Thien Huu Nguyen
PhoGPT: Generative Pre-training for Vietnamese
Nov 06, 2023
Abstract:We open-source a state-of-the-art 7.5B-parameter generative model series named PhoGPT for Vietnamese, which includes the base pre-trained monolingual model PhoGPT-7B5 and its instruction-following variant, PhoGPT-7B5-Instruct. In addition, we also demonstrate its superior performance compared to previous open-source models through a human evaluation experiment. GitHub: https://github.com/VinAIResearch/PhoGPT
CulturaX: A Cleaned, Enormous, and Multilingual Dataset for Large Language Models in 167 Languages
Sep 17, 2023

Abstract:The driving factors behind the development of large language models (LLMs) with impressive learning capabilities are their colossal model sizes and extensive training datasets. Along with the progress in natural language processing, LLMs have been frequently made accessible to the public to foster deeper investigation and applications. However, when it comes to training datasets for these LLMs, especially the recent state-of-the-art models, they are often not fully disclosed. Creating training data for high-performing LLMs involves extensive cleaning and deduplication to ensure the necessary level of quality. The lack of transparency for training data has thus hampered research on attributing and addressing hallucination and bias issues in LLMs, hindering replication efforts and further advancements in the community. These challenges become even more pronounced in multilingual learning scenarios, where the available multilingual text datasets are often inadequately collected and cleaned. Consequently, there is a lack of open-source and readily usable dataset to effectively train LLMs in multiple languages. To overcome this issue, we present CulturaX, a substantial multilingual dataset with 6.3 trillion tokens in 167 languages, tailored for LLM development. Our dataset undergoes meticulous cleaning and deduplication through a rigorous pipeline of multiple stages to accomplish the best quality for model training, including language identification, URL-based filtering, metric-based cleaning, document refinement, and data deduplication. CulturaX is fully released to the public in HuggingFace to facilitate research and advancements in multilingual LLMs: https://huggingface.co/datasets/uonlp/CulturaX.
Okapi: Instruction-tuned Large Language Models in Multiple Languages with Reinforcement Learning from Human Feedback
Aug 02, 2023



Abstract:A key technology for the development of large language models (LLMs) involves instruction tuning that helps align the models' responses with human expectations to realize impressive learning abilities. Two major approaches for instruction tuning characterize supervised fine-tuning (SFT) and reinforcement learning from human feedback (RLHF), which are currently applied to produce the best commercial LLMs (e.g., ChatGPT). To improve the accessibility of LLMs for research and development efforts, various instruction-tuned open-source LLMs have also been introduced recently, e.g., Alpaca, Vicuna, to name a few. However, existing open-source LLMs have only been instruction-tuned for English and a few popular languages, thus hindering their impacts and accessibility to many other languages in the world. Among a few very recent work to explore instruction tuning for LLMs in multiple languages, SFT has been used as the only approach to instruction-tune LLMs for multiple languages. This has left a significant gap for fine-tuned LLMs based on RLHF in diverse languages and raised important questions on how RLHF can boost the performance of multilingual instruction tuning. To overcome this issue, we present Okapi, the first system with instruction-tuned LLMs based on RLHF for multiple languages. Okapi introduces instruction and response-ranked data in 26 diverse languages to facilitate the experiments and development of future multilingual LLM research. We also present benchmark datasets to enable the evaluation of generative LLMs in multiple languages. Our experiments demonstrate the advantages of RLHF for multilingual instruction over SFT for different base models and datasets. Our framework and resources are released at https://github.com/nlp-uoregon/Okapi.
Boosting Punctuation Restoration with Data Generation and Reinforcement Learning
Jul 24, 2023Abstract:Punctuation restoration is an important task in automatic speech recognition (ASR) which aim to restore the syntactic structure of generated ASR texts to improve readability. While punctuated texts are abundant from written documents, the discrepancy between written punctuated texts and ASR texts limits the usability of written texts in training punctuation restoration systems for ASR texts. This paper proposes a reinforcement learning method to exploit in-topic written texts and recent advances in large pre-trained generative language models to bridge this gap. The experiments show that our method achieves state-of-the-art performance on the ASR test set on two benchmark datasets for punctuation restoration.
Question-Context Alignment and Answer-Context Dependencies for Effective Answer Sentence Selection
Jun 03, 2023



Abstract:Answer sentence selection (AS2) in open-domain question answering finds answer for a question by ranking candidate sentences extracted from web documents. Recent work exploits answer context, i.e., sentences around a candidate, by incorporating them as additional input string to the Transformer models to improve the correctness scoring. In this paper, we propose to improve the candidate scoring by explicitly incorporating the dependencies between question-context and answer-context into the final representation of a candidate. Specifically, we use Optimal Transport to compute the question-based dependencies among sentences in the passage where the answer is extracted from. We then represent these dependencies as edges in a graph and use Graph Convolutional Network to derive the representation of a candidate, a node in the graph. Our proposed model achieves significant improvements on popular AS2 benchmarks, i.e., WikiQA and WDRASS, obtaining new state-of-the-art on all benchmarks.
ChatGPT Beyond English: Towards a Comprehensive Evaluation of Large Language Models in Multilingual Learning
Apr 12, 2023



Abstract:Over the last few years, large language models (LLMs) have emerged as the most important breakthroughs in natural language processing (NLP) that fundamentally transform research and developments in the field. ChatGPT represents one of the most exciting LLM systems developed recently to showcase impressive skills for language generation and highly attract public attention. Among various exciting applications discovered for ChatGPT in English, the model can process and generate texts for multiple languages due to its multilingual training data. Given the broad adoption of ChatGPT for English in different problems and areas, a natural question is whether ChatGPT can also be applied effectively for other languages or it is necessary to develop more language-specific technologies. The answer to this question requires a thorough evaluation of ChatGPT over multiple tasks with diverse languages and large datasets (i.e., beyond reported anecdotes), which is still missing or limited in current research. Our work aims to fill this gap for the evaluation of ChatGPT and similar LLMs to provide more comprehensive information for multilingual NLP applications. While this work will be an ongoing effort to include additional experiments in the future, our current paper evaluates ChatGPT on 7 different tasks, covering 37 diverse languages with high, medium, low, and extremely low resources. We also focus on the zero-shot learning setting for ChatGPT to improve reproducibility and better simulate the interactions of general users. Compared to the performance of previous models, our extensive experimental results demonstrate a worse performance of ChatGPT for different NLP tasks and languages, calling for further research to develop better models and understanding for multilingual learning.
MEE: A Novel Multilingual Event Extraction Dataset
Nov 17, 2022



Abstract:Event Extraction (EE) is one of the fundamental tasks in Information Extraction (IE) that aims to recognize event mentions and their arguments (i.e., participants) from text. Due to its importance, extensive methods and resources have been developed for Event Extraction. However, one limitation of current research for EE involves the under-exploration for non-English languages in which the lack of high-quality multilingual EE datasets for model training and evaluation has been the main hindrance. To address this limitation, we propose a novel Multilingual Event Extraction dataset (MEE) that provides annotation for more than 50K event mentions in 8 typologically different languages. MEE comprehensively annotates data for entity mentions, event triggers and event arguments. We conduct extensive experiments on the proposed dataset to reveal challenges and opportunities for multilingual EE.
MINION: a Large-Scale and Diverse Dataset for Multilingual Event Detection
Nov 17, 2022



Abstract:Event Detection (ED) is the task of identifying and classifying trigger words of event mentions in text. Despite considerable research efforts in recent years for English text, the task of ED in other languages has been significantly less explored. Switching to non-English languages, important research questions for ED include how well existing ED models perform on different languages, how challenging ED is in other languages, and how well ED knowledge and annotation can be transferred across languages. To answer those questions, it is crucial to obtain multilingual ED datasets that provide consistent event annotation for multiple languages. There exist some multilingual ED datasets; however, they tend to cover a handful of languages and mainly focus on popular ones. Many languages are not covered in existing multilingual ED datasets. In addition, the current datasets are often small and not accessible to the public. To overcome those shortcomings, we introduce a new large-scale multilingual dataset for ED (called MINION) that consistently annotates events for 8 different languages; 5 of them have not been supported by existing multilingual datasets. We also perform extensive experiments and analysis to demonstrate the challenges and transferability of ED across languages in MINION that in all call for more research effort in this area.
Textual Data Augmentation for Patient Outcomes Prediction
Nov 13, 2022



Abstract:Deep learning models have demonstrated superior performance in various healthcare applications. However, the major limitation of these deep models is usually the lack of high-quality training data due to the private and sensitive nature of this field. In this study, we propose a novel textual data augmentation method to generate artificial clinical notes in patients' Electronic Health Records (EHRs) that can be used as additional training data for patient outcomes prediction. Essentially, we fine-tune the generative language model GPT-2 to synthesize labeled text with the original training data. More specifically, We propose a teacher-student framework where we first pre-train a teacher model on the original data, and then train a student model on the GPT-augmented data under the guidance of the teacher. We evaluate our method on the most common patient outcome, i.e., the 30-day readmission rate. The experimental results show that deep models can improve their predictive performance with the augmented data, indicating the effectiveness of the proposed architecture.
Tutorial Recommendation for Livestream Videos using Discourse-Level Consistency and Ontology-Based Filtering
Sep 11, 2022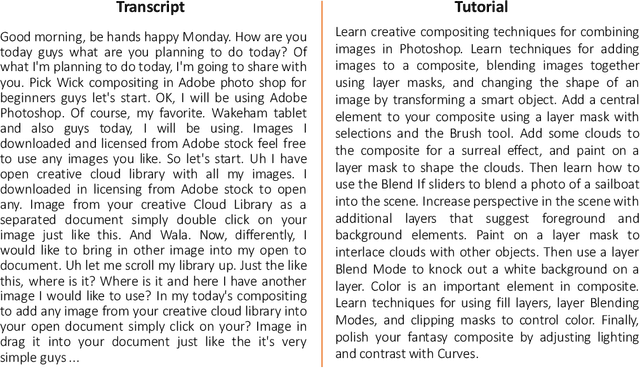
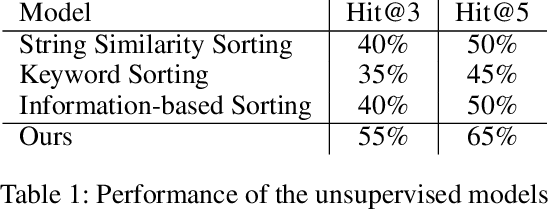
Abstract:Streaming videos is one of the methods for creators to share their creative works with their audience. In these videos, the streamer share how they achieve their final objective by using various tools in one or several programs for creative projects. To this end, the steps required to achieve the final goal can be discussed. As such, these videos could provide substantial educational content that can be used to learn how to employ the tools used by the streamer. However, one of the drawbacks is that the streamer might not provide enough details for every step. Therefore, for the learners, it might be difficult to catch up with all the steps. In order to alleviate this issue, one solution is to link the streaming videos with the relevant tutorial available for the tools used in the streaming video. More specifically, a system can analyze the content of the live streaming video and recommend the most relevant tutorials. Since the existing document recommendation models cannot handle this situation, in this work, we present a novel dataset and model for the task of tutorial recommendation for live-streamed videos. We conduct extensive analyses on the proposed dataset and models, revealing the challenging nature of this task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge