Sang Hyun Park
A Meta-Learning Approach for Medical Image Registration
Apr 21, 2021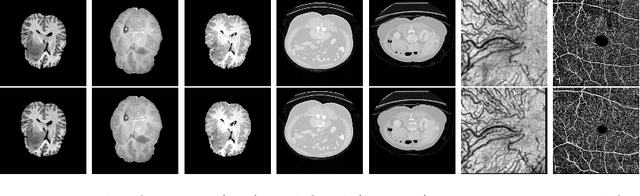
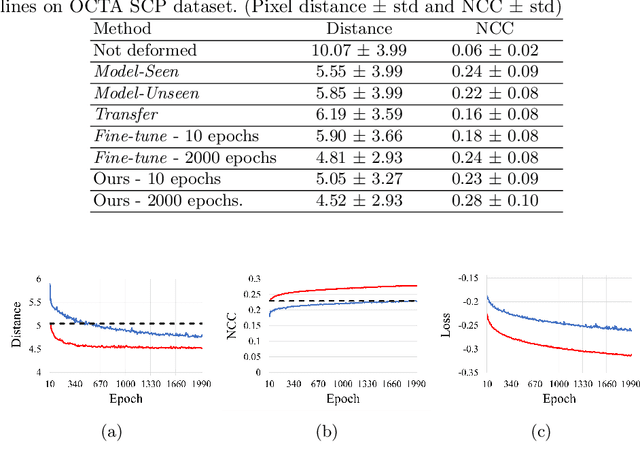
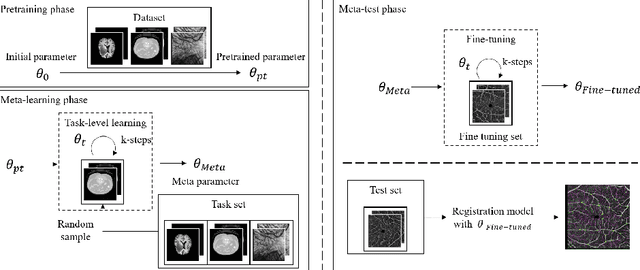
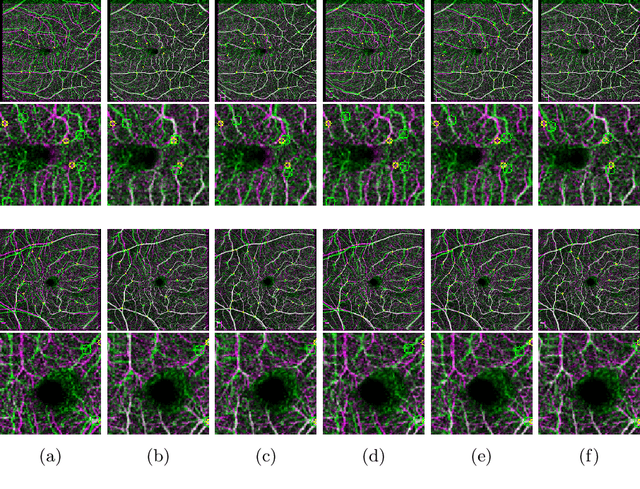
Abstract:Non-rigid registration is a necessary but challenging task in medical imaging studies. Recently, unsupervised registration models have shown good performance, but they often require a large-scale training dataset and long training times. Therefore, in real world application where only dozens to hundreds of image pairs are available, existing models cannot be practically used. To address these limitations, we propose a novel unsupervised registration model which is integrated with a gradient-based meta learning framework. In particular, we train a meta learner which finds an initialization point of parameters by utilizing a variety of existing registration datasets. To quickly adapt to various tasks, the meta learner was updated to get close to the center of parameters which are fine-tuned for each registration task. Thereby, our model can adapt to unseen domain tasks via a short fine-tuning process and perform accurate registration. To verify the superiority of our model, we train the model for various 2D medical image registration tasks such as retinal choroid Optical Coherence Tomography Angiography (OCTA), CT organs, and brain MRI scans and test on registration of retinal OCTA Superficial Capillary Plexus (SCP). In our experiments, the proposed model obtained significantly improved performance in terms of accuracy and training time compared to other registration models.
Self-Supervised Learning based CT Denoising using Pseudo-CT Image Pairs
Apr 06, 2021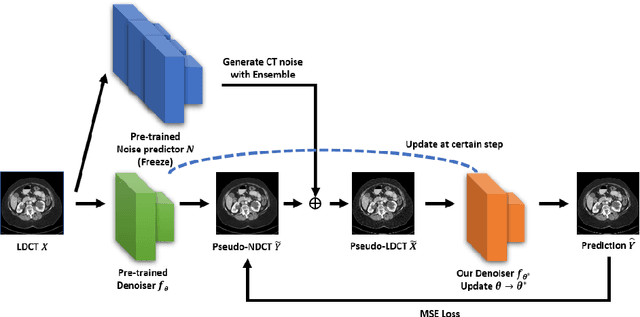
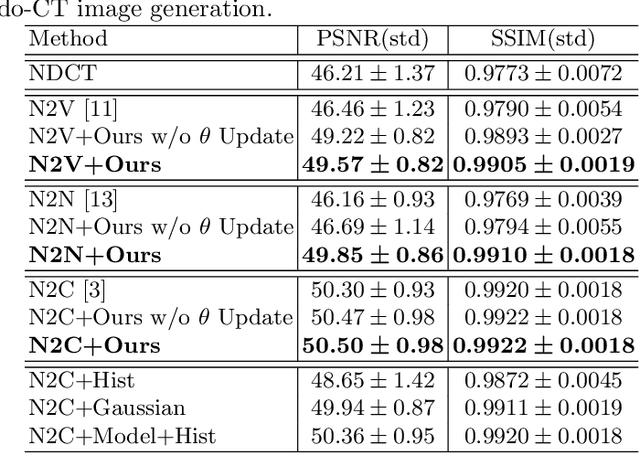
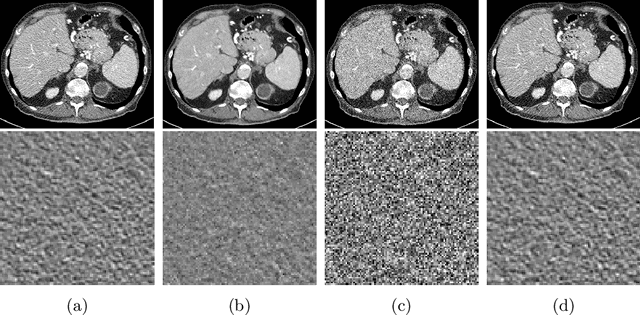
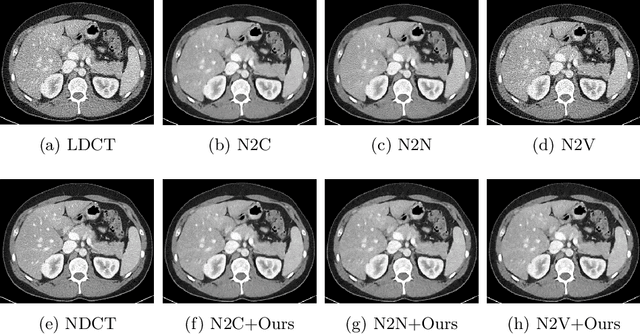
Abstract:Recently, Self-supervised learning methods able to perform image denoising without ground truth labels have been proposed. These methods create low-quality images by adding random or Gaussian noise to images and then train a model for denoising. Ideally, it would be beneficial if one can generate high-quality CT images with only a few training samples via self-supervision. However, the performance of CT denoising is generally limited due to the complexity of CT noise. To address this problem, we propose a novel self-supervised learning-based CT denoising method. In particular, we train pre-train CT denoising and noise models that can predict CT noise from Low-dose CT (LDCT) using available LDCT and Normal-dose CT (NDCT) pairs. For a given test LDCT, we generate Pseudo-LDCT and NDCT pairs using the pre-trained denoising and noise models and then update the parameters of the denoising model using these pairs to remove noise in the test LDCT. To make realistic Pseudo LDCT, we train multiple noise models from individual images and generate the noise using the ensemble of noise models. We evaluate our method on the 2016 AAPM Low-Dose CT Grand Challenge dataset. The proposed ensemble noise model can generate realistic CT noise, and thus our method significantly improves the denoising performance existing denoising models trained by supervised- and self-supervised learning.
Mixing-AdaSIN: Constructing a de-biased dataset using Adaptive Structural Instance Normalization and texture Mixing
Mar 26, 2021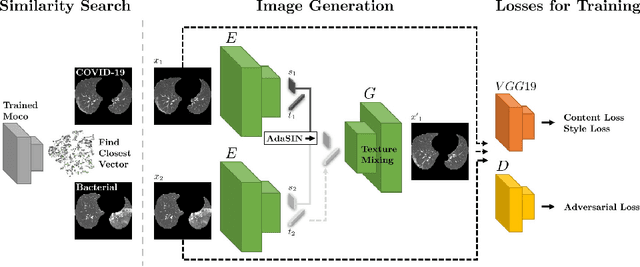
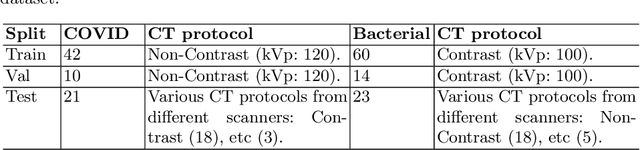
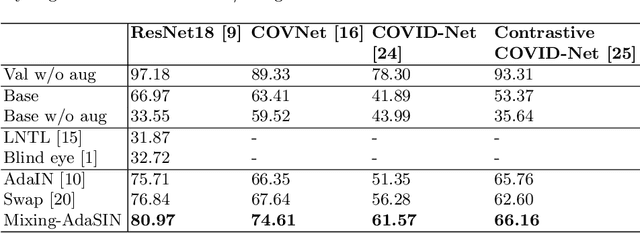
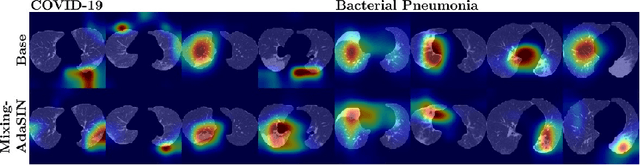
Abstract:Following the pandemic outbreak, several works have proposed to diagnose COVID-19 with deep learning in computed tomography (CT); reporting performance on-par with experts. However, models trained/tested on the same in-distribution data may rely on the inherent data biases for successful prediction, failing to generalize on out-of-distribution samples or CT with different scanning protocols. Early attempts have partly addressed bias-mitigation and generalization through augmentation or re-sampling, but are still limited by collection costs and the difficulty of quantifying bias in medical images. In this work, we propose Mixing-AdaSIN; a bias mitigation method that uses a generative model to generate de-biased images by mixing texture information between different labeled CT scans with semantically similar features. Here, we use Adaptive Structural Instance Normalization (AdaSIN) to enhance de-biasing generation quality and guarantee structural consistency. Following, a classifier trained with the generated images learns to correctly predict the label without bias and generalizes better. To demonstrate the efficacy of our method, we construct a biased COVID-19 vs. bacterial pneumonia dataset based on CT protocols and compare with existing state-of-the-art de-biasing methods. Our experiments show that classifiers trained with de-biased generated images report improved in-distribution performance and generalization on an external COVID-19 dataset.
Bidirectional RNN-based Few Shot Learning for 3D Medical Image Segmentation
Nov 19, 2020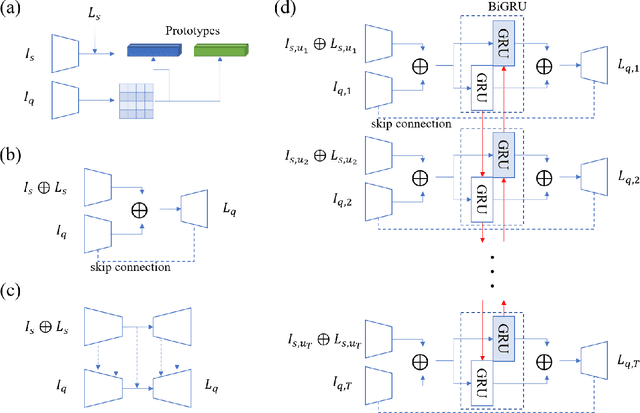
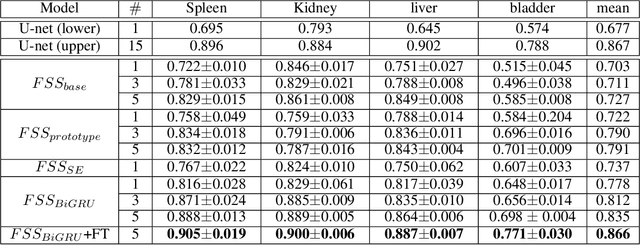
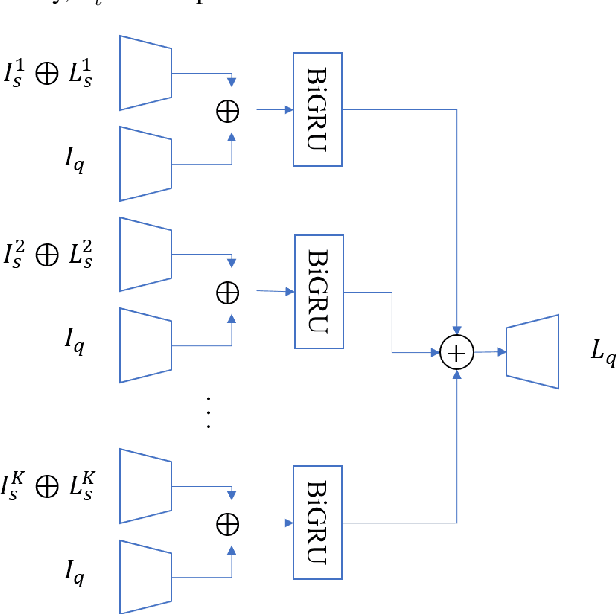
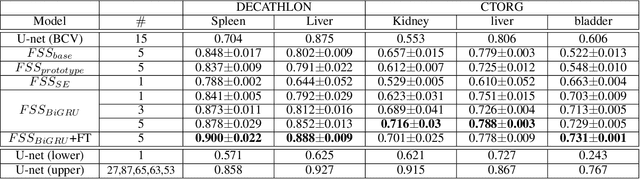
Abstract:Segmentation of organs of interest in 3D medical images is necessary for accurate diagnosis and longitudinal studies. Though recent advances using deep learning have shown success for many segmentation tasks, large datasets are required for high performance and the annotation process is both time consuming and labor intensive. In this paper, we propose a 3D few shot segmentation framework for accurate organ segmentation using limited training samples of the target organ annotation. To achieve this, a U-Net like network is designed to predict segmentation by learning the relationship between 2D slices of support data and a query image, including a bidirectional gated recurrent unit (GRU) that learns consistency of encoded features between adjacent slices. Also, we introduce a transfer learning method to adapt the characteristics of the target image and organ by updating the model before testing with arbitrary support and query data sampled from the support data. We evaluate our proposed model using three 3D CT datasets with annotations of different organs. Our model yielded significantly improved performance over state-of-the-art few shot segmentation models and was comparable to a fully supervised model trained with more target training data.
Few-Shot Relation Learning with Attention for EEG-based Motor Imagery Classification
Mar 03, 2020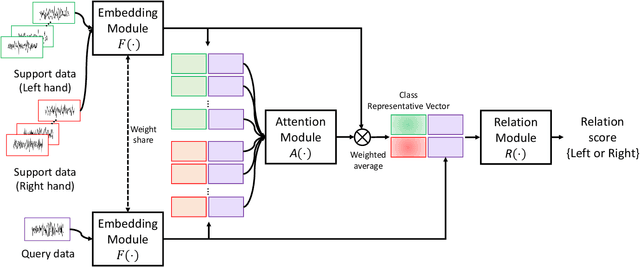
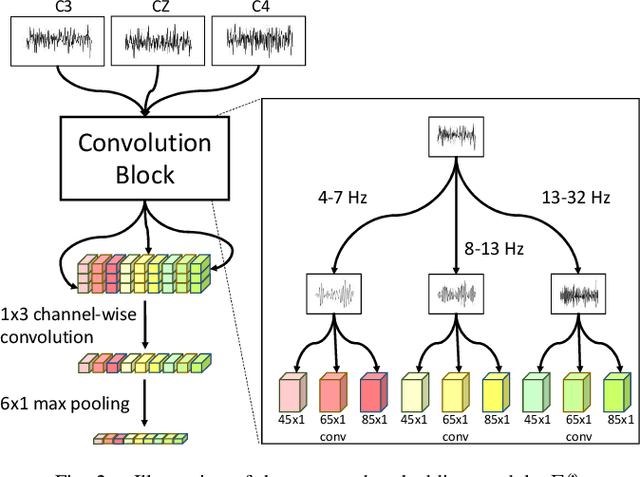
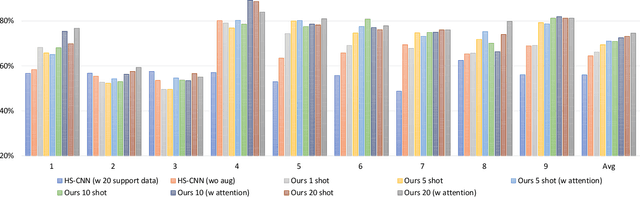
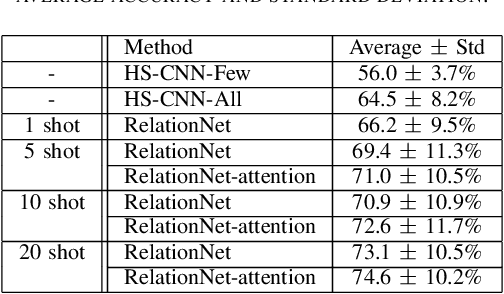
Abstract:Brain-Computer Interfaces (BCI) based on Electroencephalography (EEG) signals, in particular motor imagery (MI) data have received a lot of attention and show the potential towards the design of key technologies both in healthcare and other industries. MI data is generated when a subject imagines movement of limbs and can be used to aid rehabilitation as well as in autonomous driving scenarios. Thus, classification of MI signals is vital for EEG-based BCI systems. Recently, MI EEG classification techniques using deep learning have shown improved performance over conventional techniques. However, due to inter-subject variability, the scarcity of unseen subject data, and low signal-to-noise ratio, extracting robust features and improving accuracy is still challenging. In this context, we propose a novel two-way few shot network that is able to efficiently learn how to learn representative features of unseen subject categories and how to classify them with limited MI EEG data. The pipeline includes an embedding module that learns feature representations from a set of samples, an attention mechanism for key signal feature discovery, and a relation module for final classification based on relation scores between a support set and a query signal. In addition to the unified learning of feature similarity and a few shot classifier, our method leads to emphasize informative features in support data relevant to the query data, which generalizes better on unseen subjects. For evaluation, we used the BCI competition IV 2b dataset and achieved an 9.3% accuracy improvement in the 20-shot classification task with state-of-the-art performance. Experimental results demonstrate the effectiveness of employing attention and the overall generality of our method.
2018 Robotic Scene Segmentation Challenge
Feb 03, 2020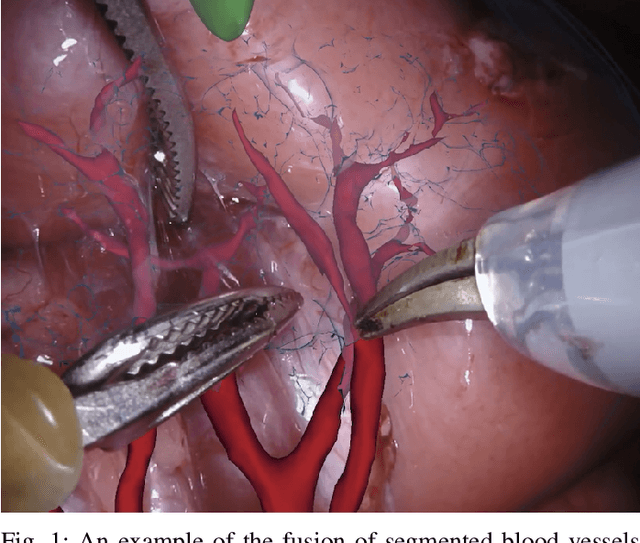
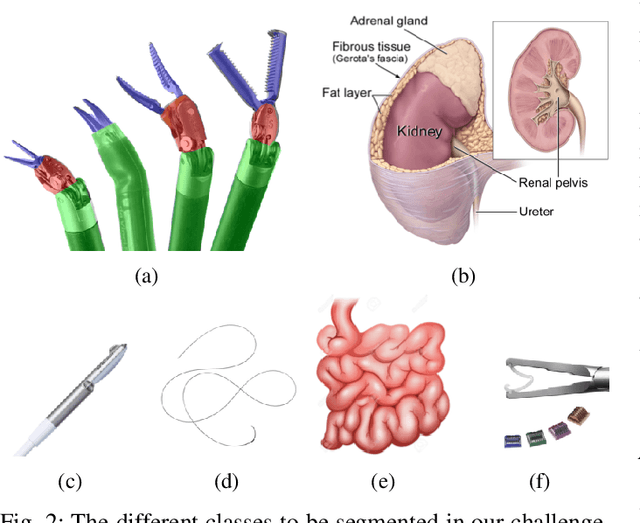
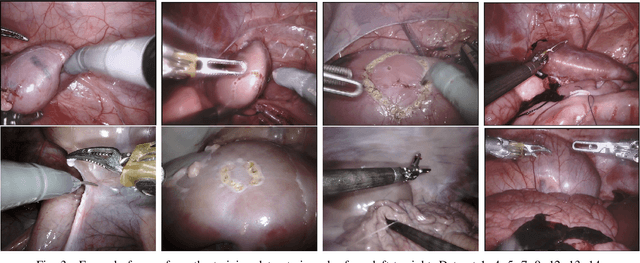
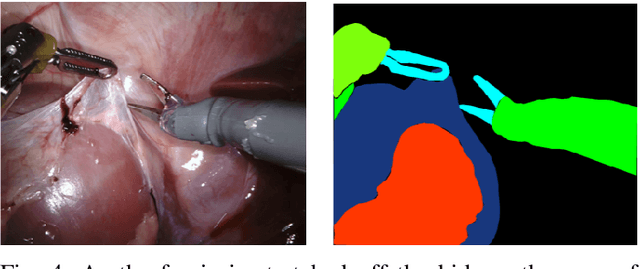
Abstract:In 2015 we began a sub-challenge at the EndoVis workshop at MICCAI in Munich using endoscope images of ex-vivo tissue with automatically generated annotations from robot forward kinematics and instrument CAD models. However, the limited background variation and simple motion rendered the dataset uninformative in learning about which techniques would be suitable for segmentation in real surgery. In 2017, at the same workshop in Quebec we introduced the robotic instrument segmentation dataset with 10 teams participating in the challenge to perform binary, articulating parts and type segmentation of da Vinci instruments. This challenge included realistic instrument motion and more complex porcine tissue as background and was widely addressed with modifications on U-Nets and other popular CNN architectures. In 2018 we added to the complexity by introducing a set of anatomical objects and medical devices to the segmented classes. To avoid over-complicating the challenge, we continued with porcine data which is dramatically simpler than human tissue due to the lack of fatty tissue occluding many organs.
Standardized Assessment of Automatic Segmentation of White Matter Hyperintensities and Results of the WMH Segmentation Challenge
Apr 01, 2019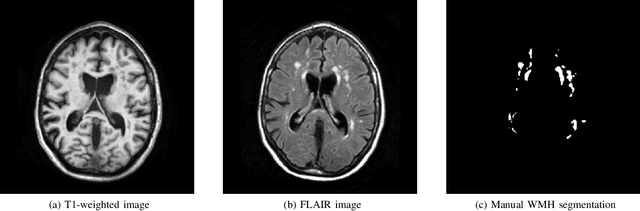
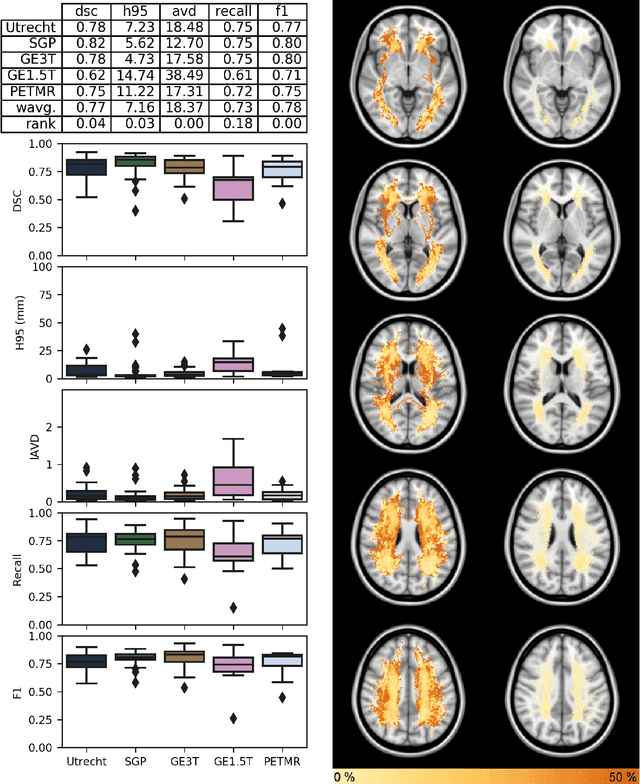
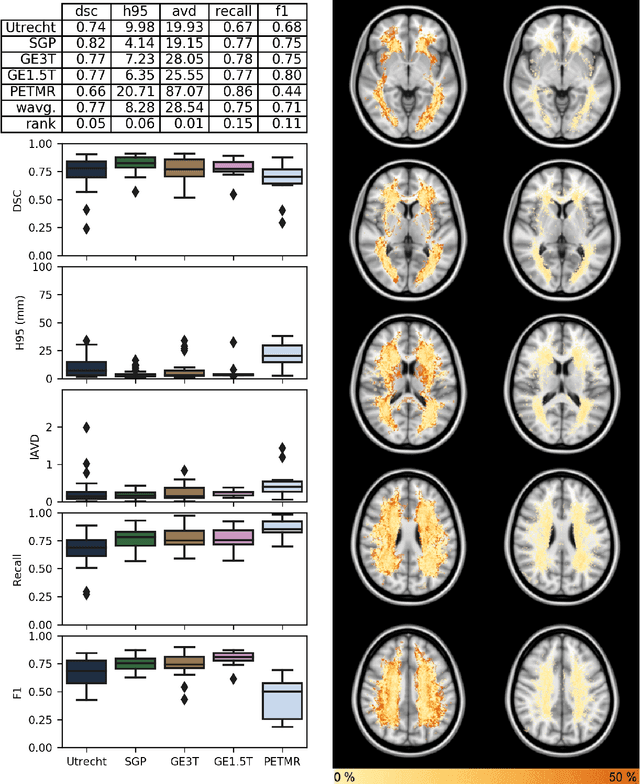
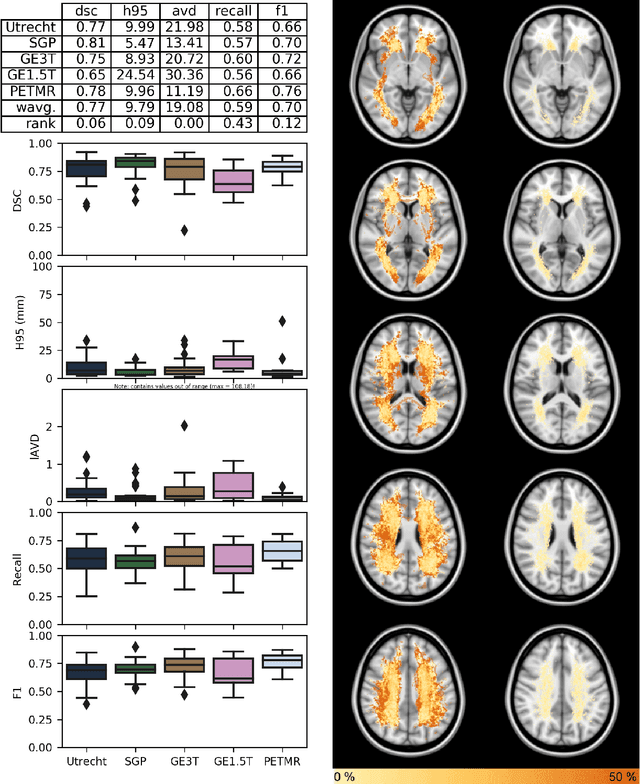
Abstract:Quantification of cerebral white matter hyperintensities (WMH) of presumed vascular origin is of key importance in many neurological research studies. Currently, measurements are often still obtained from manual segmentations on brain MR images, which is a laborious procedure. Automatic WMH segmentation methods exist, but a standardized comparison of the performance of such methods is lacking. We organized a scientific challenge, in which developers could evaluate their method on a standardized multi-center/-scanner image dataset, giving an objective comparison: the WMH Segmentation Challenge (https://wmh.isi.uu.nl/). Sixty T1+FLAIR images from three MR scanners were released with manual WMH segmentations for training. A test set of 110 images from five MR scanners was used for evaluation. Segmentation methods had to be containerized and submitted to the challenge organizers. Five evaluation metrics were used to rank the methods: (1) Dice similarity coefficient, (2) modified Hausdorff distance (95th percentile), (3) absolute log-transformed volume difference, (4) sensitivity for detecting individual lesions, and (5) F1-score for individual lesions. Additionally, methods were ranked on their inter-scanner robustness. Twenty participants submitted their method for evaluation. This paper provides a detailed analysis of the results. In brief, there is a cluster of four methods that rank significantly better than the other methods, with one clear winner. The inter-scanner robustness ranking shows that not all methods generalize to unseen scanners. The challenge remains open for future submissions and provides a public platform for method evaluation.
Facial expression recognition based on local region specific features and support vector machines
Apr 15, 2016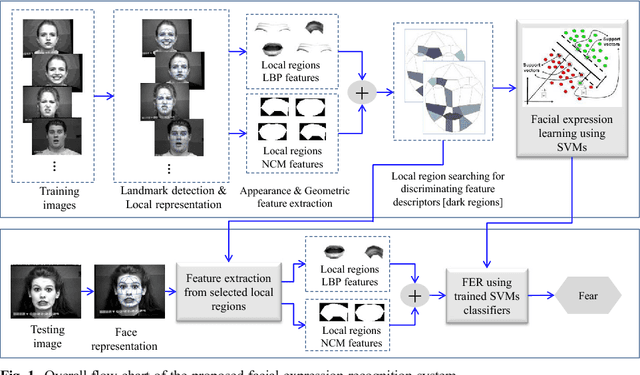
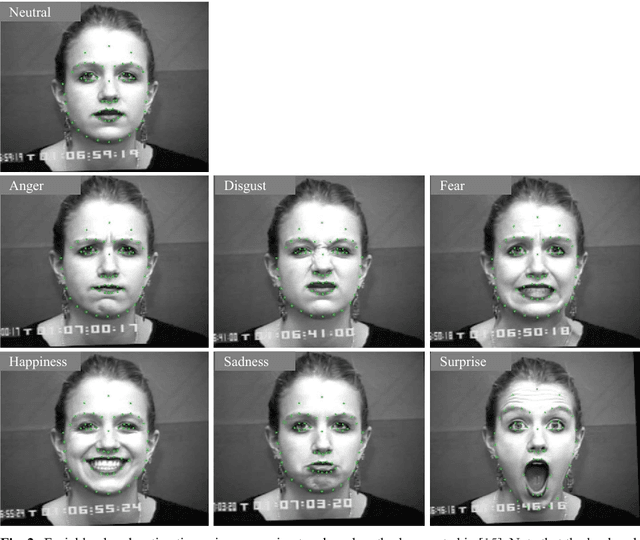
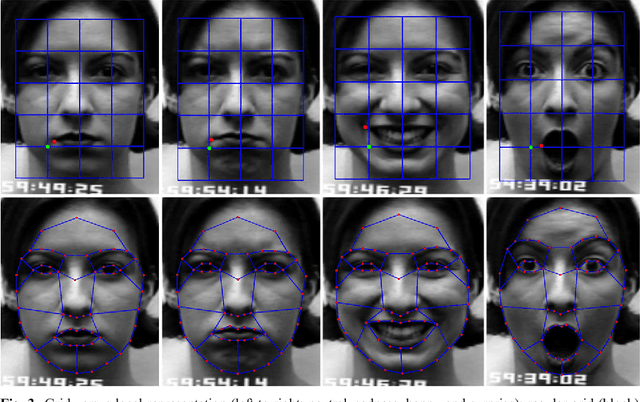
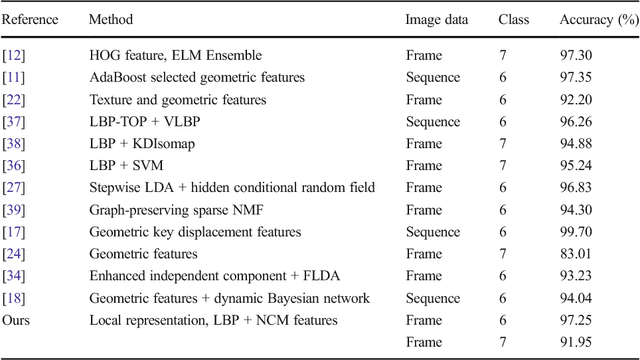


Abstract:Facial expressions are one of the most powerful, natural and immediate means for human being to communicate their emotions and intensions. Recognition of facial expression has many applications including human-computer interaction, cognitive science, human emotion analysis, personality development etc. In this paper, we propose a new method for the recognition of facial expressions from single image frame that uses combination of appearance and geometric features with support vector machines classification. In general, appearance features for the recognition of facial expressions are computed by dividing face region into regular grid (holistic representation). But, in this paper we extracted region specific appearance features by dividing the whole face region into domain specific local regions. Geometric features are also extracted from corresponding domain specific regions. In addition, important local regions are determined by using incremental search approach which results in the reduction of feature dimension and improvement in recognition accuracy. The results of facial expressions recognition using features from domain specific regions are also compared with the results obtained using holistic representation. The performance of the proposed facial expression recognition system has been validated on publicly available extended Cohn-Kanade (CK+) facial expression data sets.
* Facial expressions, Local representation, Appearance features, Geometric features, Support vector machines
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge