Songxiao Yang
RAM-H1200: A Unified Evaluation and Dataset on Hand Radiographs for Rheumatoid Arthritis
May 07, 2026Abstract:Rheumatoid arthritis (RA) assessment from hand radiographs requires multi-level analysis and modeling of anatomical structures and fine-grained local pathological changes. However, existing public resources do not support such unified multi-level analysis, often lacking full-hand coverage, fine-grained annotations, and consistent integration with clinical scoring systems. In particular, annotations that enable quantitative analysis of bone erosion (BE) remain scarce. RAM-H1200 contains 1,200 hand radiographs collected from six medical centers, with multi-level annotations including (i) whole-hand bone structure instance segmentation, (ii) pixel-level BE masks, (iii) SvdH-defined joint regions of interest, and (iv) joint-level SvdH scores for both BE and joint space narrowing (JSN). It is designed to evaluate whether models can jointly capture anatomical structure, localized erosive pathology, and clinically standardized RA severity from hand radiographs. The proposed BE masks enable, for the first time, quantitative BE analysis beyond coarse categorical grading by providing explicit spatial supervision for lesion extent and morphology. To our knowledge, RAM-H1200 is the first public large-scale benchmark that jointly supports whole-hand bone structure instance segmentation, pixel-level BE delineation, and clinically grounded joint-level SvdH scoring for both BE and JSN. Results across benchmark tasks show that anatomical modeling is substantially more mature than quantitative BE analysis: whole-hand bone segmentation achieves strong performance, whereas BE segmentation remains a major open challenge. By unifying anatomical structure modeling, quantitative lesion analysis, and clinically grounded SvdH scoring, RAM-H1200 provides a single benchmark for comprehensive RA analysis on hand radiographs.
PipeMFL-240K: A Large-scale Dataset and Benchmark for Object Detection in Pipeline Magnetic Flux Leakage Imaging
Feb 04, 2026Abstract:Pipeline integrity is critical to industrial safety and environmental protection, with Magnetic Flux Leakage (MFL) detection being a primary non-destructive testing technology. Despite the promise of deep learning for automating MFL interpretation, progress toward reliable models has been constrained by the absence of a large-scale public dataset and benchmark, making fair comparison and reproducible evaluation difficult. We introduce \textbf{PipeMFL-240K}, a large-scale, meticulously annotated dataset and benchmark for complex object detection in pipeline MFL pseudo-color images. PipeMFL-240K reflects real-world inspection complexity and poses several unique challenges: (i) an extremely long-tailed distribution over \textbf{12} categories, (ii) a high prevalence of tiny objects that often comprise only a handful of pixels, and (iii) substantial intra-class variability. The dataset contains \textbf{240,320} images and \textbf{191,530} high-quality bounding-box annotations, collected from 11 pipelines spanning approximately \textbf{1,480} km. Extensive experiments are conducted with state-of-the-art object detectors to establish baselines. Results show that modern detectors still struggle with the intrinsic properties of MFL data, highlighting considerable headroom for improvement, while PipeMFL-240K provides a reliable and challenging testbed to drive future research. As the first public dataset and the first benchmark of this scale and scope for pipeline MFL inspection, it provides a critical foundation for efficient pipeline diagnostics as well as maintenance planning and is expected to accelerate algorithmic innovation and reproducible research in MFL-based pipeline integrity assessment.
Efficient MedSAMs: Segment Anything in Medical Images on Laptop
Dec 20, 2024


Abstract:Promptable segmentation foundation models have emerged as a transformative approach to addressing the diverse needs in medical images, but most existing models require expensive computing, posing a big barrier to their adoption in clinical practice. In this work, we organized the first international competition dedicated to promptable medical image segmentation, featuring a large-scale dataset spanning nine common imaging modalities from over 20 different institutions. The top teams developed lightweight segmentation foundation models and implemented an efficient inference pipeline that substantially reduced computational requirements while maintaining state-of-the-art segmentation accuracy. Moreover, the post-challenge phase advanced the algorithms through the design of performance booster and reproducibility tasks, resulting in improved algorithms and validated reproducibility of the winning solution. Furthermore, the best-performing algorithms have been incorporated into the open-source software with a user-friendly interface to facilitate clinical adoption. The data and code are publicly available to foster the further development of medical image segmentation foundation models and pave the way for impactful real-world applications.
Fusing Medical Image Features and Clinical Features with Deep Learning for Computer-Aided Diagnosis
Mar 10, 2021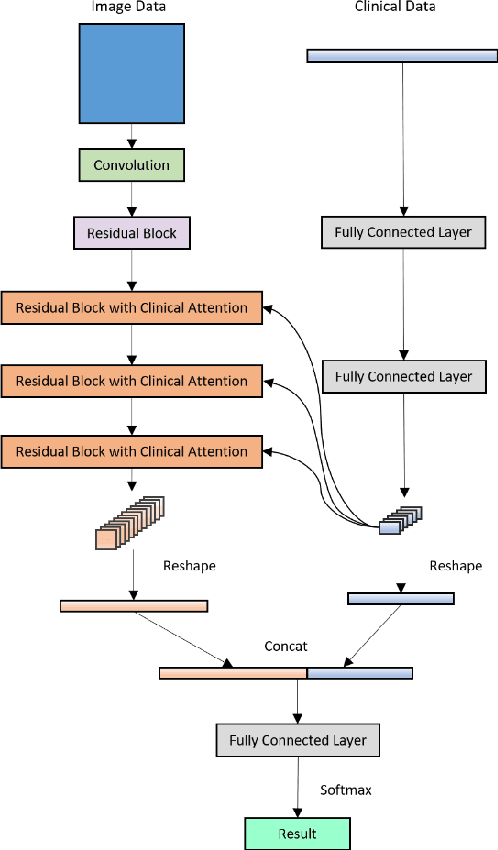
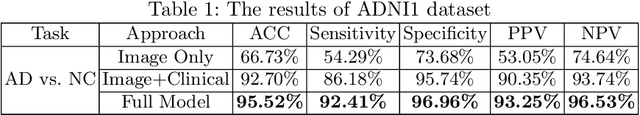
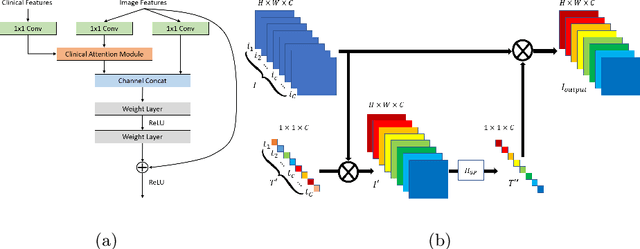
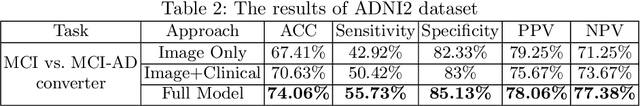
Abstract:Current Computer-Aided Diagnosis (CAD) methods mainly depend on medical images. The clinical information, which usually needs to be considered in practical clinical diagnosis, has not been fully employed in CAD. In this paper, we propose a novel deep learning-based method for fusing Magnetic Resonance Imaging (MRI)/Computed Tomography (CT) images and clinical information for diagnostic tasks. Two paths of neural layers are performed to extract image features and clinical features, respectively, and at the same time clinical features are employed as the attention to guide the extraction of image features. Finally, these two modalities of features are concatenated to make decisions. We evaluate the proposed method on its applications to Alzheimer's disease diagnosis, mild cognitive impairment converter prediction and hepatic microvascular invasion diagnosis. The encouraging experimental results prove the values of the image feature extraction guided by clinical features and the concatenation of two modalities of features for classification, which improve the performance of diagnosis effectively and stably.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge