Stephanie L. Hyland
ML4H Abstract Track 2020
Nov 19, 2020Abstract:A collection of the accepted abstracts for the Machine Learning for Health (ML4H) workshop at NeurIPS 2020. This index is not complete, as some accepted abstracts chose to opt-out of inclusion.
On the Intrinsic Privacy of Stochastic Gradient Descent
Dec 05, 2019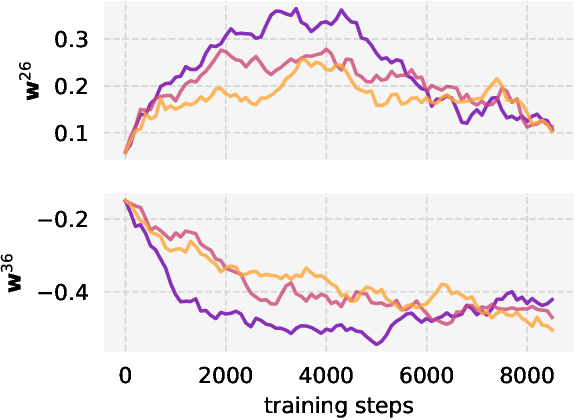
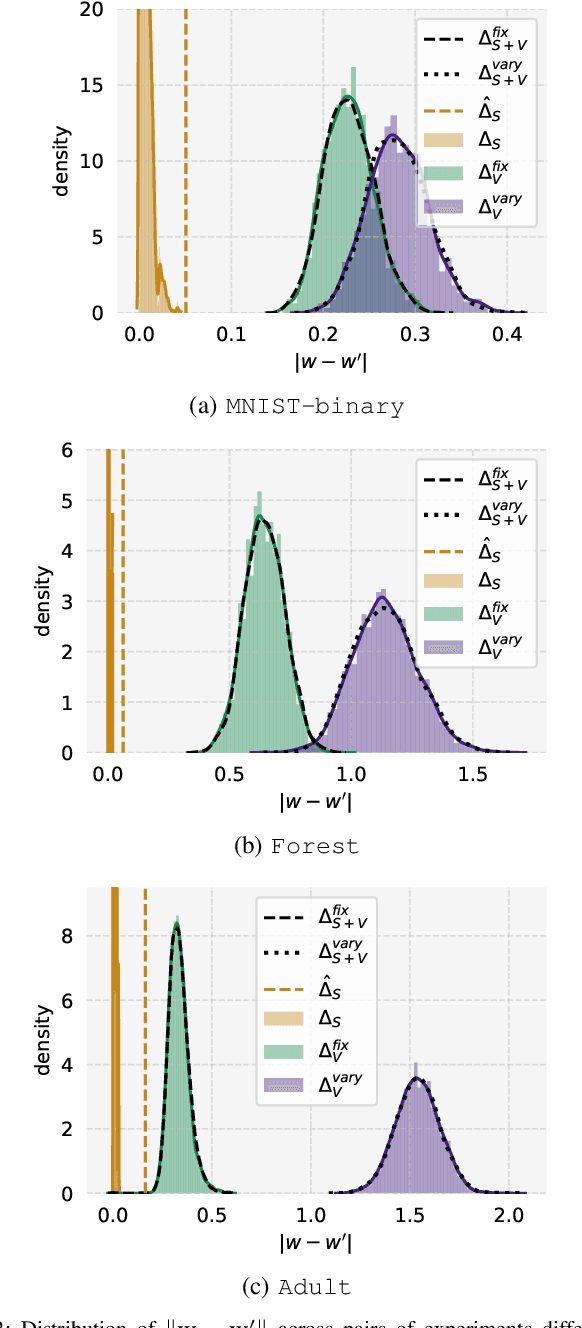
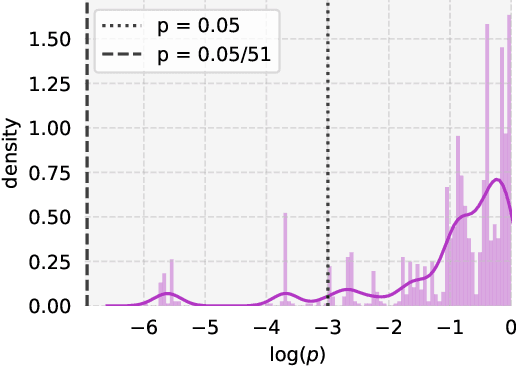
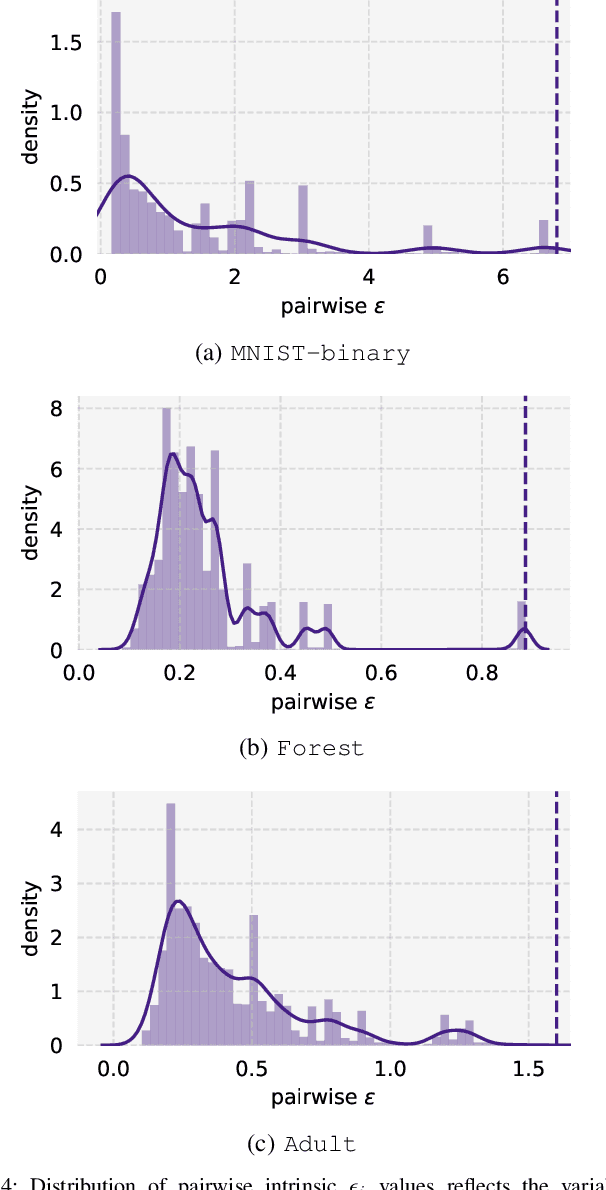
Abstract:Private learning algorithms have been proposed that ensure strong differential-privacy (DP) guarantees, however they often come at a cost to utility. Meanwhile, stochastic gradient descent (SGD) contains intrinsic randomness which has not been leveraged for privacy. In this work, we take the first step towards analysing the intrinsic privacy properties of SGD. Our primary contribution is a large-scale empirical analysis of SGD on convex and non-convex objectives. We evaluate the inherent variability due to the stochasticity in SGD on 3 datasets and calculate the $\epsilon$ values due to the intrinsic noise. First, we show that the variability in model parameters due to the random sampling almost always exceeds that due to changes in the data. We observe that SGD provides intrinsic $\epsilon$ values of 7.8, 6.9, and 2.8 on MNIST, Adult, and Forest Covertype datasets respectively. Next, we propose a method to augment the intrinsic noise of SGD to achieve the desired $\epsilon$. Our augmented SGD outputs models that outperform existing approaches with the same privacy guarantee, closing the gap to noiseless utility between 0.19% and 10.07%. Finally, we show that the existing theoretical bound on the sensitivity of SGD is not tight. By estimating the tightest bound empirically, we achieve near-noiseless performance at $\epsilon = 1$, closing the utility gap to the noiseless model between 3.13% and 100%. Our experiments provide concrete evidence that changing the seed in SGD is likely to have a far greater impact on the model than excluding any given training example. By accounting for this intrinsic randomness, higher utility can be achieved without sacrificing further privacy. With these results, we hope to inspire the research community to further explore and characterise the randomness in SGD, its impact on privacy, and the parallels with generalisation in machine learning.
Unsupervised Extraction of Phenotypes from Cancer Clinical Notes for Association Studies
May 03, 2019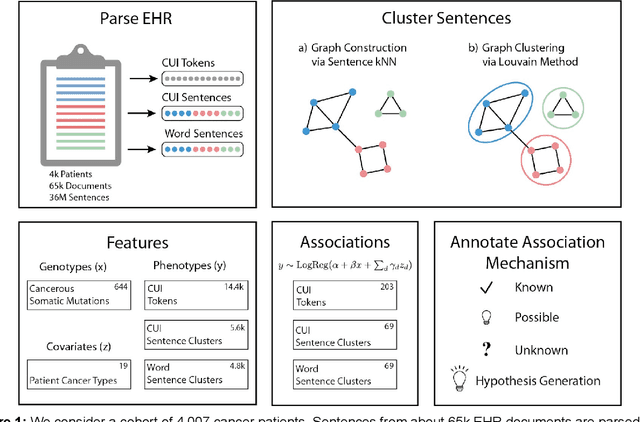
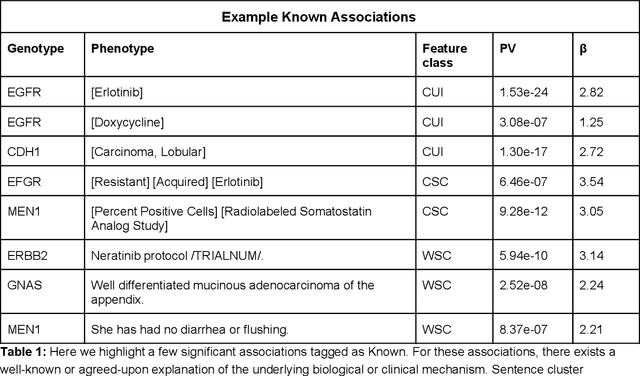
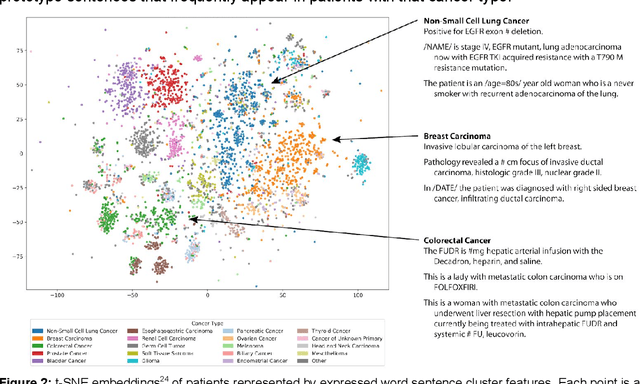
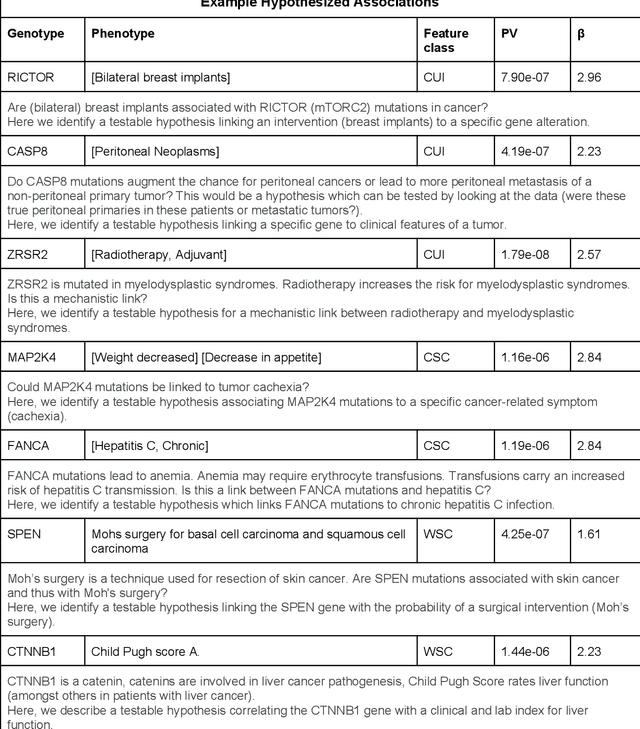
Abstract:The recent adoption of Electronic Health Records (EHRs) by health care providers has introduced an important source of data that provides detailed and highly specific insights into patient phenotypes over large cohorts. These datasets, in combination with machine learning and statistical approaches, generate new opportunities for research and clinical care. However, many methods require the patient representations to be in structured formats, while the information in the EHR is often locked in unstructured texts designed for human readability. In this work, we develop the methodology to automatically extract clinical features from clinical narratives from large EHR corpora without the need for prior knowledge. We consider medical terms and sentences appearing in clinical narratives as atomic information units. We propose an efficient clustering strategy suitable for the analysis of large text corpora and to utilize the clusters to represent information about the patient compactly. To demonstrate the utility of our approach, we perform an association study of clinical features with somatic mutation profiles from 4,007 cancer patients and their tumors. We apply the proposed algorithm to a dataset consisting of about 65 thousand documents with a total of about 3.2 million sentences. We identify 341 significant statistical associations between the presence of somatic mutations and clinical features. We annotated these associations according to their novelty, and report several known associations. We also propose 32 testable hypotheses where the underlying biological mechanism does not appear to be known but plausible. These results illustrate that the automated discovery of clinical features is possible and the joint analysis of clinical and genetic datasets can generate appealing new hypotheses.
Machine learning for early prediction of circulatory failure in the intensive care unit
Apr 19, 2019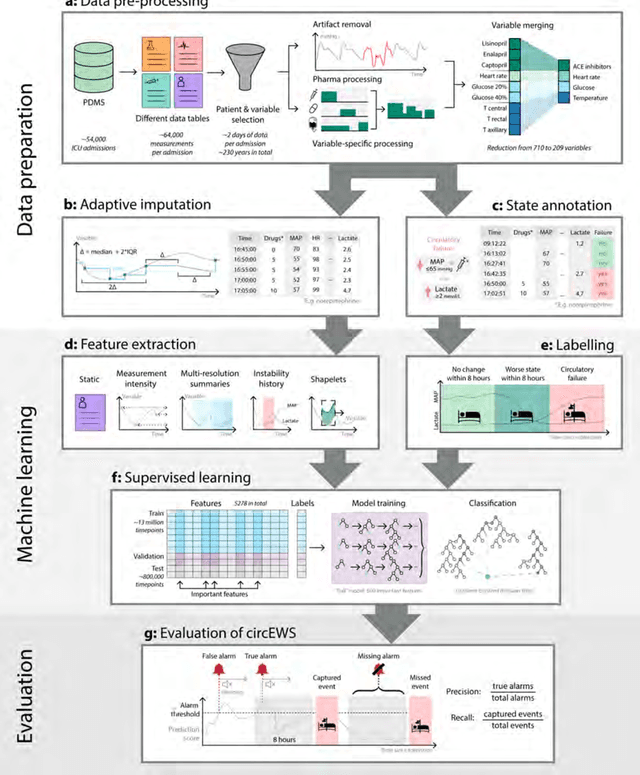
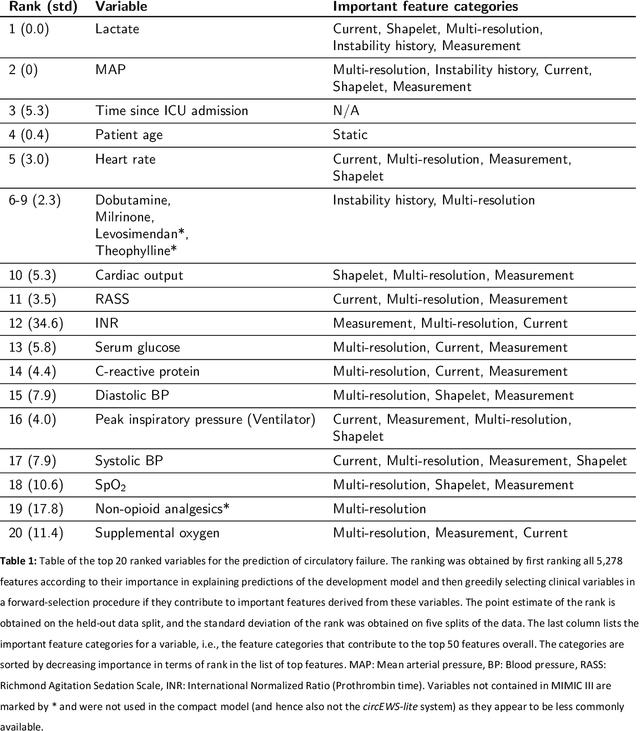
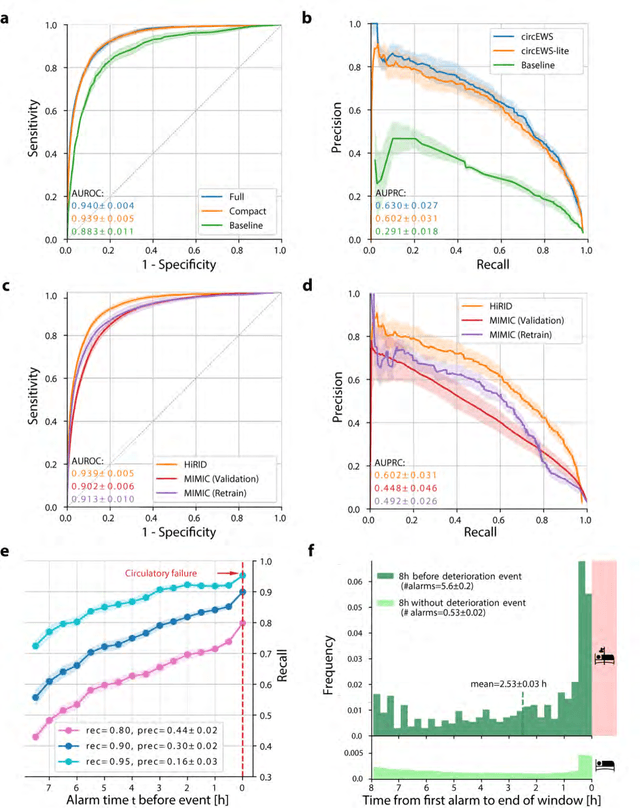
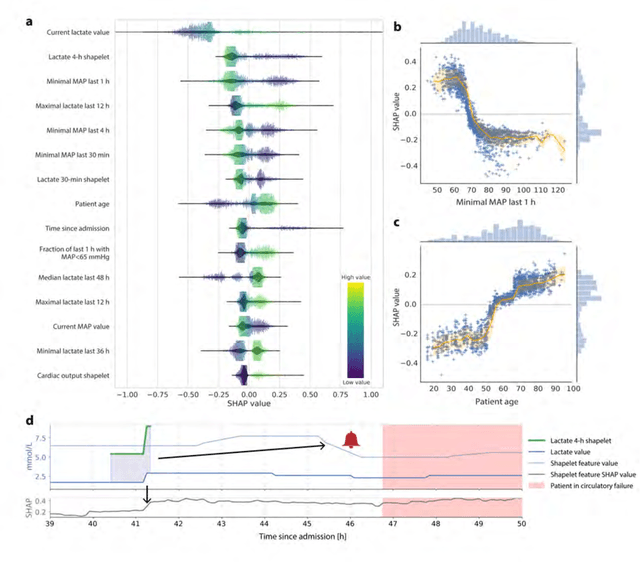
Abstract:Intensive care clinicians are presented with large quantities of patient information and measurements from a multitude of monitoring systems. The limited ability of humans to process such complex information hinders physicians to readily recognize and act on early signs of patient deterioration. We used machine learning to develop an early warning system for circulatory failure based on a high-resolution ICU database with 240 patient years of data. This automatic system predicts 90.0% of circulatory failure events (prevalence 3.1%), with 81.8% identified more than two hours in advance, resulting in an area under the receiver operating characteristic curve of 94.0% and area under the precision-recall curve of 63.0%. The model was externally validated in a large independent patient cohort.
Improving Clinical Predictions through Unsupervised Time Series Representation Learning
Dec 02, 2018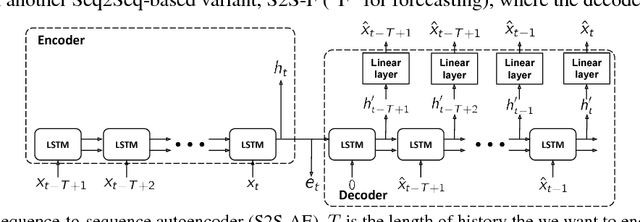
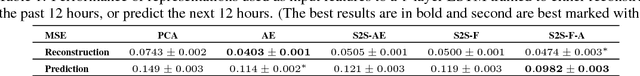
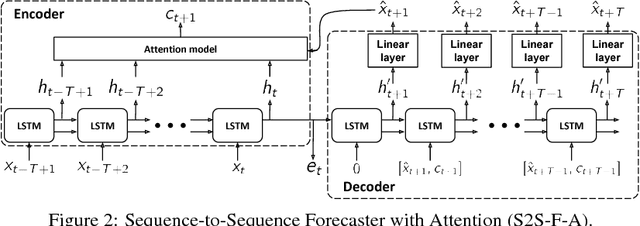
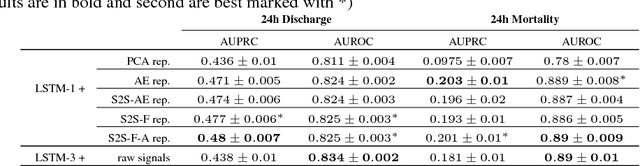
Abstract:In this work, we investigate unsupervised representation learning on medical time series, which bears the promise of leveraging copious amounts of existing unlabeled data in order to eventually assist clinical decision making. By evaluating on the prediction of clinically relevant outcomes, we show that in a practical setting, unsupervised representation learning can offer clear performance benefits over end-to-end supervised architectures. We experiment with using sequence-to-sequence (Seq2Seq) models in two different ways, as an autoencoder and as a forecaster, and show that the best performance is achieved by a forecasting Seq2Seq model with an integrated attention mechanism, proposed here for the first time in the setting of unsupervised learning for medical time series.
Real-valued (Medical) Time Series Generation with Recurrent Conditional GANs
Dec 04, 2017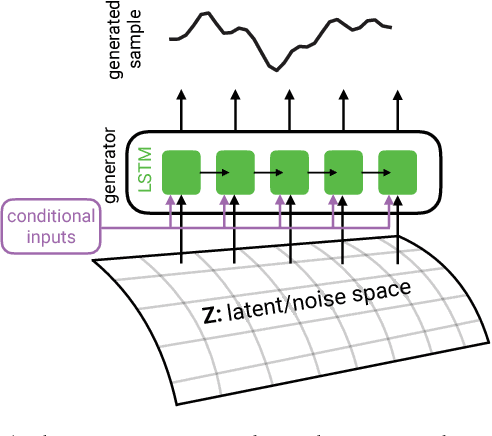
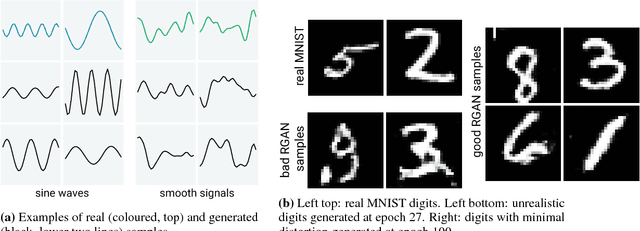
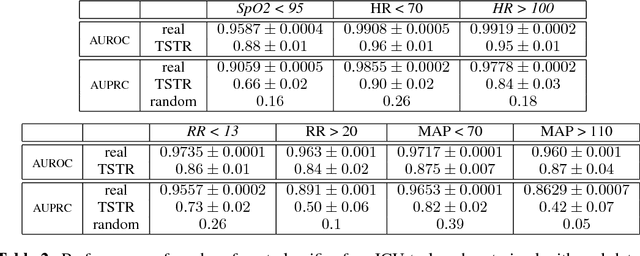
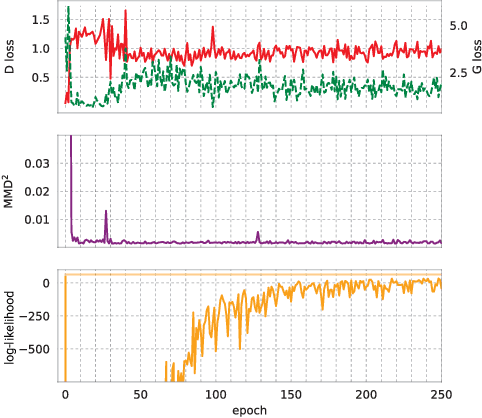
Abstract:Generative Adversarial Networks (GANs) have shown remarkable success as a framework for training models to produce realistic-looking data. In this work, we propose a Recurrent GAN (RGAN) and Recurrent Conditional GAN (RCGAN) to produce realistic real-valued multi-dimensional time series, with an emphasis on their application to medical data. RGANs make use of recurrent neural networks in the generator and the discriminator. In the case of RCGANs, both of these RNNs are conditioned on auxiliary information. We demonstrate our models in a set of toy datasets, where we show visually and quantitatively (using sample likelihood and maximum mean discrepancy) that they can successfully generate realistic time-series. We also describe novel evaluation methods for GANs, where we generate a synthetic labelled training dataset, and evaluate on a real test set the performance of a model trained on the synthetic data, and vice-versa. We illustrate with these metrics that RCGANs can generate time-series data useful for supervised training, with only minor degradation in performance on real test data. This is demonstrated on digit classification from 'serialised' MNIST and by training an early warning system on a medical dataset of 17,000 patients from an intensive care unit. We further discuss and analyse the privacy concerns that may arise when using RCGANs to generate realistic synthetic medical time series data.
Learning Unitary Operators with Help From u
Jan 10, 2017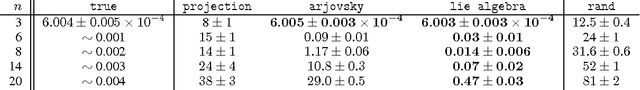
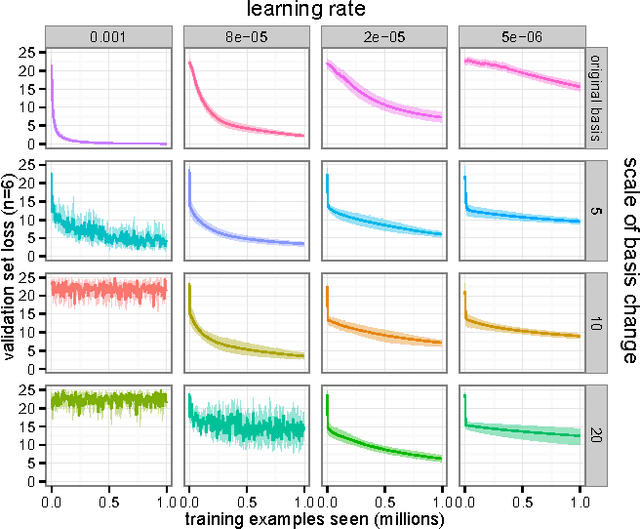

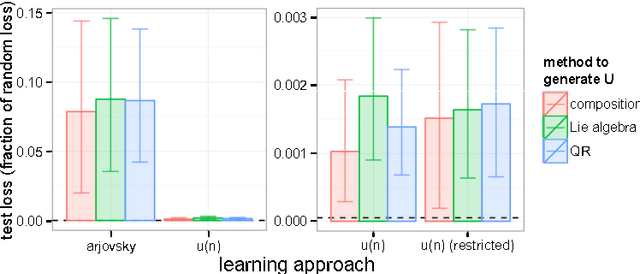
Abstract:A major challenge in the training of recurrent neural networks is the so-called vanishing or exploding gradient problem. The use of a norm-preserving transition operator can address this issue, but parametrization is challenging. In this work we focus on unitary operators and describe a parametrization using the Lie algebra $\mathfrak{u}(n)$ associated with the Lie group $U(n)$ of $n \times n$ unitary matrices. The exponential map provides a correspondence between these spaces, and allows us to define a unitary matrix using $n^2$ real coefficients relative to a basis of the Lie algebra. The parametrization is closed under additive updates of these coefficients, and thus provides a simple space in which to do gradient descent. We demonstrate the effectiveness of this parametrization on the problem of learning arbitrary unitary operators, comparing to several baselines and outperforming a recently-proposed lower-dimensional parametrization. We additionally use our parametrization to generalize a recently-proposed unitary recurrent neural network to arbitrary unitary matrices, using it to solve standard long-memory tasks.
Neural Document Embeddings for Intensive Care Patient Mortality Prediction
Dec 01, 2016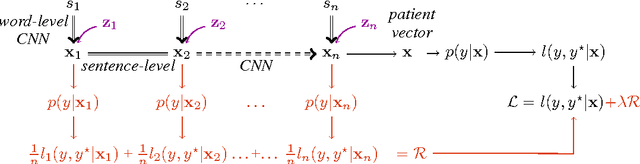
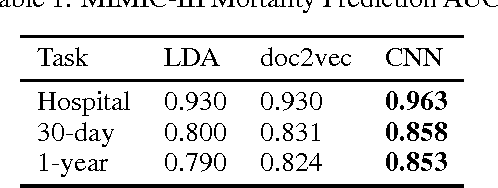
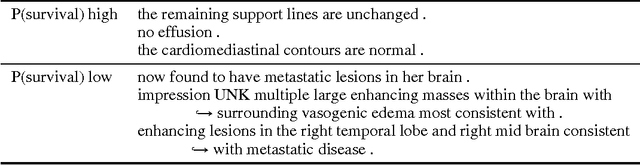



Abstract:We present an automatic mortality prediction scheme based on the unstructured textual content of clinical notes. Proposing a convolutional document embedding approach, our empirical investigation using the MIMIC-III intensive care database shows significant performance gains compared to previously employed methods such as latent topic distributions or generic doc2vec embeddings. These improvements are especially pronounced for the difficult problem of post-discharge mortality prediction.
Knowledge Transfer with Medical Language Embeddings
Feb 10, 2016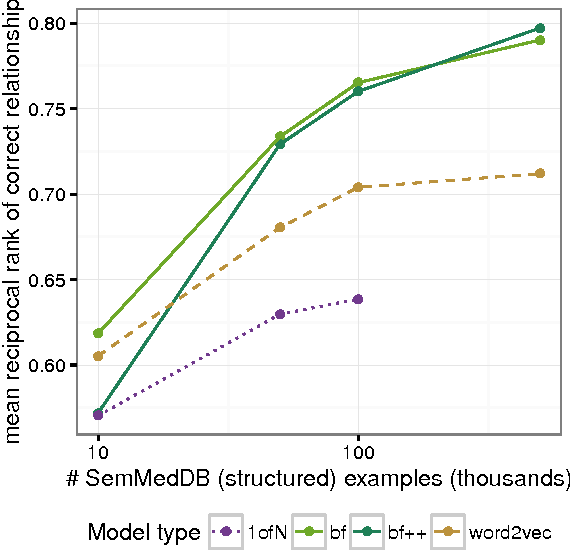
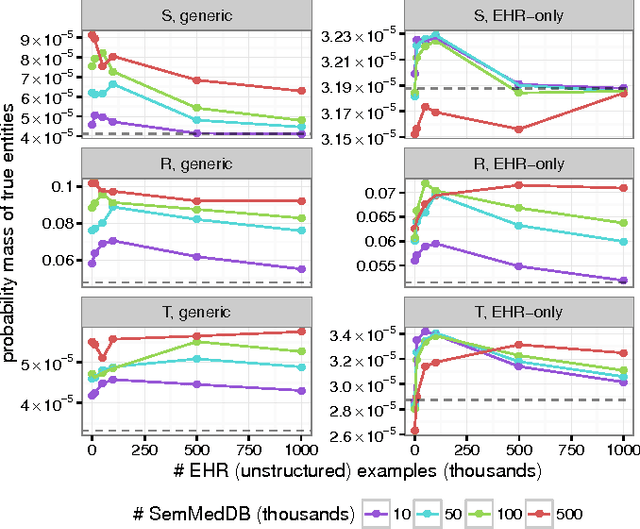
Abstract:Identifying relationships between concepts is a key aspect of scientific knowledge synthesis. Finding these links often requires a researcher to laboriously search through scien- tific papers and databases, as the size of these resources grows ever larger. In this paper we describe how distributional semantics can be used to unify structured knowledge graphs with unstructured text to predict new relationships between medical concepts, using a probabilistic generative model. Our approach is also designed to ameliorate data sparsity and scarcity issues in the medical domain, which make language modelling more challenging. Specifically, we integrate the medical relational database (SemMedDB) with text from electronic health records (EHRs) to perform knowledge graph completion. We further demonstrate the ability of our model to predict relationships between tokens not appearing in the relational database.
A Generative Model of Words and Relationships from Multiple Sources
Dec 03, 2015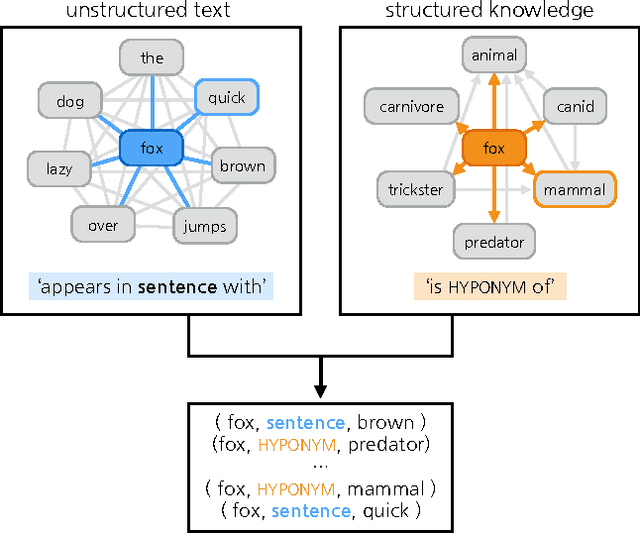
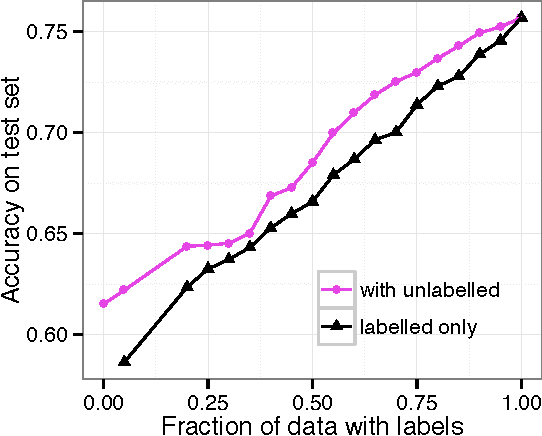
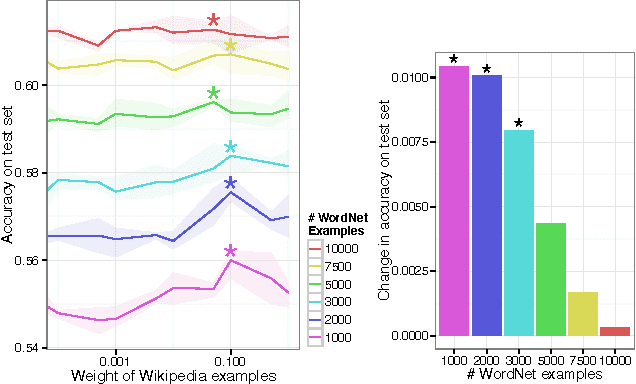
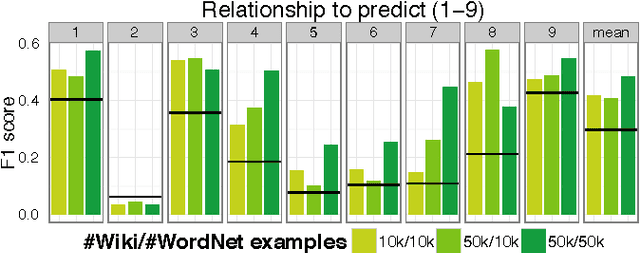
Abstract:Neural language models are a powerful tool to embed words into semantic vector spaces. However, learning such models generally relies on the availability of abundant and diverse training examples. In highly specialised domains this requirement may not be met due to difficulties in obtaining a large corpus, or the limited range of expression in average use. Such domains may encode prior knowledge about entities in a knowledge base or ontology. We propose a generative model which integrates evidence from diverse data sources, enabling the sharing of semantic information. We achieve this by generalising the concept of co-occurrence from distributional semantics to include other relationships between entities or words, which we model as affine transformations on the embedding space. We demonstrate the effectiveness of this approach by outperforming recent models on a link prediction task and demonstrating its ability to profit from partially or fully unobserved data training labels. We further demonstrate the usefulness of learning from different data sources with overlapping vocabularies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge