Sergios Theodoridis
Robert
Early soft and flexible fusion of EEG and fMRI via tensor decompositions
May 12, 2020
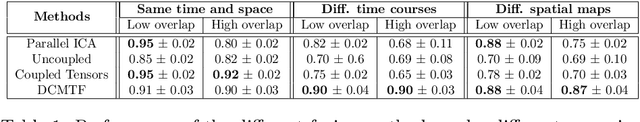
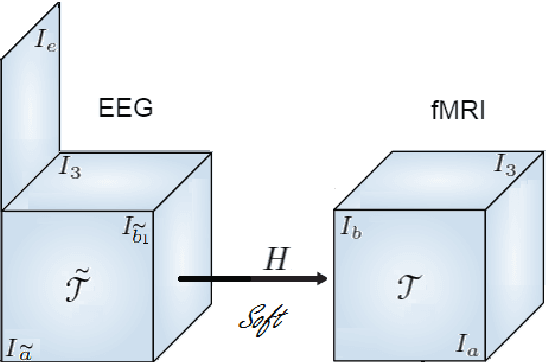
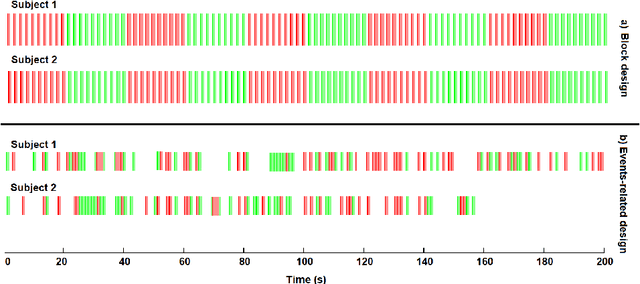
Abstract:Data fusion refers to the joint analysis of multiple datasets which provide complementary views of the same task. In this preprint, the problem of jointly analyzing electroencephalography (EEG) and functional Magnetic Resonance Imaging (fMRI) data is considered. Jointly analyzing EEG and fMRI measurements is highly beneficial for studying brain function because these modalities have complementary spatiotemporal resolution: EEG offers good temporal resolution while fMRI is better in its spatial resolution. The fusion methods reported so far ignore the underlying multi-way nature of the data in at least one of the modalities and/or rely on very strong assumptions about the relation of the two datasets. In this preprint, these two points are addressed by adopting for the first time tensor models in the two modalities while also exploring double coupled tensor decompositions and by following soft and flexible coupling approaches to implement the multi-modal analysis. To cope with the Event Related Potential (ERP) variability in EEG, the PARAFAC2 model is adopted. The results obtained are compared against those of parallel Independent Component Analysis (ICA) and hard coupling alternatives in both simulated and real data. Our results confirm the superiority of tensorial methods over methods based on ICA. In scenarios that do not meet the assumptions underlying hard coupling, the advantage of soft and flexible coupled decompositions is clearly demonstrated.
FedLoc: Federated Learning Framework for Cooperative Localization and Location Data Processing
Mar 08, 2020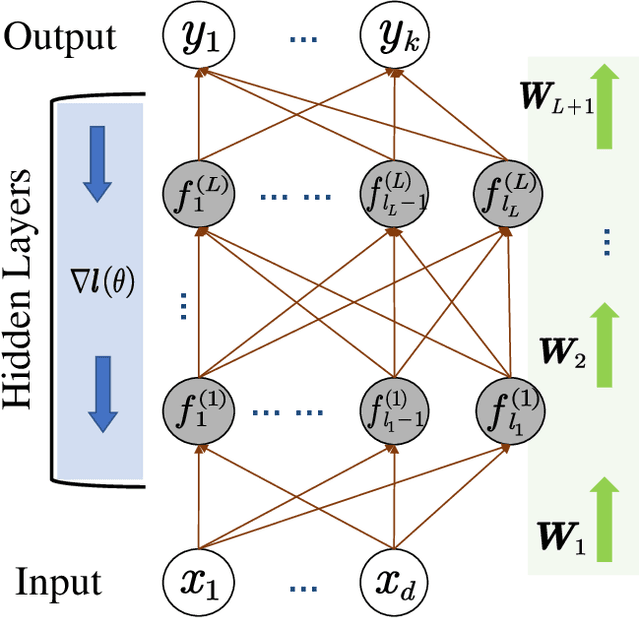
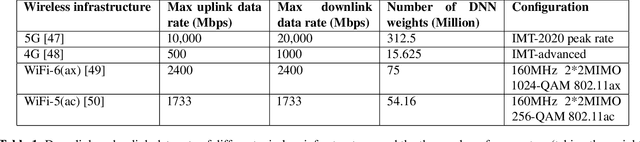

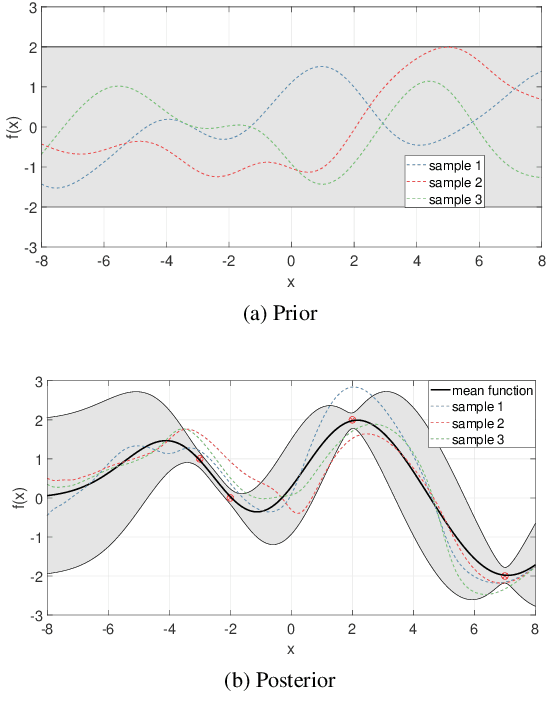
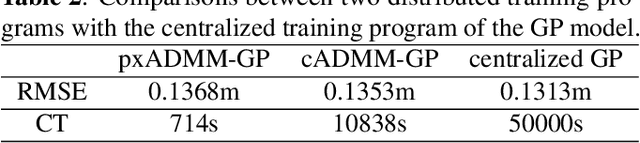
Abstract:In this paper, we propose a new localization framework in which mobile users or smart agents can cooperate to build accurate location services without sacrificing privacy, in particular, information related to their trajectories. The proposed framework is called Federated Localization (FedLoc), simply because it adopts the recently proposed federated learning. Apart from the new FedLoc framework, this paper can be deemed as an overview paper, in which we review the state-of-the-art federated learning framework, two widely used learning models, various distributed model hyper-parameter optimization schemes, and some practical use cases that fall under the FedLoc framework. The use cases, summarized from a mixture of standard, recently published, and unpublished works, cover a broad range of location services, including collaborative static localization/fingerprinting, indoor target tracking, outdoor navigation using low-sampling GPS, and spatio-temporal wireless traffic data modeling and prediction. The obtained primary results confirm that the proposed FedLoc framework well suits data-driven, machine learning-based localization and spatio-temporal data modeling. Future research directions are discussed at the end of this paper.
Variational Conditional-Dependence Hidden Markov Models for Human Action Recognition
Feb 13, 2020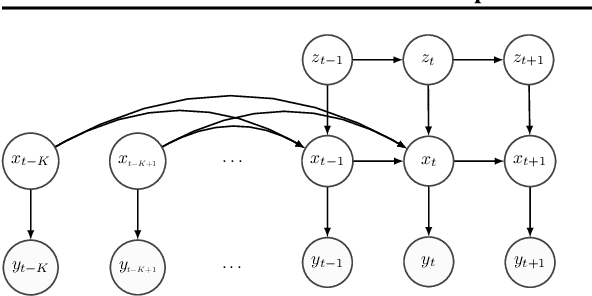
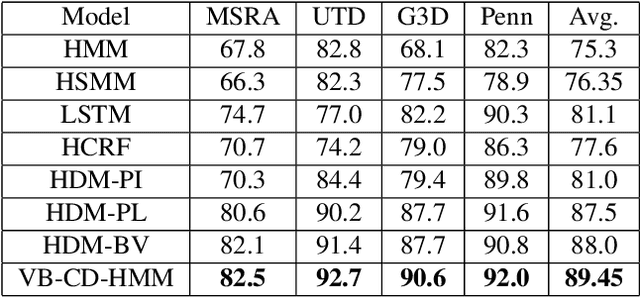
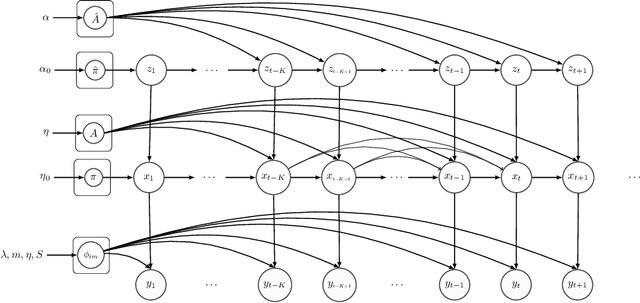
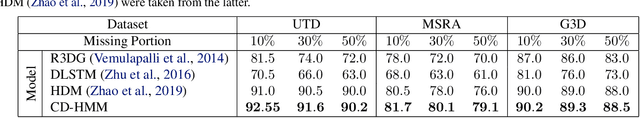
Abstract:Hidden Markov Models (HMMs) are a powerful generative approach for modeling sequential data and time-series in general. However, the commonly employed assumption of the dependence of the current time frame to a single or multiple immediately preceding frames is unrealistic; more complicated dynamics potentially exist in real world scenarios. Human Action Recognition constitutes such a scenario, and has attracted increased attention with the advent of low-cost 3D sensors. The naturally arising variations and complex temporal dependencies have established this task as a challenging problem in the community. This paper revisits conventional sequential modeling approaches, aiming to address the problem of capturing time-varying temporal dependency patterns. To this end, we propose a different formulation of HMMs, whereby the dependence on past frames is dynamically inferred from the data. Specifically, we introduce a hierarchical extension by postulating an additional latent variable layer; therein, the (time-varying) temporal dependence patterns are treated as latent variables over which inference is performed. We leverage solid arguments from the Variational Bayes framework and derive a tractable inference algorithm based on the forward-backward algorithm. As we experimentally show using benchmark datasets, our approach yields competitive recognition accuracy and can effectively handle data with missing values.
Linear Multiple Low-Rank Kernel Based Stationary Gaussian Processes Regression for Time Series
Apr 21, 2019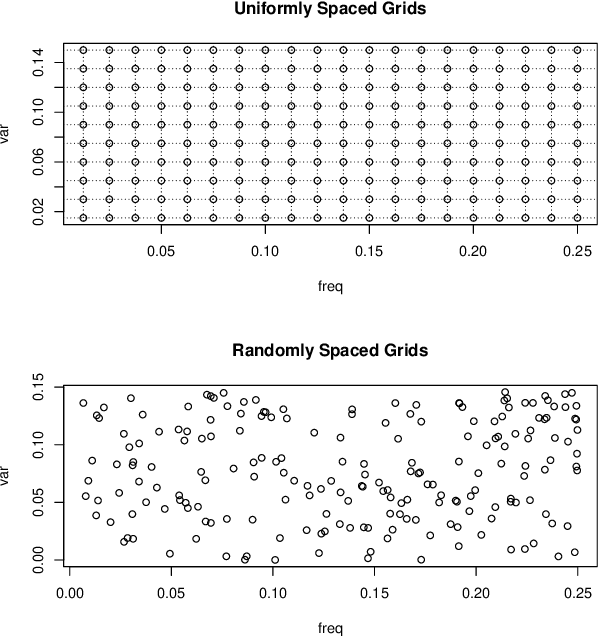
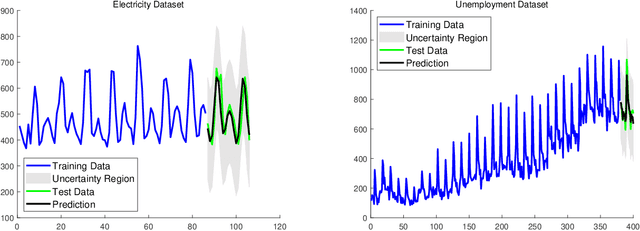
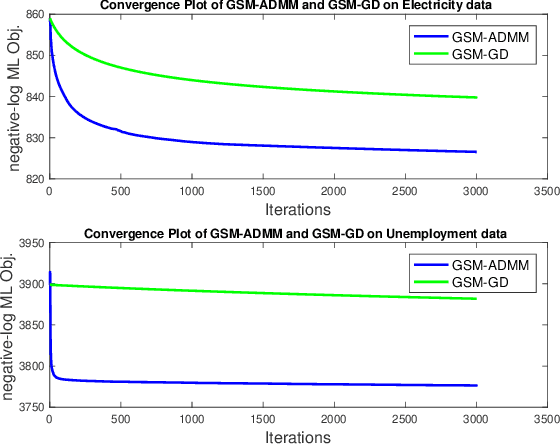
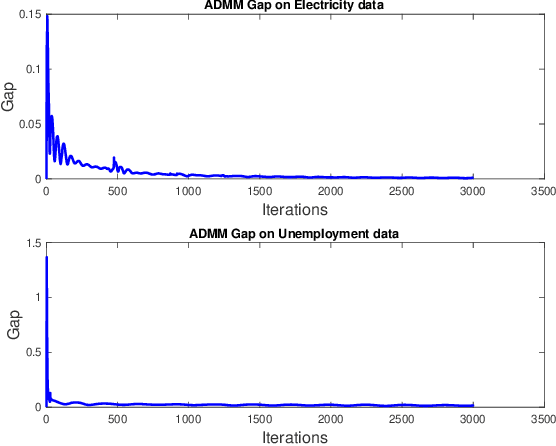
Abstract:Gaussian processes (GP) for machine learning have been studied systematically over the past two decades and they are by now widely used in a number of diverse applications. However, GP kernel design and the associated hyper-parameter optimization are still hard and to a large extend open problems. In this paper, we consider the task of GP regression for time series modeling and analysis. The underlying stationary kernel can be approximated arbitrarily close by a new proposed grid spectral mixture (GSM) kernel, which turns out to be a linear combination of low-rank sub-kernels. In the case where a large number of the sub-kernels are used, either the Nystr\"{o}m or the random Fourier feature approximations can be adopted to deal efficiently with the computational demands. The unknown GP hyper-parameters consist of the non-negative weights of all sub-kernels as well as the noise variance; their estimation is performed via the maximum-likelihood (ML) estimation framework. Two efficient numerical optimization methods for solving the unknown hyper-parameters are derived, including a sequential majorization-minimization (MM) method and a non-linearly constrained alternating direction of multiplier method (ADMM). The MM matches perfectly with the proven low-rank property of the proposed GSM sub-kernels and turns out to be a part of efficiency, stable, and efficient solver, while the ADMM has the potential to generate better local minimum in terms of the test MSE. Experimental results, based on various classic time series data sets, corroborate that the proposed GSM kernel-based GP regression model outperforms several salient competitors of similar kind in terms of prediction mean-squared-error and numerical stability.
Nonparametric Bayesian Deep Networks with Local Competition
May 19, 2018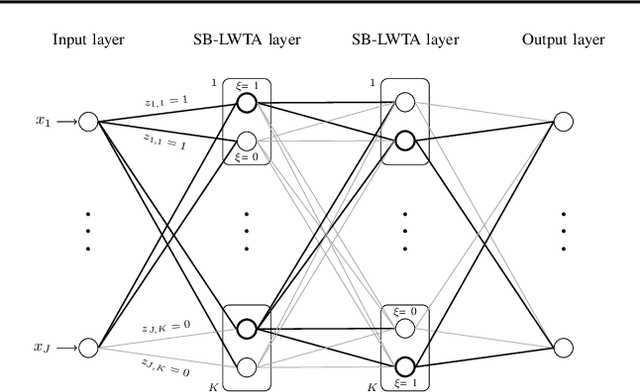
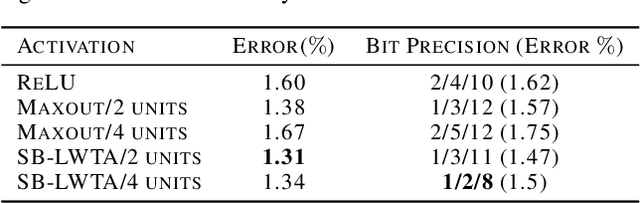

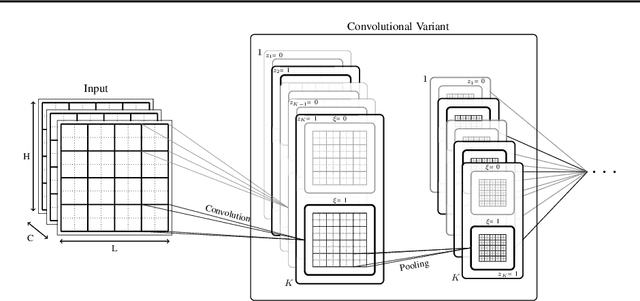
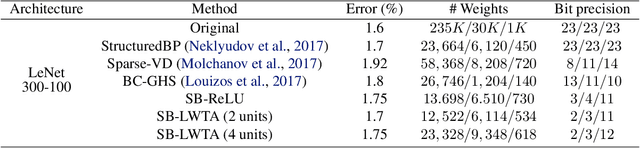
Abstract:Local competition among neighboring neurons is a common procedure taking place in biological systems. This finding has inspired research on more biologically plausible deep networks that comprise competing linear units; such models can be effectively trained by means of gradient-based backpropagation. This is in contrast to traditional deep networks, built of nonlinear units that do not entail any form of (local) competition. However, for the case of competition-based networks, the problem of data-driven inference of their most appropriate configuration, including the needed number of connections or locally competing sets of units, has not been touched upon by the research community. This work constitutes the first attempt to address this shortcoming; to this end, we leverage solid arguments from the field of Bayesian nonparametrics. Specifically, we introduce auxiliary discrete latent variables of model component utility, and perform Bayesian inference over them. We impose appropriate stick-breaking priors over the introduced discrete latent variables; these give rise to an well-established sparsity-inducing mechanism. We devise efficient inference algorithms for our model by resorting to stochastic gradient variational Bayes. We perform an extensive experimental evaluation of our approach using benchmark data. Our results verify that we obtain state-of-the-art accuracy albeit via networks of much smaller memory and computational footprint than the competition.
Information Assisted Dictionary Learning for fMRI data analysis
May 11, 2018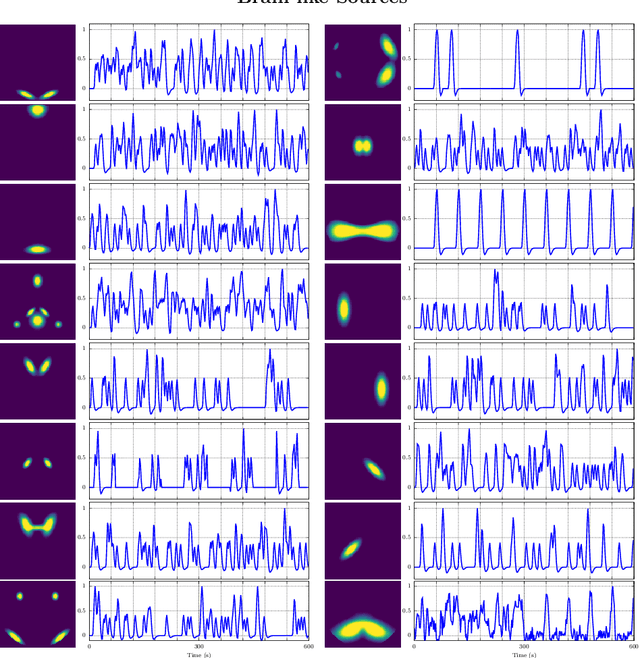
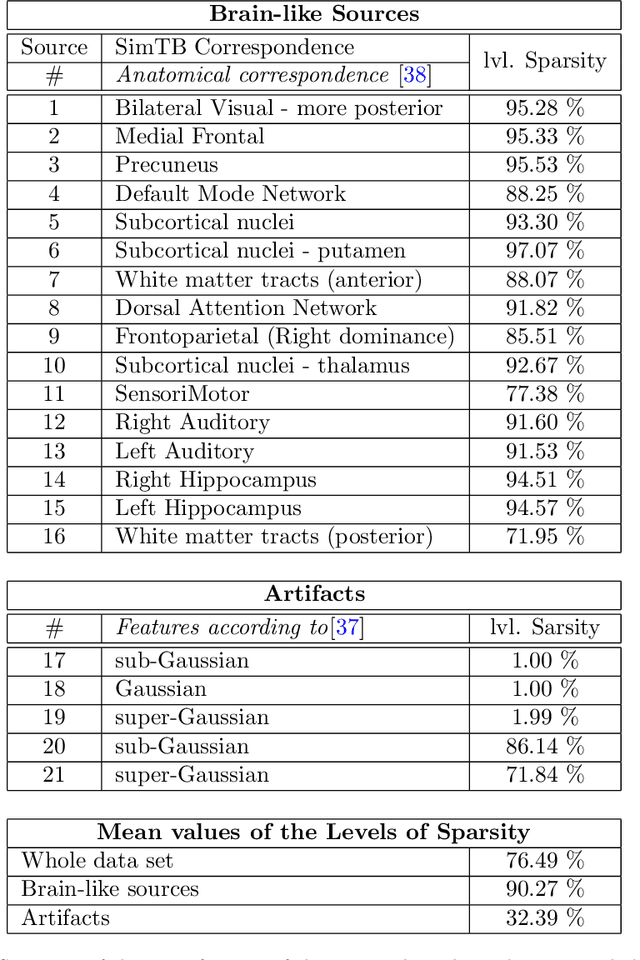
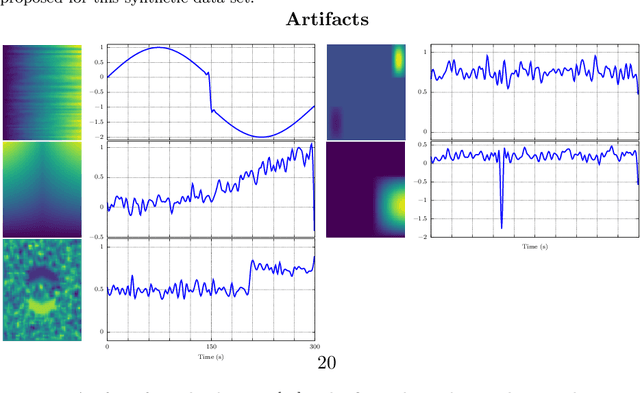

Abstract:In this paper, the task-related fMRI problem is treated in its matrix factorization formulation. The focus of the reported work is on the dictionary learning (DL) matrix factorization approach. A major novelty of the paper lies in the incorporation of well-established assumptions associated with the GLM technique, which is currently in use by the neuroscientists. These assumptions are embedded as constraints in the DL formulation. In this way, our approach provides a framework of combining well-established and understood techniques with a more ``modern'' and powerful tool. Furthermore, this paper offers a way to relax a major drawback associated with DL techniques; that is, the proper tuning of the DL regularization parameter. This parameter plays a critical role in DL-based fMRI analysis since it essentially determines the shape and structures of the estimated functional brain networks. However, in actual fMRI data analysis, the lack of ground truth renders the a priori choice of the regularization parameter a truly challenging task. Indeed, the values of the DL regularization parameter, associated with the $\ell_1$ sparsity promoting norm, do not convey any tangible physical meaning. So it is practically difficult to guess its proper value. In this paper, the DL problem is reformulated around a sparsity-promoting constraint that can directly be related to the minimum amount of voxels that the spatial maps of the functional brain networks occupy. Such information is documented and it is readily available to neuroscientists and experts in the field. The proposed method is tested against a number of other popular techniques and the obtained performance gains are reported using a number of synthetic fMRI data. Results with real data have also been obtained in the context of a number of experiments and will be soon reported in a different publication.
Unsupervised learning of the brain connectivity dynamic using residual D-net
Apr 20, 2018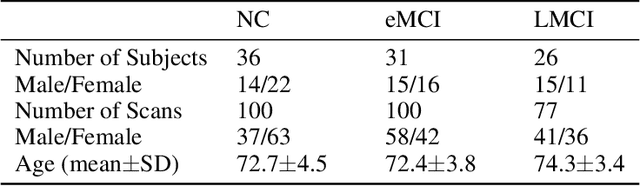
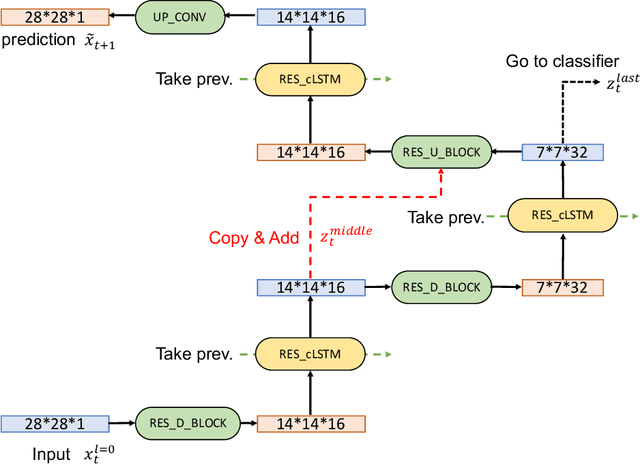
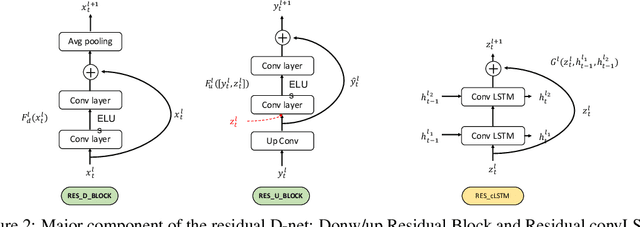
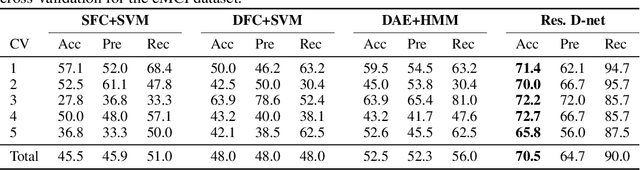
Abstract:In this paper, we propose a novel unsupervised learning method to learn the brain dynamics using a deep learning architecture named residual D-net. As it is often the case in medical research, in contrast to typical deep learning tasks, the size of the resting-state functional Magnetic Resonance Image (rs-fMRI) datasets for training is limited. Thus, the available data should be very efficiently used to learn the complex patterns underneath the brain connectivity dynamics. To address this issue, we use residual connections to alleviate the training complexity through recurrent multi-scale representation. We conduct two classification tasks to differentiate early and late stage Mild Cognitive Impairment (MCI) from Normal healthy Control (NC) subjects. The experiments verify that our proposed residual D-net indeed learns the brain connectivity dynamics, leading to significantly higher classification accuracy compared to previously published techniques.
Online Distributed Learning Over Networks in RKH Spaces Using Random Fourier Features
Mar 24, 2017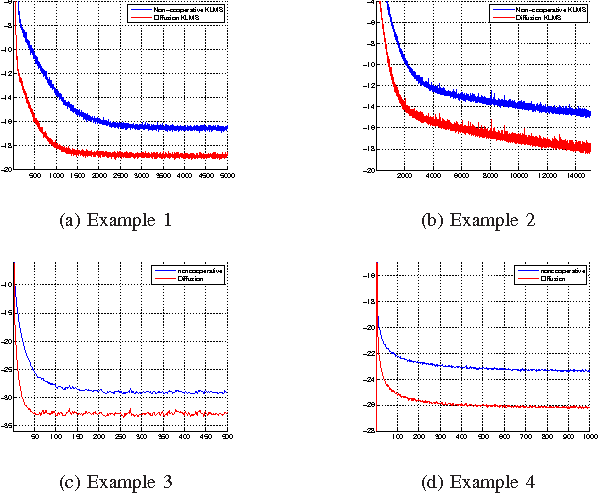
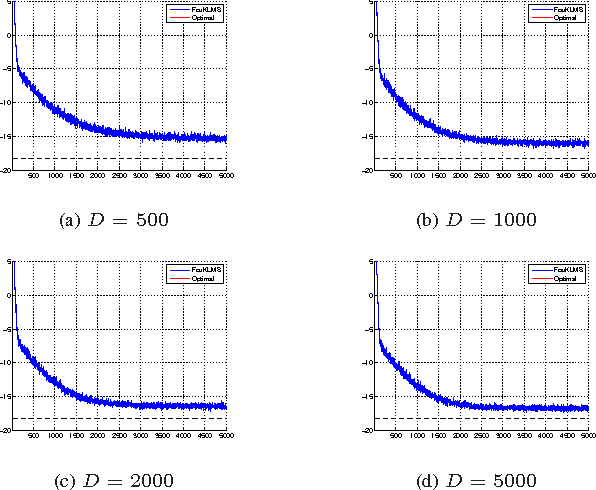
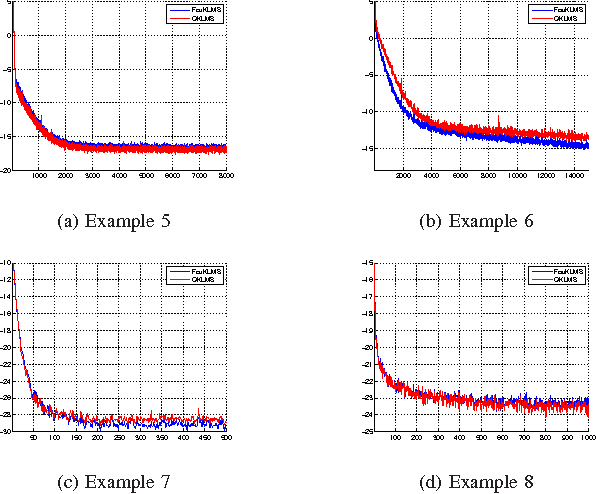
Abstract:We present a novel diffusion scheme for online kernel-based learning over networks. So far, a major drawback of any online learning algorithm, operating in a reproducing kernel Hilbert space (RKHS), is the need for updating a growing number of parameters as time iterations evolve. Besides complexity, this leads to an increased need of communication resources, in a distributed setting. In contrast, the proposed method approximates the solution as a fixed-size vector (of larger dimension than the input space) using Random Fourier Features. This paves the way to use standard linear combine-then-adapt techniques. To the best of our knowledge, this is the first time that a complete protocol for distributed online learning in RKHS is presented. Conditions for asymptotic convergence and boundness of the networkwise regret are also provided. The simulated tests illustrate the performance of the proposed scheme.
Assisted Dictionary Learning for fMRI Data Analysis
Oct 11, 2016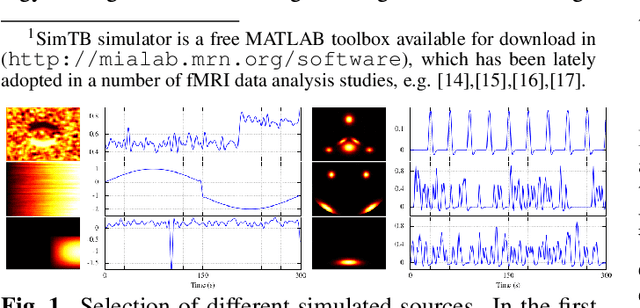
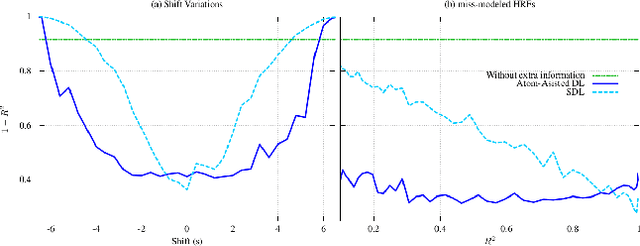
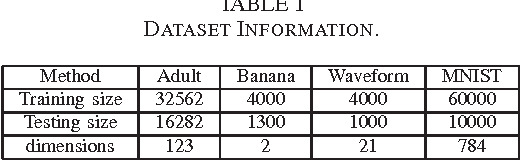
Abstract:Extracting information from functional magnetic resonance (fMRI) images has been a major area of research for more than two decades. The goal of this work is to present a new method for the analysis of fMRI data sets, that is capable to incorporate a priori available information, via an efficient optimization framework. Tests on synthetic data sets demonstrate significant performance gains over existing methods of this kind.
Robust Non-linear Regression: A Greedy Approach Employing Kernels with Application to Image Denoising
Aug 03, 2016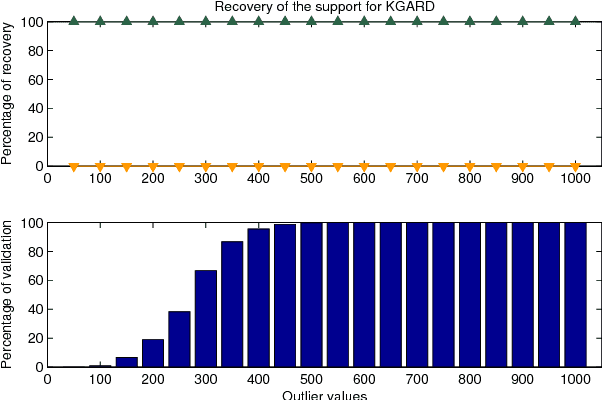
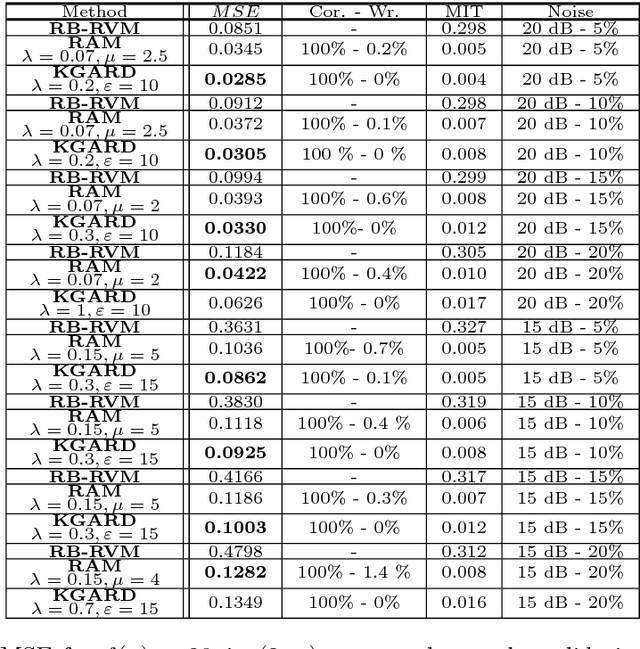
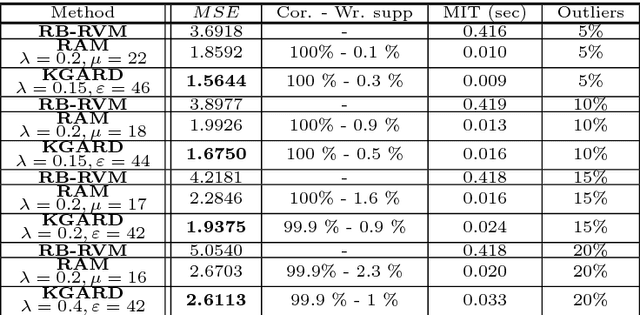
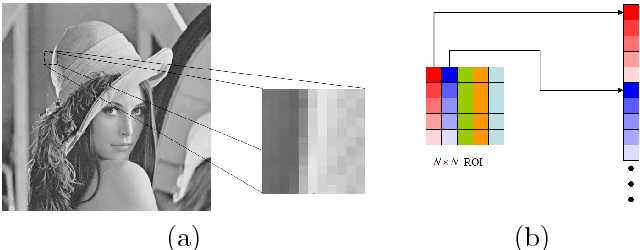
Abstract:We consider the task of robust non-linear regression in the presence of both inlier noise and outliers. Assuming that the unknown non-linear function belongs to a Reproducing Kernel Hilbert Space (RKHS), our goal is to estimate the set of the associated unknown parameters. Due to the presence of outliers, common techniques such as the Kernel Ridge Regression (KRR) or the Support Vector Regression (SVR) turn out to be inadequate. Instead, we employ sparse modeling arguments to explicitly model and estimate the outliers, adopting a greedy approach. The proposed robust scheme, i.e., Kernel Greedy Algorithm for Robust Denoising (KGARD), is inspired by the classical Orthogonal Matching Pursuit (OMP) algorithm. Specifically, the proposed method alternates between a KRR task and an OMP-like selection step. Theoretical results concerning the identification of the outliers are provided. Moreover, KGARD is compared against other cutting edge methods, where its performance is evaluated via a set of experiments with various types of noise. Finally, the proposed robust estimation framework is applied to the task of image denoising, and its enhanced performance in the presence of outliers is demonstrated.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge