Seongjun Yang
Predictive Pipelined Decoding: A Compute-Latency Trade-off for Exact LLM Decoding
Jul 12, 2023



Abstract:This paper presents "Predictive Pipelined Decoding (PPD)," an approach that speeds up greedy decoding in Large Language Models (LLMs) while maintaining the exact same output as the original decoding. Unlike conventional strategies, PPD employs additional compute resources to parallelize the initiation of subsequent token decoding during the current token decoding. This innovative method reduces decoding latency and reshapes the understanding of trade-offs in LLM decoding strategies. We have developed a theoretical framework that allows us to analyze the trade-off between computation and latency. Using this framework, we can analytically estimate the potential reduction in latency associated with our proposed method, achieved through the assessment of the match rate, represented as p_correct. The results demonstrate that the use of extra computational resources has the potential to accelerate LLM greedy decoding.
EHRSQL: A Practical Text-to-SQL Benchmark for Electronic Health Records
Jan 16, 2023



Abstract:We present a new text-to-SQL dataset for electronic health records (EHRs). The utterances were collected from 222 hospital staff, including physicians, nurses, insurance review and health records teams, and more. To construct the QA dataset on structured EHR data, we conducted a poll at a university hospital and templatized the responses to create seed questions. Then, we manually linked them to two open-source EHR databases, MIMIC-III and eICU, and included them with various time expressions and held-out unanswerable questions in the dataset, which were all collected from the poll. Our dataset poses a unique set of challenges: the model needs to 1) generate SQL queries that reflect a wide range of needs in the hospital, including simple retrieval and complex operations such as calculating survival rate, 2) understand various time expressions to answer time-sensitive questions in healthcare, and 3) distinguish whether a given question is answerable or unanswerable based on the prediction confidence. We believe our dataset, EHRSQL, could serve as a practical benchmark to develop and assess QA models on structured EHR data and take one step further towards bridging the gap between text-to-SQL research and its real-life deployment in healthcare. EHRSQL is available at https://github.com/glee4810/EHRSQL.
Universal EHR Federated Learning Framework
Nov 14, 2022



Abstract:Federated learning (FL) is the most practical multi-source learning method for electronic healthcare records (EHR). Despite its guarantee of privacy protection, the wide application of FL is restricted by two large challenges: the heterogeneous EHR systems, and the non-i.i.d. data characteristic. A recent research proposed a framework that unifies heterogeneous EHRs, named UniHPF. We attempt to address both the challenges simultaneously by combining UniHPF and FL. Our study is the first approach to unify heterogeneous EHRs into a single FL framework. This combination provides an average of 3.4% performance gain compared to local learning. We believe that our framework is practically applicable in the real-world FL.
Task Agnostic and Post-hoc Unseen Distribution Detection
Jul 26, 2022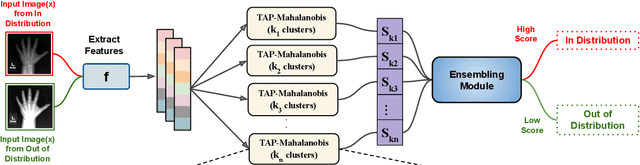
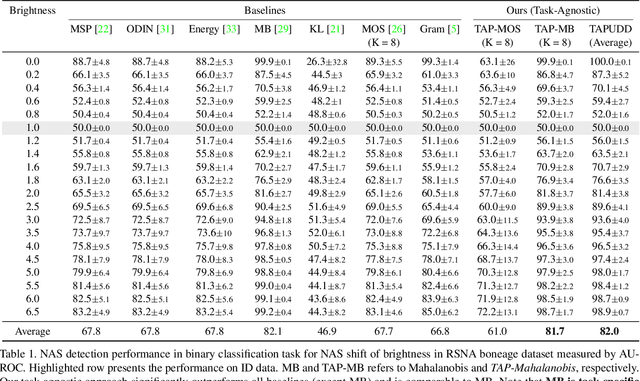
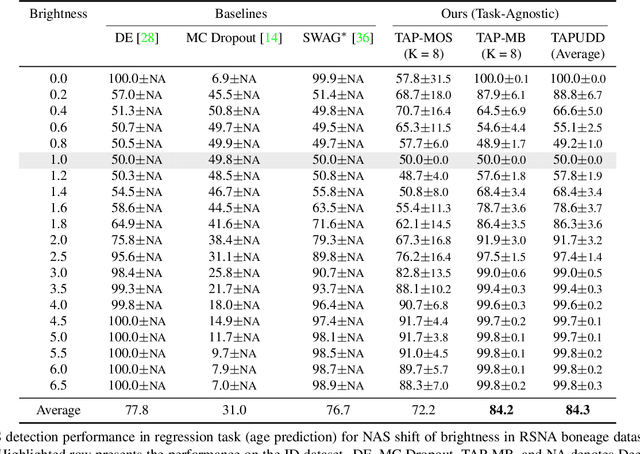
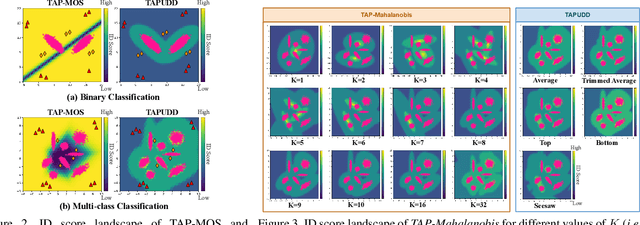
Abstract:Despite the recent advances in out-of-distribution(OOD) detection, anomaly detection, and uncertainty estimation tasks, there do not exist a task-agnostic and post-hoc approach. To address this limitation, we design a novel clustering-based ensembling method, called Task Agnostic and Post-hoc Unseen Distribution Detection (TAPUDD) that utilizes the features extracted from the model trained on a specific task. Explicitly, it comprises of TAP-Mahalanobis, which clusters the training datasets' features and determines the minimum Mahalanobis distance of the test sample from all clusters. Further, we propose the Ensembling module that aggregates the computation of iterative TAP-Mahalanobis for a different number of clusters to provide reliable and efficient cluster computation. Through extensive experiments on synthetic and real-world datasets, we observe that our approach can detect unseen samples effectively across diverse tasks and performs better or on-par with the existing baselines. To this end, we eliminate the necessity of determining the optimal value of the number of clusters and demonstrate that our method is more viable for large-scale classification tasks.
Towards the Practical Utility of Federated Learning in the Medical Domain
Jul 14, 2022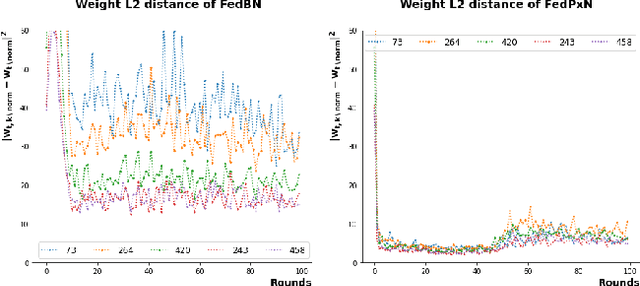
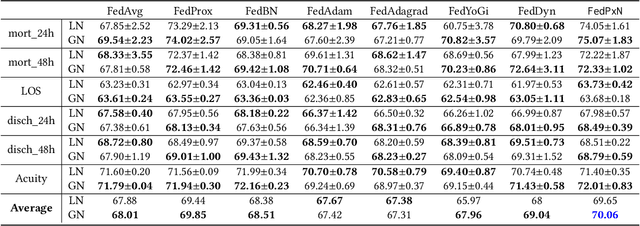
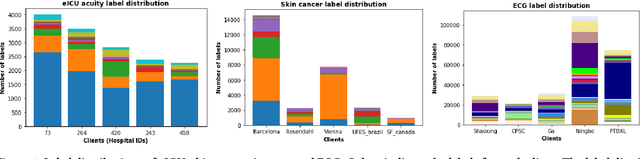
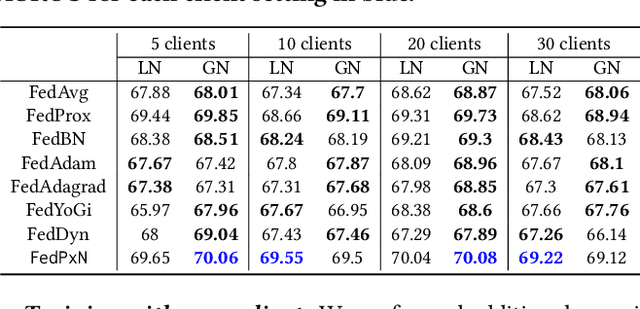
Abstract:Federated learning (FL) is an active area of research. One of the most suitable areas for adopting FL is the medical domain, where patient privacy must be respected. Previous research, however, does not fully consider who will most likely use FL in the medical domain. It is not the hospitals who are eager to adopt FL, but the service providers such as IT companies who want to develop machine learning models with real patient records. Moreover, service providers would prefer to focus on maximizing the performance of the models at the lowest cost possible. In this work, we propose empirical benchmarks of FL methods considering both performance and monetary cost with three real-world datasets: electronic health records, skin cancer images, and electrocardiogram datasets. We also propose Federated learning with Proximal regularization eXcept local Normalization (FedPxN), which, using a simple combination of FedProx and FedBN, outperforms all other FL algorithms while consuming only slightly more power than the most power efficient method.
Improving Lexically Constrained Neural Machine Translation with Source-Conditioned Masked Span Prediction
May 31, 2021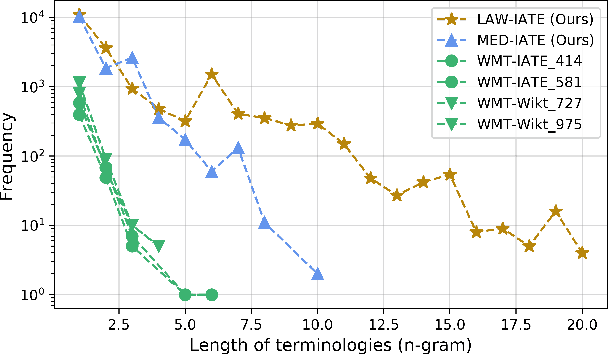
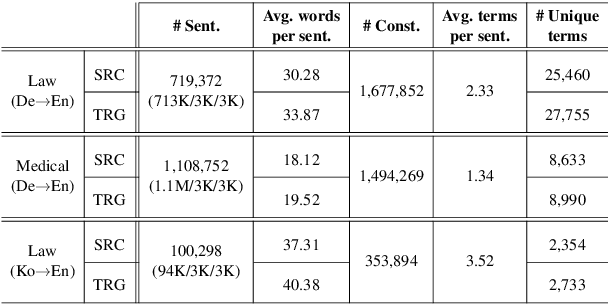
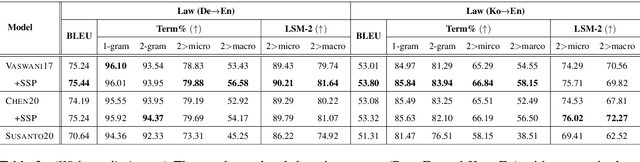
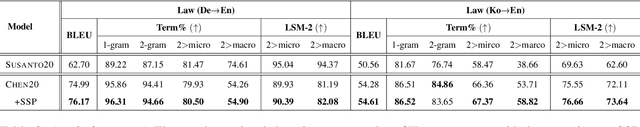
Abstract:Accurate terminology translation is crucial for ensuring the practicality and reliability of neural machine translation (NMT) systems. To address this, lexically constrained NMT explores various methods to ensure pre-specified words and phrases appear in the translation output. However, in many cases, those methods are studied on general domain corpora, where the terms are mostly uni- and bi-grams (>98%). In this paper, we instead tackle a more challenging setup consisting of domain-specific corpora with much longer n-gram and highly specialized terms. Inspired by the recent success of masked span prediction models, we propose a simple and effective training strategy that achieves consistent improvements on both terminology and sentence-level translation for three domain-specific corpora in two language pairs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge