Shujun Wang
ADAgent: LLM Agent for Alzheimer's Disease Analysis with Collaborative Coordinator
Jun 16, 2025Abstract:Alzheimer's disease (AD) is a progressive and irreversible neurodegenerative disease. Early and precise diagnosis of AD is crucial for timely intervention and treatment planning to alleviate the progressive neurodegeneration. However, most existing methods rely on single-modality data, which contrasts with the multifaceted approach used by medical experts. While some deep learning approaches process multi-modal data, they are limited to specific tasks with a small set of input modalities and cannot handle arbitrary combinations. This highlights the need for a system that can address diverse AD-related tasks, process multi-modal or missing input, and integrate multiple advanced methods for improved performance. In this paper, we propose ADAgent, the first specialized AI agent for AD analysis, built on a large language model (LLM) to address user queries and support decision-making. ADAgent integrates a reasoning engine, specialized medical tools, and a collaborative outcome coordinator to facilitate multi-modal diagnosis and prognosis tasks in AD. Extensive experiments demonstrate that ADAgent outperforms SOTA methods, achieving significant improvements in accuracy, including a 2.7% increase in multi-modal diagnosis, a 0.7% improvement in multi-modal prognosis, and enhancements in MRI and PET diagnosis tasks.
CorBenchX: Large-Scale Chest X-Ray Error Dataset and Vision-Language Model Benchmark for Report Error Correction
May 17, 2025Abstract:AI-driven models have shown great promise in detecting errors in radiology reports, yet the field lacks a unified benchmark for rigorous evaluation of error detection and further correction. To address this gap, we introduce CorBenchX, a comprehensive suite for automated error detection and correction in chest X-ray reports, designed to advance AI-assisted quality control in clinical practice. We first synthesize a large-scale dataset of 26,326 chest X-ray error reports by injecting clinically common errors via prompting DeepSeek-R1, with each corrupted report paired with its original text, error type, and human-readable description. Leveraging this dataset, we benchmark both open- and closed-source vision-language models,(e.g., InternVL, Qwen-VL, GPT-4o, o4-mini, and Claude-3.7) for error detection and correction under zero-shot prompting. Among these models, o4-mini achieves the best performance, with 50.6 % detection accuracy and correction scores of BLEU 0.853, ROUGE 0.924, BERTScore 0.981, SembScore 0.865, and CheXbertF1 0.954, remaining below clinical-level accuracy, highlighting the challenge of precise report correction. To advance the state of the art, we propose a multi-step reinforcement learning (MSRL) framework that optimizes a multi-objective reward combining format compliance, error-type accuracy, and BLEU similarity. We apply MSRL to QwenVL2.5-7B, the top open-source model in our benchmark, achieving an improvement of 38.3% in single-error detection precision and 5.2% in single-error correction over the zero-shot baseline.
Predicting Diabetic Macular Edema Treatment Responses Using OCT: Dataset and Methods of APTOS Competition
May 09, 2025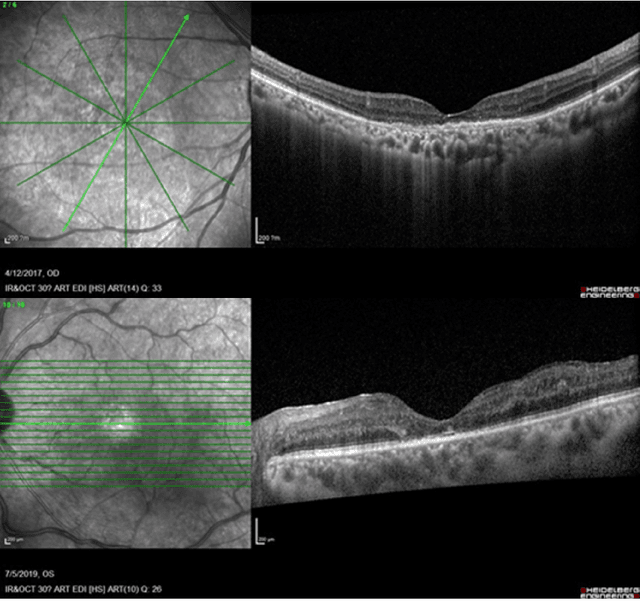
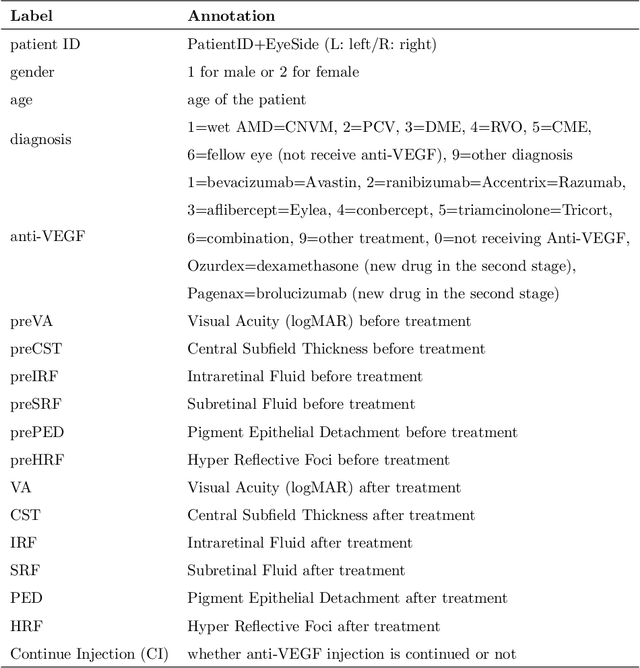
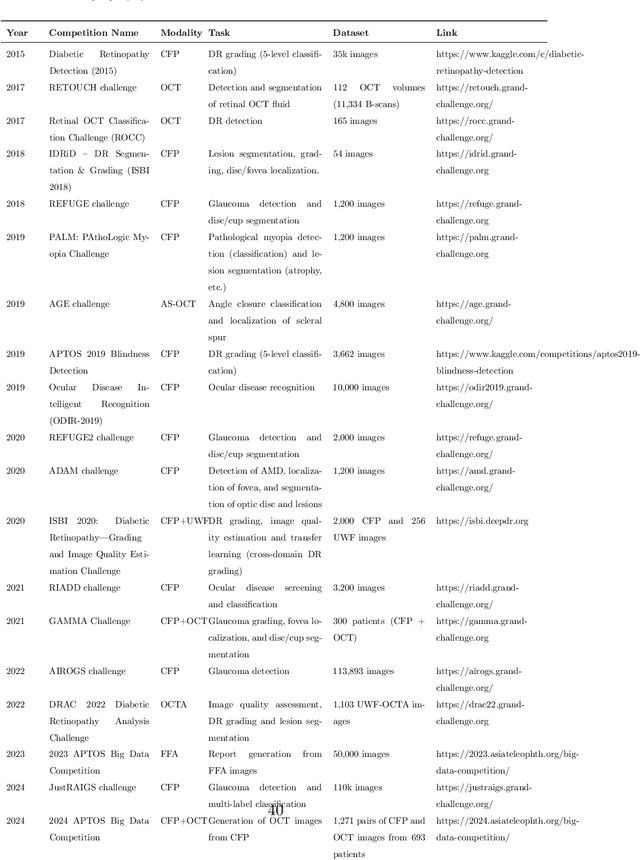
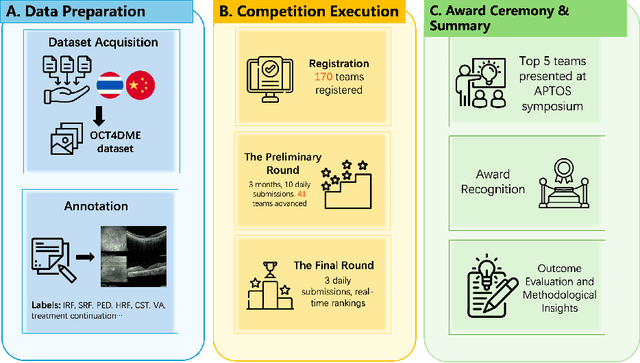

Abstract:Diabetic macular edema (DME) significantly contributes to visual impairment in diabetic patients. Treatment responses to intravitreal therapies vary, highlighting the need for patient stratification to predict therapeutic benefits and enable personalized strategies. To our knowledge, this study is the first to explore pre-treatment stratification for predicting DME treatment responses. To advance this research, we organized the 2nd Asia-Pacific Tele-Ophthalmology Society (APTOS) Big Data Competition in 2021. The competition focused on improving predictive accuracy for anti-VEGF therapy responses using ophthalmic OCT images. We provided a dataset containing tens of thousands of OCT images from 2,000 patients with labels across four sub-tasks. This paper details the competition's structure, dataset, leading methods, and evaluation metrics. The competition attracted strong scientific community participation, with 170 teams initially registering and 41 reaching the final round. The top-performing team achieved an AUC of 80.06%, highlighting the potential of AI in personalized DME treatment and clinical decision-making.
AOR: Anatomical Ontology-Guided Reasoning for Medical Large Multimodal Model in Chest X-Ray Interpretation
May 05, 2025Abstract:Chest X-rays (CXRs) are the most frequently performed imaging examinations in clinical settings. Recent advancements in Large Multimodal Models (LMMs) have enabled automated CXR interpretation, enhancing diagnostic accuracy and efficiency. However, despite their strong visual understanding, current Medical LMMs (MLMMs) still face two major challenges: (1) Insufficient region-level understanding and interaction, and (2) Limited accuracy and interpretability due to single-step reasoning. In this paper, we empower MLMMs with anatomy-centric reasoning capabilities to enhance their interactivity and explainability. Specifically, we first propose an Anatomical Ontology-Guided Reasoning (AOR) framework, which centers on cross-modal region-level information to facilitate multi-step reasoning. Next, under the guidance of expert physicians, we develop AOR-Instruction, a large instruction dataset for MLMMs training. Our experiments demonstrate AOR's superior performance in both VQA and report generation tasks.
Synchronized Video-to-Audio Generation via Mel Quantization-Continuum Decomposition
Mar 10, 2025Abstract:Video-to-audio generation is essential for synthesizing realistic audio tracks that synchronize effectively with silent videos. Following the perspective of extracting essential signals from videos that can precisely control the mature text-to-audio generative diffusion models, this paper presents how to balance the representation of mel-spectrograms in terms of completeness and complexity through a new approach called Mel Quantization-Continuum Decomposition (Mel-QCD). We decompose the mel-spectrogram into three distinct types of signals, employing quantization or continuity to them, we can effectively predict them from video by a devised video-to-all (V2X) predictor. Then, the predicted signals are recomposed and fed into a ControlNet, along with a textual inversion design, to control the audio generation process. Our proposed Mel-QCD method demonstrates state-of-the-art performance across eight metrics, evaluating dimensions such as quality, synchronization, and semantic consistency. Our codes and demos will be released at \href{Website}{https://wjc2830.github.io/MelQCD/}.
ReFocus: Reinforcing Mid-Frequency and Key-Frequency Modeling for Multivariate Time Series Forecasting
Feb 24, 2025



Abstract:Recent advancements have progressively incorporated frequency-based techniques into deep learning models, leading to notable improvements in accuracy and efficiency for time series analysis tasks. However, the Mid-Frequency Spectrum Gap in the real-world time series, where the energy is concentrated at the low-frequency region while the middle-frequency band is negligible, hinders the ability of existing deep learning models to extract the crucial frequency information. Additionally, the shared Key-Frequency in multivariate time series, where different time series share indistinguishable frequency patterns, is rarely exploited by existing literature. This work introduces a novel module, Adaptive Mid-Frequency Energy Optimizer, based on convolution and residual learning, to emphasize the significance of mid-frequency bands. We also propose an Energy-based Key-Frequency Picking Block to capture shared Key-Frequency, which achieves superior inter-series modeling performance with fewer parameters. A novel Key-Frequency Enhanced Training strategy is employed to further enhance Key-Frequency modeling, where spectral information from other channels is randomly introduced into each channel. Our approach advanced multivariate time series forecasting on the challenging Traffic, ECL, and Solar benchmarks, reducing MSE by 4%, 6%, and 5% compared to the previous SOTA iTransformer. Code is available at this GitHub Repository: https://github.com/Levi-Ackman/ReFocus.
LiNo: Advancing Recursive Residual Decomposition of Linear and Nonlinear Patterns for Robust Time Series Forecasting
Oct 22, 2024



Abstract:Forecasting models are pivotal in a data-driven world with vast volumes of time series data that appear as a compound of vast Linear and Nonlinear patterns. Recent deep time series forecasting models struggle to utilize seasonal and trend decomposition to separate the entangled components. Such a strategy only explicitly extracts simple linear patterns like trends, leaving the other linear modes and vast unexplored nonlinear patterns to the residual. Their flawed linear and nonlinear feature extraction models and shallow-level decomposition limit their adaptation to the diverse patterns present in real-world scenarios. Given this, we innovate Recursive Residual Decomposition by introducing explicit extraction of both linear and nonlinear patterns. This deeper-level decomposition framework, which is named LiNo, captures linear patterns using a Li block which can be a moving average kernel, and models nonlinear patterns using a No block which can be a Transformer encoder. The extraction of these two patterns is performed alternatively and recursively. To achieve the full potential of LiNo, we develop the current simple linear pattern extractor to a general learnable autoregressive model, and design a novel No block that can handle all essential nonlinear patterns. Remarkably, the proposed LiNo achieves state-of-the-art on thirteen real-world benchmarks under univariate and multivariate forecasting scenarios. Experiments show that current forecasting models can deliver more robust and precise results through this advanced Recursive Residual Decomposition. We hope this work could offer insight into designing more effective forecasting models. Code is available at this Repository: https://github.com/Levi-Ackman/LiNo.
MMR-Mamba: Multi-Modal MRI Reconstruction with Mamba and Spatial-Frequency Information Fusion
Jul 07, 2024Abstract:Multi-modal MRI offers valuable complementary information for diagnosis and treatment; however, its utility is limited by prolonged scanning times. To accelerate the acquisition process, a practical approach is to reconstruct images of the target modality, which requires longer scanning times, from under-sampled k-space data using the fully-sampled reference modality with shorter scanning times as guidance. The primary challenge of this task is comprehensively and efficiently integrating complementary information from different modalities to achieve high-quality reconstruction. Existing methods struggle with this: 1) convolution-based models fail to capture long-range dependencies; 2) transformer-based models, while excelling in global feature modeling, struggle with quadratic computational complexity. To address this, we propose MMR-Mamba, a novel framework that thoroughly and efficiently integrates multi-modal features for MRI reconstruction, leveraging Mamba's capability to capture long-range dependencies with linear computational complexity while exploiting global properties of the Fourier domain. Specifically, we first design a Target modality-guided Cross Mamba (TCM) module in the spatial domain, which maximally restores the target modality information by selectively incorporating relevant information from the reference modality. Then, we introduce a Selective Frequency Fusion (SFF) module to efficiently integrate global information in the Fourier domain and recover high-frequency signals for the reconstruction of structural details. Furthermore, we devise an Adaptive Spatial-Frequency Fusion (ASFF) module, which mutually enhances the spatial and frequency domains by supplementing less informative channels from one domain with corresponding channels from the other.
MMR-Mamba: Multi-Contrast MRI Reconstruction with Mamba and Spatial-Frequency Information Fusion
Jun 27, 2024Abstract:Multi-contrast MRI acceleration has become prevalent in MR imaging, enabling the reconstruction of high-quality MR images from under-sampled k-space data of the target modality, using guidance from a fully-sampled auxiliary modality. The main crux lies in efficiently and comprehensively integrating complementary information from the auxiliary modality. Existing methods either suffer from quadratic computational complexity or fail to capture long-range correlated features comprehensively. In this work, we propose MMR-Mamba, a novel framework that achieves comprehensive integration of multi-contrast features through Mamba and spatial-frequency information fusion. Firstly, we design the \textit{Target modality-guided Cross Mamba} (TCM) module in the spatial domain, which maximally restores the target modality information by selectively absorbing useful information from the auxiliary modality. Secondly, leveraging global properties of the Fourier domain, we introduce the \textit{Selective Frequency Fusion} (SFF) module to efficiently integrate global information in the frequency domain and recover high-frequency signals for the reconstruction of structure details. Additionally, we present the \textit{Adaptive Spatial-Frequency Fusion} (ASFF) module, which enhances fused features by supplementing less informative features from one domain with corresponding features from the other domain. These innovative strategies ensure efficient feature fusion across spatial and frequency domains, avoiding the introduction of redundant information and facilitating the reconstruction of high-quality target images. Extensive experiments on the BraTS and fastMRI knee datasets demonstrate the superiority of the proposed MMR-Mamba over state-of-the-art MRI reconstruction methods.
CMViM: Contrastive Masked Vim Autoencoder for 3D Multi-modal Representation Learning for AD classification
Mar 25, 2024Abstract:Alzheimer's disease (AD) is an incurable neurodegenerative condition leading to cognitive and functional deterioration. Given the lack of a cure, prompt and precise AD diagnosis is vital, a complex process dependent on multiple factors and multi-modal data. While successful efforts have been made to integrate multi-modal representation learning into medical datasets, scant attention has been given to 3D medical images. In this paper, we propose Contrastive Masked Vim Autoencoder (CMViM), the first efficient representation learning method tailored for 3D multi-modal data. Our proposed framework is built on a masked Vim autoencoder to learn a unified multi-modal representation and long-dependencies contained in 3D medical images. We also introduce an intra-modal contrastive learning module to enhance the capability of the multi-modal Vim encoder for modeling the discriminative features in the same modality, and an inter-modal contrastive learning module to alleviate misaligned representation among modalities. Our framework consists of two main steps: 1) incorporate the Vision Mamba (Vim) into the mask autoencoder to reconstruct 3D masked multi-modal data efficiently. 2) align the multi-modal representations with contrastive learning mechanisms from both intra-modal and inter-modal aspects. Our framework is pre-trained and validated ADNI2 dataset and validated on the downstream task for AD classification. The proposed CMViM yields 2.7\% AUC performance improvement compared with other state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge