Several machine learning approaches have been developed, but none have outperformed state-of-the-art cheminformatics methods. We propose torsional diffusion, a novel diffusion framework that operates on the space of torsion angles via a diffusion process on the hypertorus and an extrinsic-to-intrinsic score model. On a standard benchmark of drug-like molecules, torsional diffusion generates superior conformer ensembles compared to machine learning and cheminformatics methods in terms of both RMSD and chemical properties, and is orders of magnitude faster than previous diffusion-based models. Moreover, our model provides exact likelihoods, which we employ to build the first generalizable Boltzmann generator. Code is available at https://github.com/gcorso/torsional-diffusion.
Subspace Diffusion Generative Models
May 03, 2022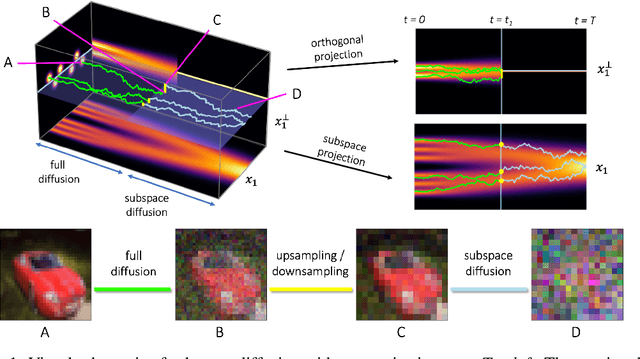
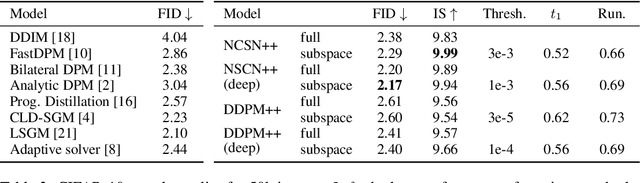
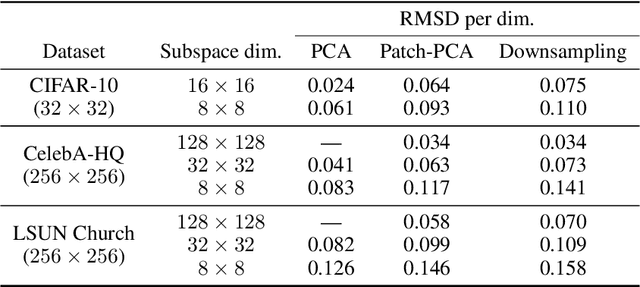
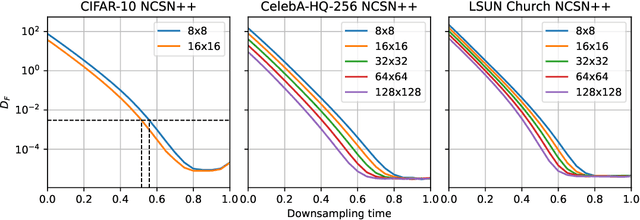

Abstract:Score-based models generate samples by mapping noise to data (and vice versa) via a high-dimensional diffusion process. We question whether it is necessary to run this entire process at high dimensionality and incur all the inconveniences thereof. Instead, we restrict the diffusion via projections onto subspaces as the data distribution evolves toward noise. When applied to state-of-the-art models, our framework simultaneously improves sample quality -- reaching an FID of 2.17 on unconditional CIFAR-10 -- and reduces the computational cost of inference for the same number of denoising steps. Our framework is fully compatible with continuous-time diffusion and retains its flexible capabilities, including exact log-likelihoods and controllable generation. Code is available at https://github.com/bjing2016/subspace-diffusion.
Simulate Time-integrated Coarse-grained Molecular Dynamics with Geometric Machine Learning
Apr 21, 2022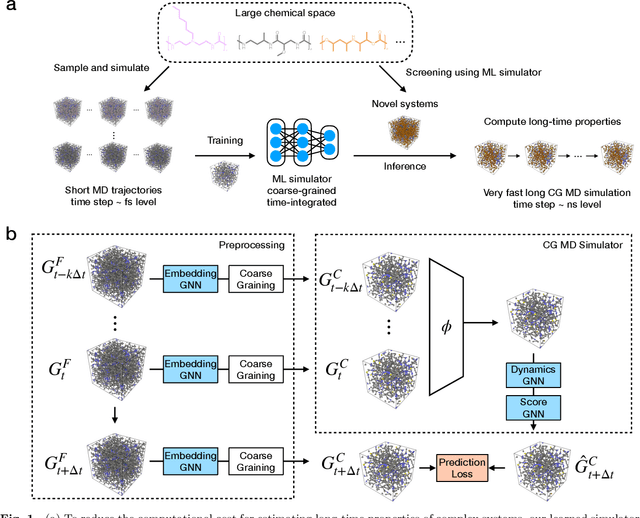

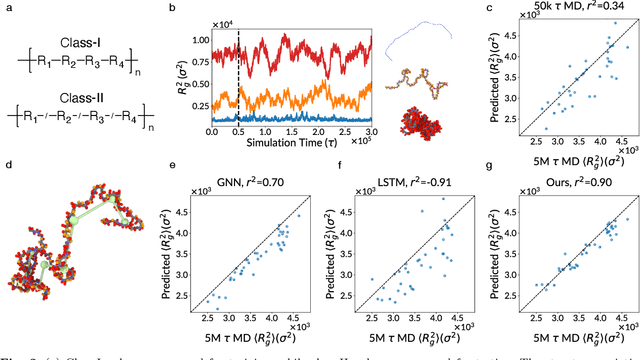
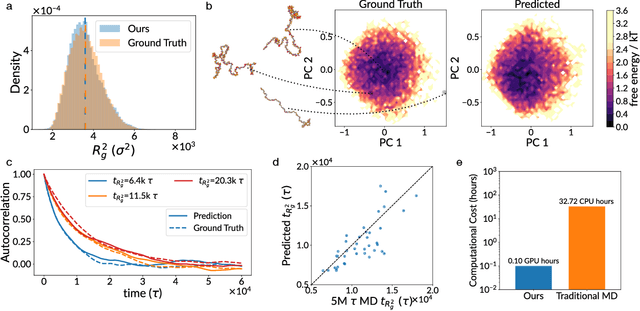
Abstract:Molecular dynamics (MD) simulation is the workhorse of various scientific domains but is limited by high computational cost. Learning-based force fields have made major progress in accelerating ab-initio MD simulation but are still not fast enough for many real-world applications that require long-time MD simulation. In this paper, we adopt a different machine learning approach where we coarse-grain a physical system using graph clustering, and model the system evolution with a very large time-integration step using graph neural networks. A novel score-based GNN refinement module resolves the long-standing challenge of long-time simulation instability. Despite only trained with short MD trajectory data, our learned simulator can generalize to unseen novel systems and simulate for much longer than the training trajectories. Properties requiring 10-100 ns level long-time dynamics can be accurately recovered at several-orders-of-magnitude higher speed than classical force fields. We demonstrate the effectiveness of our method on two realistic complex systems: (1) single-chain coarse-grained polymers in implicit solvent; (2) multi-component Li-ion polymer electrolyte systems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge