Sen Wu
Slice-based Learning: A Programming Model for Residual Learning in Critical Data Slices
Sep 13, 2019
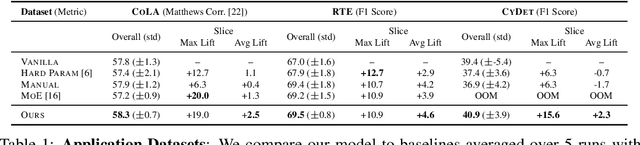
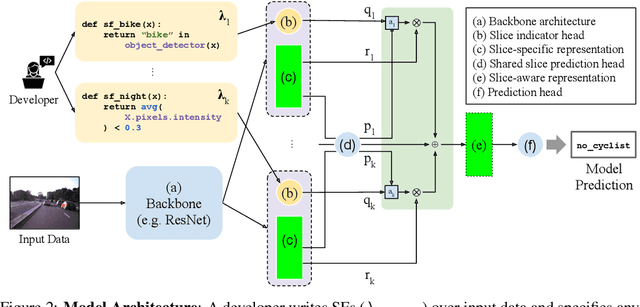
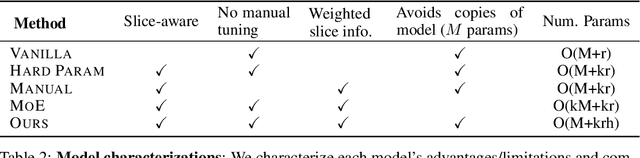
Abstract:In real-world machine learning applications, data subsets correspond to especially critical outcomes: vulnerable cyclist detections are safety-critical in an autonomous driving task, and "question" sentences might be important to a dialogue agent's language understanding for product purposes. While machine learning models can achieve high quality performance on coarse-grained metrics like F1-score and overall accuracy, they may underperform on critical subsets---we define these as slices, the key abstraction in our approach. To address slice-level performance, practitioners often train separate "expert" models on slice subsets or use multi-task hard parameter sharing. We propose Slice-based Learning, a new programming model in which the slicing function (SF), a programming interface, specifies critical data subsets for which the model should commit additional capacity. Any model can leverage SFs to learn slice expert representations, which are combined with an attention mechanism to make slice-aware predictions. We show that our approach maintains a parameter-efficient representation while improving over baselines by up to 19.0 F1 on slices and 4.6 F1 overall on datasets spanning language understanding (e.g. SuperGLUE), computer vision, and production-scale industrial systems.
Snorkel: Rapid Training Data Creation with Weak Supervision
Nov 28, 2017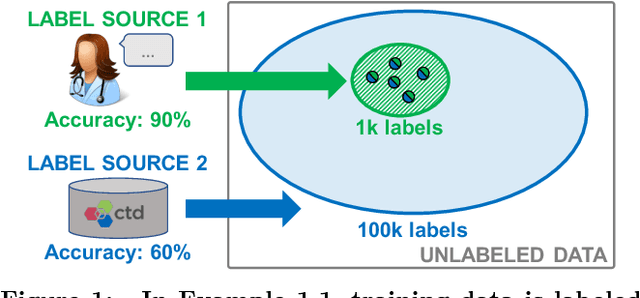
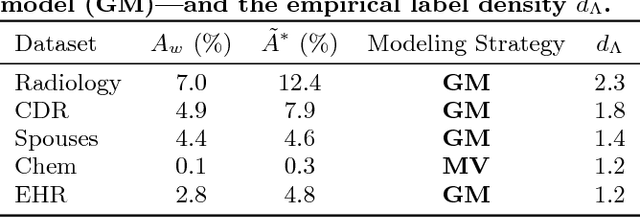
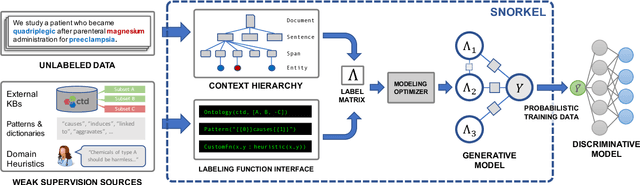
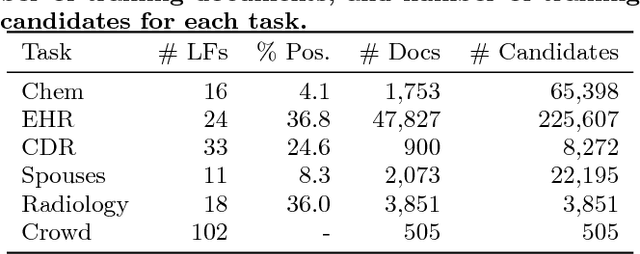
Abstract:Labeling training data is increasingly the largest bottleneck in deploying machine learning systems. We present Snorkel, a first-of-its-kind system that enables users to train state-of-the-art models without hand labeling any training data. Instead, users write labeling functions that express arbitrary heuristics, which can have unknown accuracies and correlations. Snorkel denoises their outputs without access to ground truth by incorporating the first end-to-end implementation of our recently proposed machine learning paradigm, data programming. We present a flexible interface layer for writing labeling functions based on our experience over the past year collaborating with companies, agencies, and research labs. In a user study, subject matter experts build models 2.8x faster and increase predictive performance an average 45.5% versus seven hours of hand labeling. We study the modeling tradeoffs in this new setting and propose an optimizer for automating tradeoff decisions that gives up to 1.8x speedup per pipeline execution. In two collaborations, with the U.S. Department of Veterans Affairs and the U.S. Food and Drug Administration, and on four open-source text and image data sets representative of other deployments, Snorkel provides 132% average improvements to predictive performance over prior heuristic approaches and comes within an average 3.60% of the predictive performance of large hand-curated training sets.
Robust Sparse Coding via Self-Paced Learning
Sep 10, 2017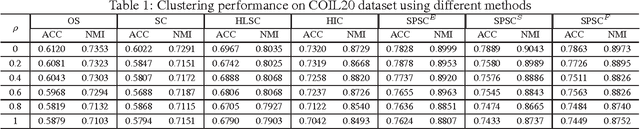

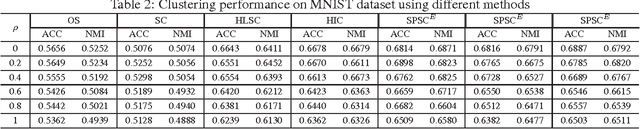
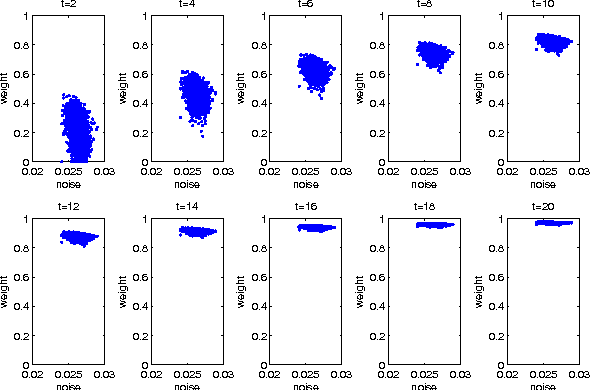
Abstract:Sparse coding (SC) is attracting more and more attention due to its comprehensive theoretical studies and its excellent performance in many signal processing applications. However, most existing sparse coding algorithms are nonconvex and are thus prone to becoming stuck into bad local minima, especially when there are outliers and noisy data. To enhance the learning robustness, in this paper, we propose a unified framework named Self-Paced Sparse Coding (SPSC), which gradually include matrix elements into SC learning from easy to complex. We also generalize the self-paced learning schema into different levels of dynamic selection on samples, features and elements respectively. Experimental results on real-world data demonstrate the efficacy of the proposed algorithms.
SwellShark: A Generative Model for Biomedical Named Entity Recognition without Labeled Data
Apr 20, 2017
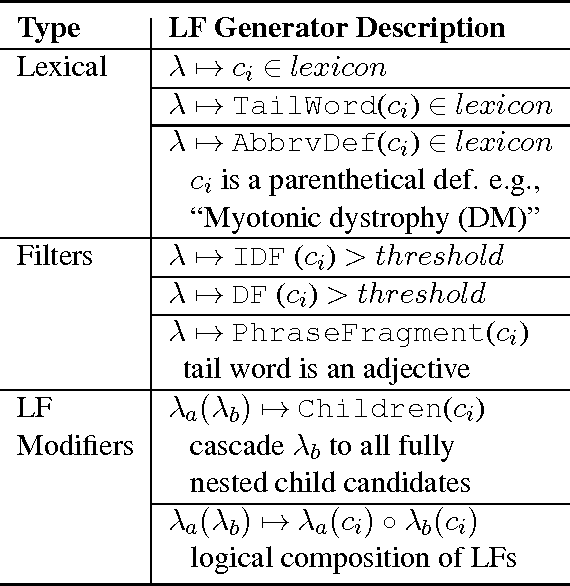

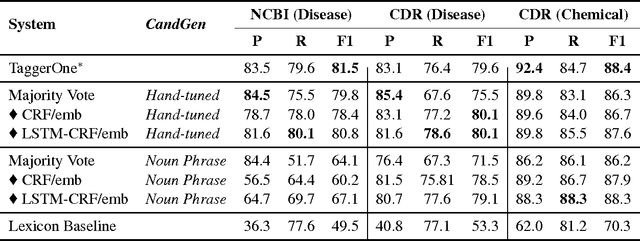
Abstract:We present SwellShark, a framework for building biomedical named entity recognition (NER) systems quickly and without hand-labeled data. Our approach views biomedical resources like lexicons as function primitives for autogenerating weak supervision. We then use a generative model to unify and denoise this supervision and construct large-scale, probabilistically labeled datasets for training high-accuracy NER taggers. In three biomedical NER tasks, SwellShark achieves competitive scores with state-of-the-art supervised benchmarks using no hand-labeled training data. In a drug name extraction task using patient medical records, one domain expert using SwellShark achieved within 5.1% of a crowdsourced annotation approach -- which originally utilized 20 teams over the course of several weeks -- in 24 hours.
Data Programming: Creating Large Training Sets, Quickly
Jan 08, 2017
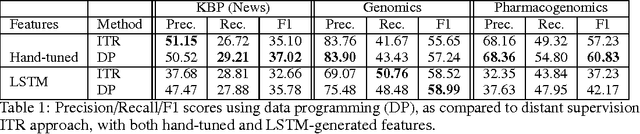
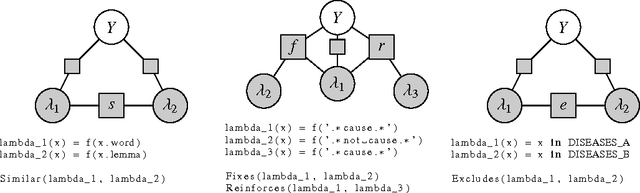
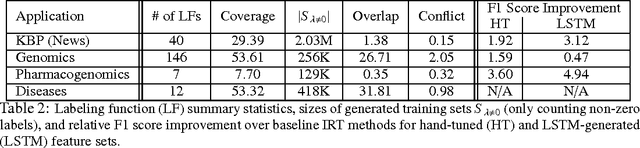
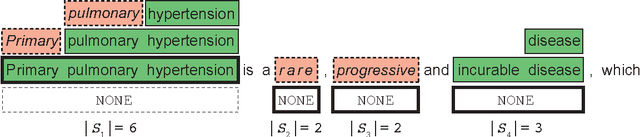
Abstract:Large labeled training sets are the critical building blocks of supervised learning methods and are key enablers of deep learning techniques. For some applications, creating labeled training sets is the most time-consuming and expensive part of applying machine learning. We therefore propose a paradigm for the programmatic creation of training sets called data programming in which users express weak supervision strategies or domain heuristics as labeling functions, which are programs that label subsets of the data, but that are noisy and may conflict. We show that by explicitly representing this training set labeling process as a generative model, we can "denoise" the generated training set, and establish theoretically that we can recover the parameters of these generative models in a handful of settings. We then show how to modify a discriminative loss function to make it noise-aware, and demonstrate our method over a range of discriminative models including logistic regression and LSTMs. Experimentally, on the 2014 TAC-KBP Slot Filling challenge, we show that data programming would have led to a new winning score, and also show that applying data programming to an LSTM model leads to a TAC-KBP score almost 6 F1 points over a state-of-the-art LSTM baseline (and into second place in the competition). Additionally, in initial user studies we observed that data programming may be an easier way for non-experts to create machine learning models when training data is limited or unavailable.
Incremental Knowledge Base Construction Using DeepDive
Jun 15, 2015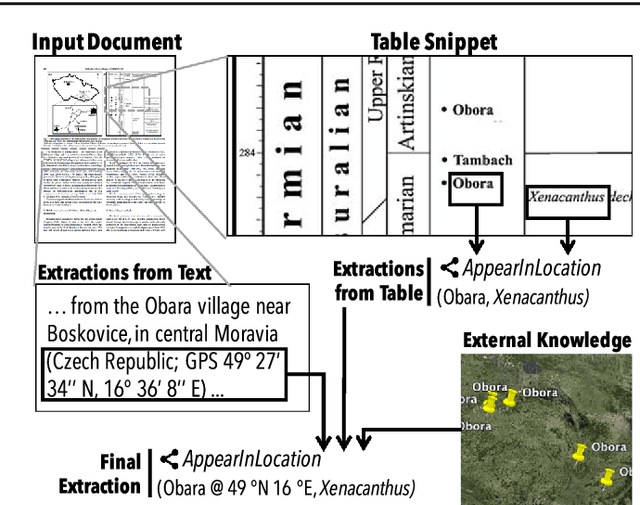
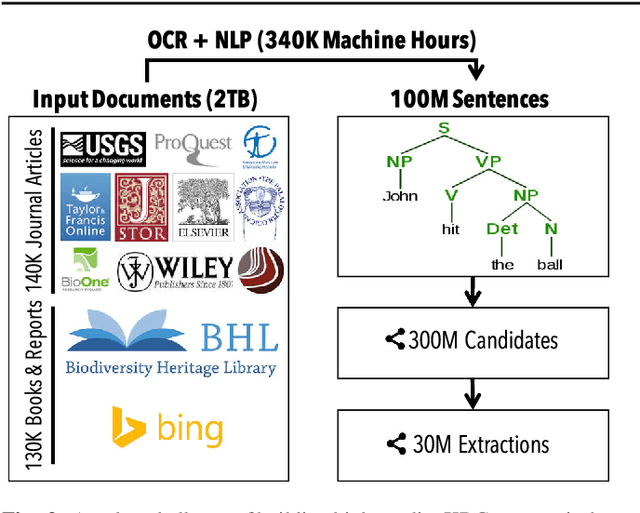
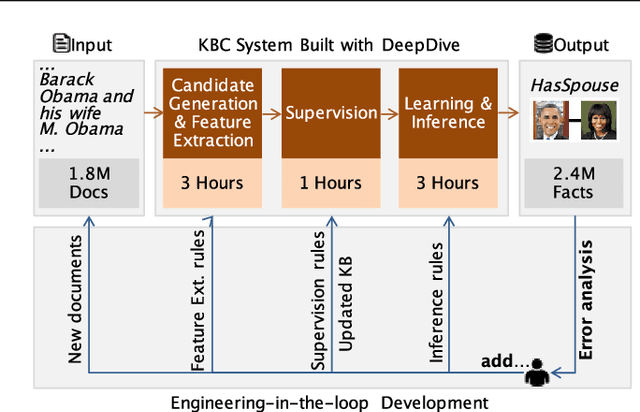
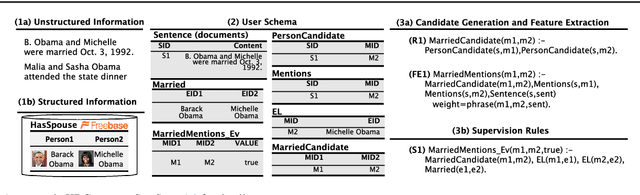
Abstract:Populating a database with unstructured information is a long-standing problem in industry and research that encompasses problems of extraction, cleaning, and integration. Recent names used for this problem include dealing with dark data and knowledge base construction (KBC). In this work, we describe DeepDive, a system that combines database and machine learning ideas to help develop KBC systems, and we present techniques to make the KBC process more efficient. We observe that the KBC process is iterative, and we develop techniques to incrementally produce inference results for KBC systems. We propose two methods for incremental inference, based respectively on sampling and variational techniques. We also study the tradeoff space of these methods and develop a simple rule-based optimizer. DeepDive includes all of these contributions, and we evaluate DeepDive on five KBC systems, showing that it can speed up KBC inference tasks by up to two orders of magnitude with negligible impact on quality.
Feature Engineering for Knowledge Base Construction
Sep 18, 2014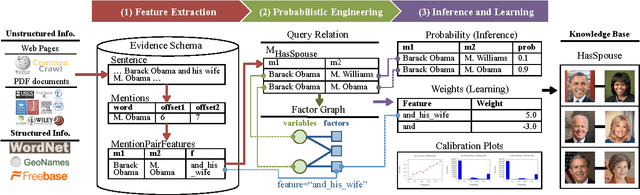
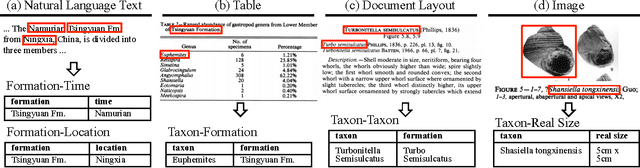
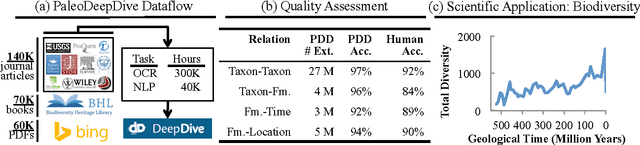

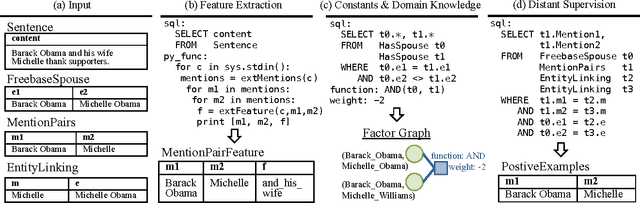
Abstract:Knowledge base construction (KBC) is the process of populating a knowledge base, i.e., a relational database together with inference rules, with information extracted from documents and structured sources. KBC blurs the distinction between two traditional database problems, information extraction and information integration. For the last several years, our group has been building knowledge bases with scientific collaborators. Using our approach, we have built knowledge bases that have comparable and sometimes better quality than those constructed by human volunteers. In contrast to these knowledge bases, which took experts a decade or more human years to construct, many of our projects are constructed by a single graduate student. Our approach to KBC is based on joint probabilistic inference and learning, but we do not see inference as either a panacea or a magic bullet: inference is a tool that allows us to be systematic in how we construct, debug, and improve the quality of such systems. In addition, inference allows us to construct these systems in a more loosely coupled way than traditional approaches. To support this idea, we have built the DeepDive system, which has the design goal of letting the user "think about features---not algorithms." We think of DeepDive as declarative in that one specifies what they want but not how to get it. We describe our approach with a focus on feature engineering, which we argue is an understudied problem relative to its importance to end-to-end quality.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge