Theodore L. Willke
Inductive Representation Learning in Large Attributed Graphs
Nov 22, 2017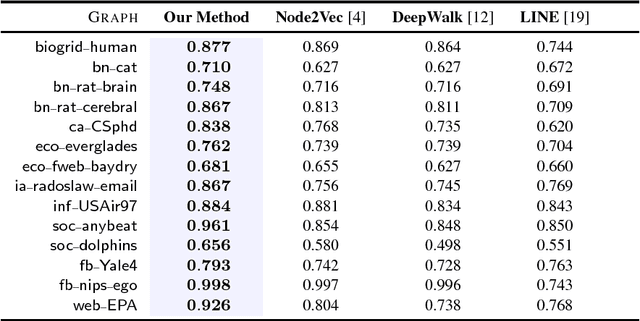
Abstract:Graphs (networks) are ubiquitous and allow us to model entities (nodes) and the dependencies (edges) between them. Learning a useful feature representation from graph data lies at the heart and success of many machine learning tasks such as classification, anomaly detection, link prediction, among many others. Many existing techniques use random walks as a basis for learning features or estimating the parameters of a graph model for a downstream prediction task. Examples include recent node embedding methods such as DeepWalk, node2vec, as well as graph-based deep learning algorithms. However, the simple random walk used by these methods is fundamentally tied to the identity of the node. This has three main disadvantages. First, these approaches are inherently transductive and do not generalize to unseen nodes and other graphs. Second, they are not space-efficient as a feature vector is learned for each node which is impractical for large graphs. Third, most of these approaches lack support for attributed graphs. To make these methods more generally applicable, we propose a framework for inductive network representation learning based on the notion of attributed random walk that is not tied to node identity and is instead based on learning a function $\Phi : \mathrm{\rm \bf x} \rightarrow w$ that maps a node attribute vector $\mathrm{\rm \bf x}$ to a type $w$. This framework serves as a basis for generalizing existing methods such as DeepWalk, node2vec, and many other previous methods that leverage traditional random walks.
Segmenting Brain Tumors with Symmetry
Nov 17, 2017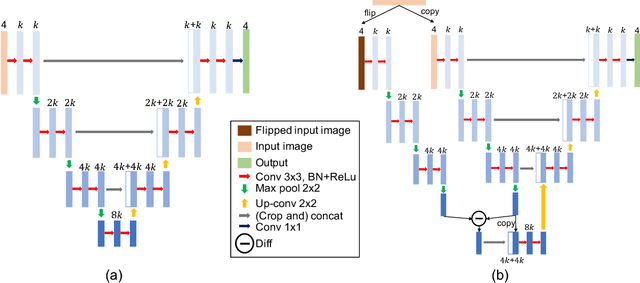
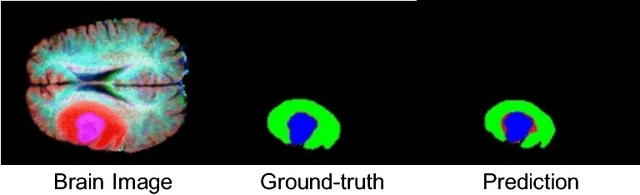

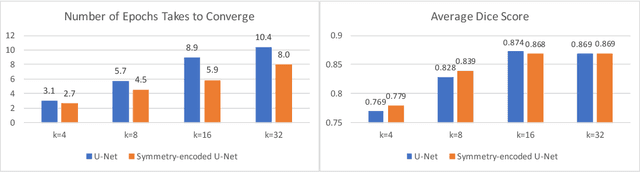

Abstract:We explore encoding brain symmetry into a neural network for a brain tumor segmentation task. A healthy human brain is symmetric at a high level of abstraction, and the high-level asymmetric parts are more likely to be tumor regions. Paying more attention to asymmetries has the potential to boost the performance in brain tumor segmentation. We propose a method to encode brain symmetry into existing neural networks and apply the method to a state-of-the-art neural network for medical imaging segmentation. We evaluate our symmetry-encoded network on the dataset from a brain tumor segmentation challenge and verify that the new model extracts information in the training images more efficiently than the original model.
A Framework for Generalizing Graph-based Representation Learning Methods
Sep 14, 2017
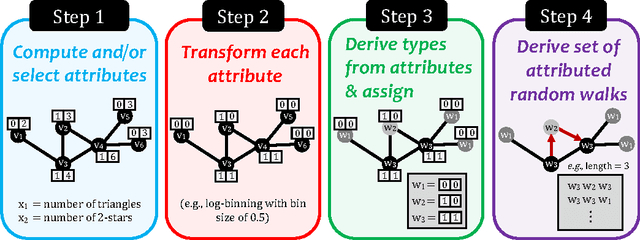
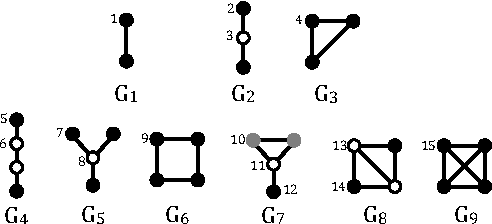
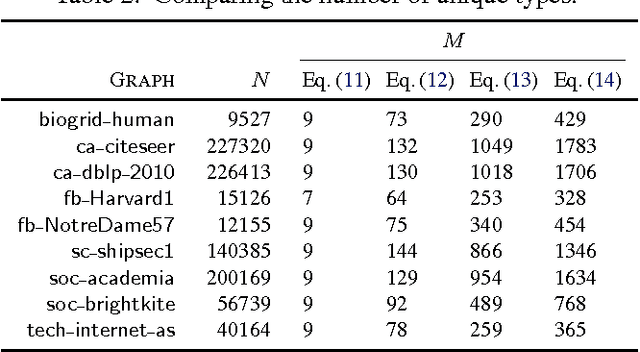
Abstract:Random walks are at the heart of many existing deep learning algorithms for graph data. However, such algorithms have many limitations that arise from the use of random walks, e.g., the features resulting from these methods are unable to transfer to new nodes and graphs as they are tied to node identity. In this work, we introduce the notion of attributed random walks which serves as a basis for generalizing existing methods such as DeepWalk, node2vec, and many others that leverage random walks. Our proposed framework enables these methods to be more widely applicable for both transductive and inductive learning as well as for use on graphs with attributes (if available). This is achieved by learning functions that generalize to new nodes and graphs. We show that our proposed framework is effective with an average AUC improvement of 16.1% while requiring on average 853 times less space than existing methods on a variety of graphs from several domains.
Revisiting Role Discovery in Networks: From Node to Edge Roles
Nov 07, 2016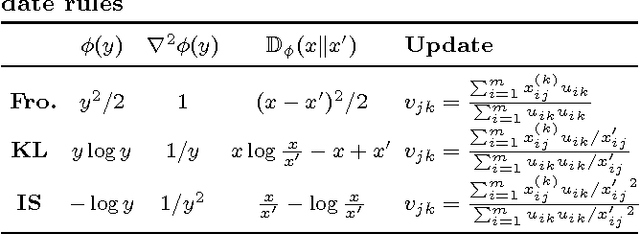
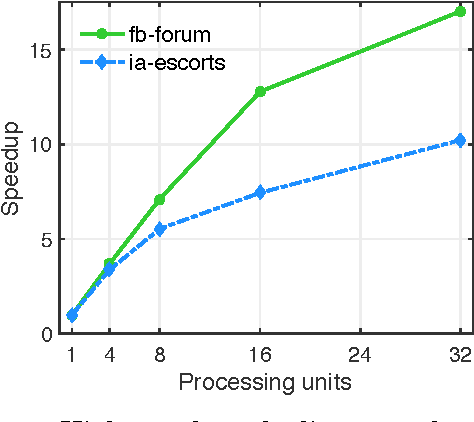
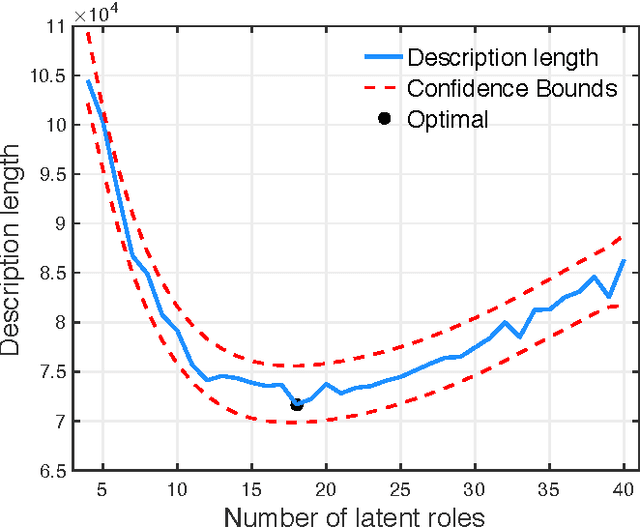
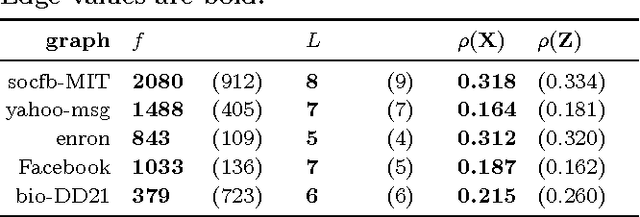
Abstract:Previous work in network analysis has focused on modeling the mixed-memberships of node roles in the graph, but not the roles of edges. We introduce the edge role discovery problem and present a generalizable framework for learning and extracting edge roles from arbitrary graphs automatically. Furthermore, while existing node-centric role models have mainly focused on simple degree and egonet features, this work also explores graphlet features for role discovery. In addition, we also develop an approach for automatically learning and extracting important and useful edge features from an arbitrary graph. The experimental results demonstrate the utility of edge roles for network analysis tasks on a variety of graphs from various problem domains.
A Searchlight Factor Model Approach for Locating Shared Information in Multi-Subject fMRI Analysis
Sep 29, 2016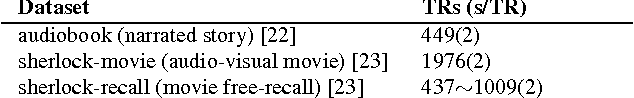
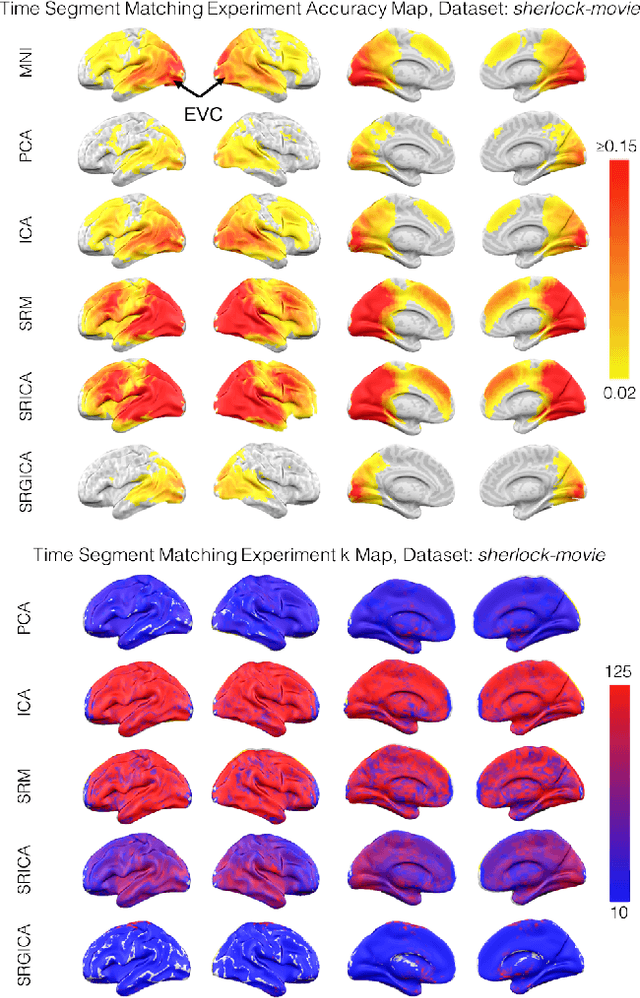
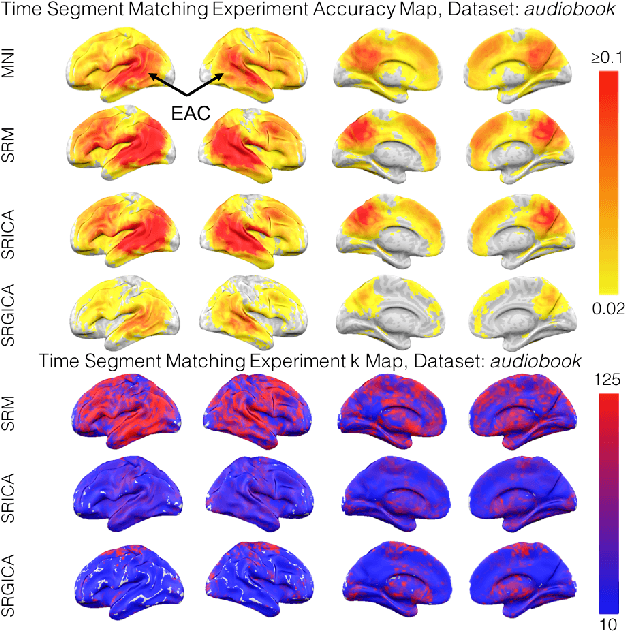
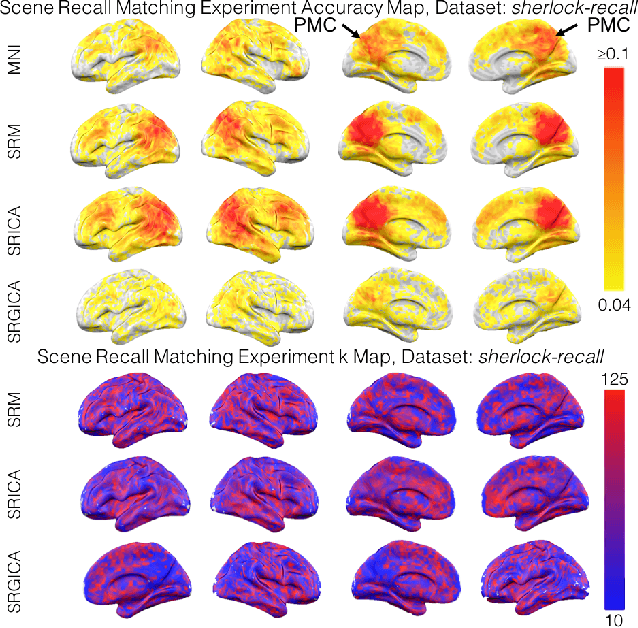
Abstract:There is a growing interest in joint multi-subject fMRI analysis. The challenge of such analysis comes from inherent anatomical and functional variability across subjects. One approach to resolving this is a shared response factor model. This assumes a shared and time synchronized stimulus across subjects. Such a model can often identify shared information, but it may not be able to pinpoint with high resolution the spatial location of this information. In this work, we examine a searchlight based shared response model to identify shared information in small contiguous regions (searchlights) across the whole brain. Validation using classification tasks demonstrates that we can pinpoint informative local regions.
Enabling Factor Analysis on Thousand-Subject Neuroimaging Datasets
Aug 18, 2016
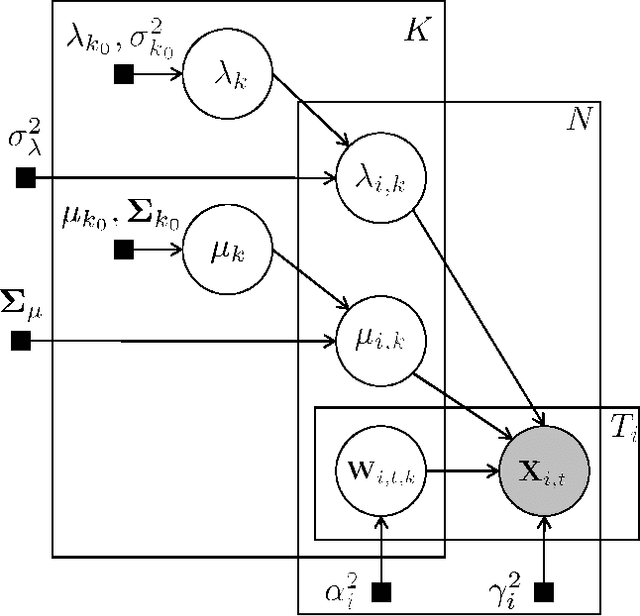
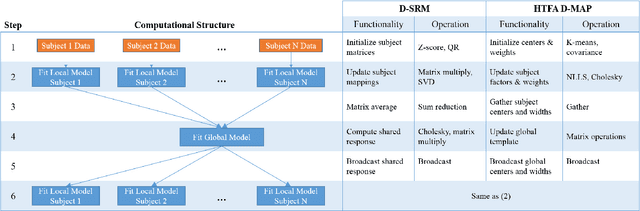
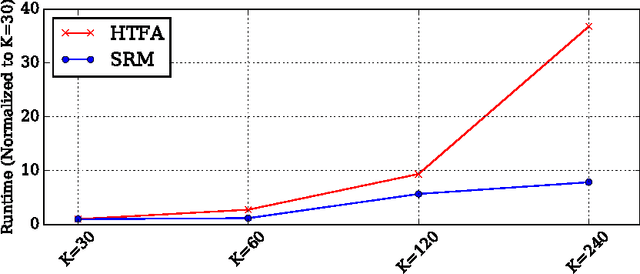
Abstract:The scale of functional magnetic resonance image data is rapidly increasing as large multi-subject datasets are becoming widely available and high-resolution scanners are adopted. The inherent low-dimensionality of the information in this data has led neuroscientists to consider factor analysis methods to extract and analyze the underlying brain activity. In this work, we consider two recent multi-subject factor analysis methods: the Shared Response Model and Hierarchical Topographic Factor Analysis. We perform analytical, algorithmic, and code optimization to enable multi-node parallel implementations to scale. Single-node improvements result in 99x and 1812x speedups on these two methods, and enables the processing of larger datasets. Our distributed implementations show strong scaling of 3.3x and 5.5x respectively with 20 nodes on real datasets. We also demonstrate weak scaling on a synthetic dataset with 1024 subjects, on up to 1024 nodes and 32,768 cores.
A Convolutional Autoencoder for Multi-Subject fMRI Data Aggregation
Aug 17, 2016
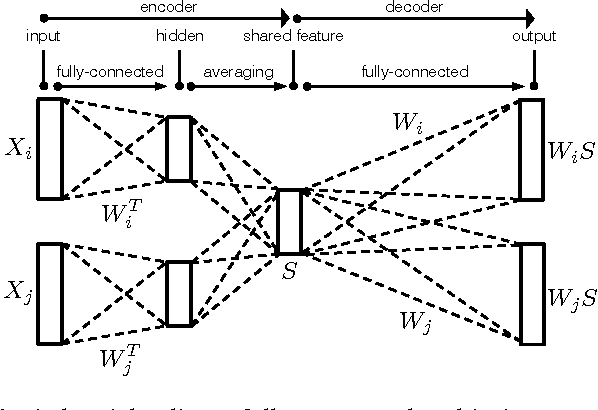
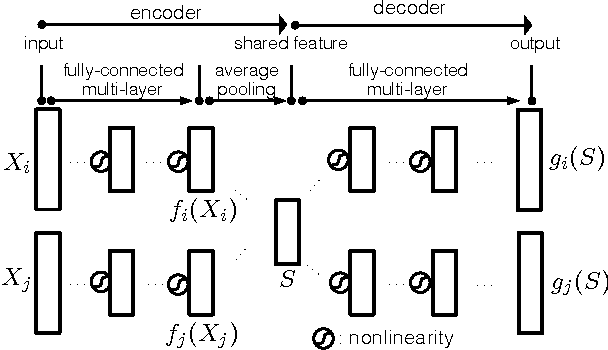
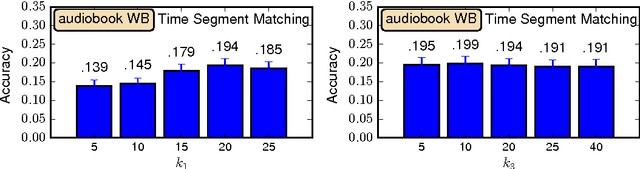
Abstract:Finding the most effective way to aggregate multi-subject fMRI data is a long-standing and challenging problem. It is of increasing interest in contemporary fMRI studies of human cognition due to the scarcity of data per subject and the variability of brain anatomy and functional response across subjects. Recent work on latent factor models shows promising results in this task but this approach does not preserve spatial locality in the brain. We examine two ways to combine the ideas of a factor model and a searchlight based analysis to aggregate multi-subject fMRI data while preserving spatial locality. We first do this directly by combining a recent factor method known as a shared response model with searchlight analysis. Then we design a multi-view convolutional autoencoder for the same task. Both approaches preserve spatial locality and have competitive or better performance compared with standard searchlight analysis and the shared response model applied across the whole brain. We also report a system design to handle the computational challenge of training the convolutional autoencoder.
Graphlet Decomposition: Framework, Algorithms, and Applications
Feb 15, 2016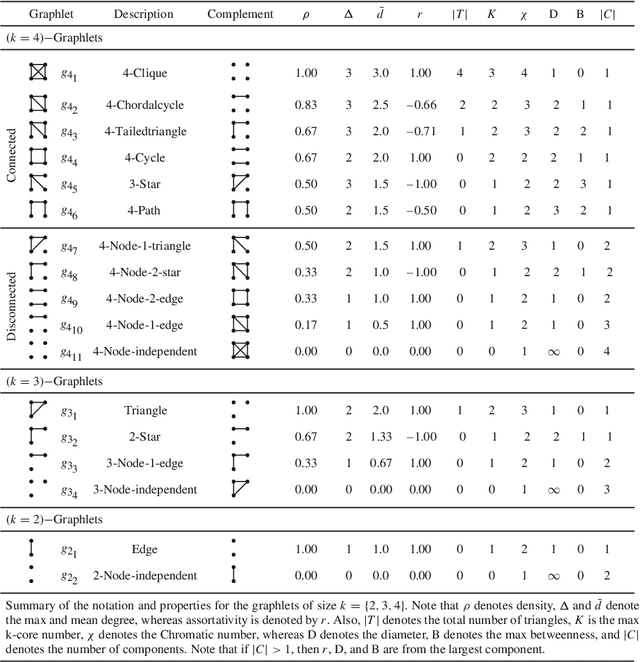
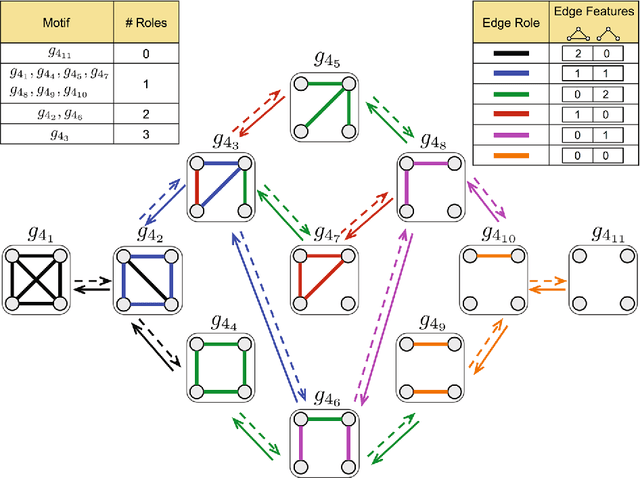
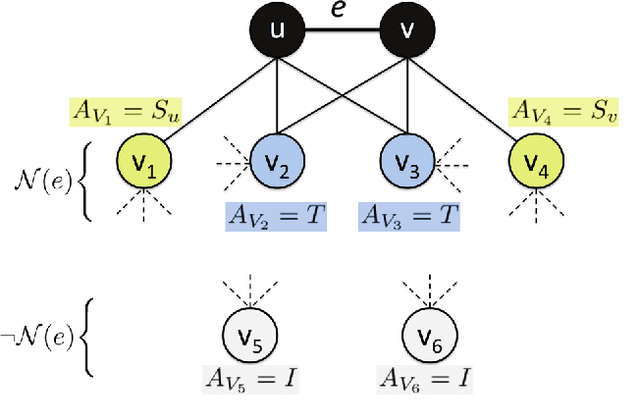
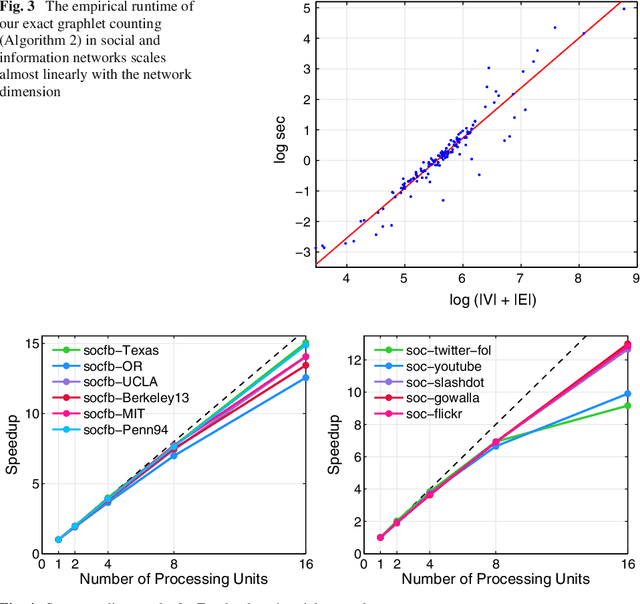
Abstract:From social science to biology, numerous applications often rely on graphlets for intuitive and meaningful characterization of networks at both the global macro-level as well as the local micro-level. While graphlets have witnessed a tremendous success and impact in a variety of domains, there has yet to be a fast and efficient approach for computing the frequencies of these subgraph patterns. However, existing methods are not scalable to large networks with millions of nodes and edges, which impedes the application of graphlets to new problems that require large-scale network analysis. To address these problems, we propose a fast, efficient, and parallel algorithm for counting graphlets of size k={3,4}-nodes that take only a fraction of the time to compute when compared with the current methods used. The proposed graphlet counting algorithms leverages a number of proven combinatorial arguments for different graphlets. For each edge, we count a few graphlets, and with these counts along with the combinatorial arguments, we obtain the exact counts of others in constant time. On a large collection of 300+ networks from a variety of domains, our graphlet counting strategies are on average 460x faster than current methods. This brings new opportunities to investigate the use of graphlets on much larger networks and newer applications as we show in the experiments. To the best of our knowledge, this paper provides the largest graphlet computations to date as well as the largest systematic investigation on over 300+ networks from a variety of domains.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge