Sion An
Few Shot Part Segmentation Reveals Compositional Logic for Industrial Anomaly Detection
Dec 21, 2023Abstract:Logical anomalies (LA) refer to data violating underlying logical constraints e.g., the quantity, arrangement, or composition of components within an image. Detecting accurately such anomalies requires models to reason about various component types through segmentation. However, curation of pixel-level annotations for semantic segmentation is both time-consuming and expensive. Although there are some prior few-shot or unsupervised co-part segmentation algorithms, they often fail on images with industrial object. These images have components with similar textures and shapes, and a precise differentiation proves challenging. In this study, we introduce a novel component segmentation model for LA detection that leverages a few labeled samples and unlabeled images sharing logical constraints. To ensure consistent segmentation across unlabeled images, we employ a histogram matching loss in conjunction with an entropy loss. As segmentation predictions play a crucial role, we propose to enhance both local and global sample validity detection by capturing key aspects from visual semantics via three memory banks: class histograms, component composition embeddings and patch-level representations. For effective LA detection, we propose an adaptive scaling strategy to standardize anomaly scores from different memory banks in inference. Extensive experiments on the public benchmark MVTec LOCO AD reveal our method achieves 98.1% AUROC in LA detection vs. 89.6% from competing methods.
Self-Supervised Learning based CT Denoising using Pseudo-CT Image Pairs
Apr 06, 2021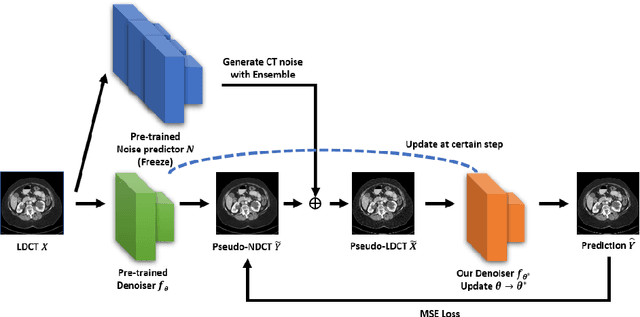
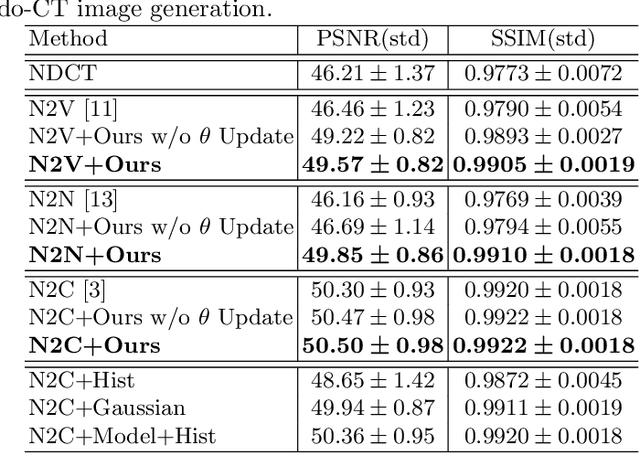
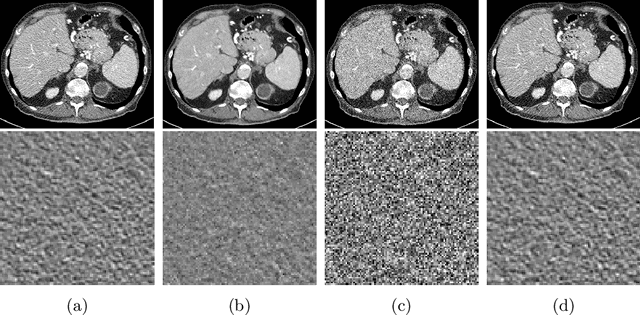
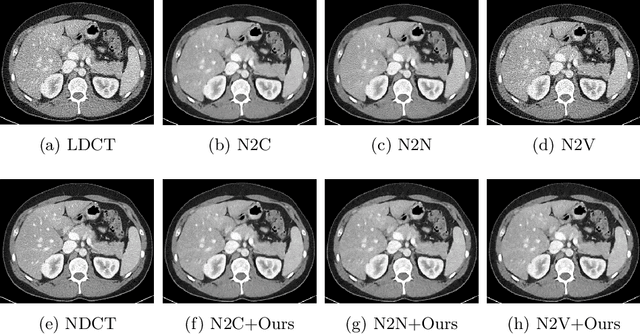
Abstract:Recently, Self-supervised learning methods able to perform image denoising without ground truth labels have been proposed. These methods create low-quality images by adding random or Gaussian noise to images and then train a model for denoising. Ideally, it would be beneficial if one can generate high-quality CT images with only a few training samples via self-supervision. However, the performance of CT denoising is generally limited due to the complexity of CT noise. To address this problem, we propose a novel self-supervised learning-based CT denoising method. In particular, we train pre-train CT denoising and noise models that can predict CT noise from Low-dose CT (LDCT) using available LDCT and Normal-dose CT (NDCT) pairs. For a given test LDCT, we generate Pseudo-LDCT and NDCT pairs using the pre-trained denoising and noise models and then update the parameters of the denoising model using these pairs to remove noise in the test LDCT. To make realistic Pseudo LDCT, we train multiple noise models from individual images and generate the noise using the ensemble of noise models. We evaluate our method on the 2016 AAPM Low-Dose CT Grand Challenge dataset. The proposed ensemble noise model can generate realistic CT noise, and thus our method significantly improves the denoising performance existing denoising models trained by supervised- and self-supervised learning.
Bidirectional RNN-based Few Shot Learning for 3D Medical Image Segmentation
Nov 19, 2020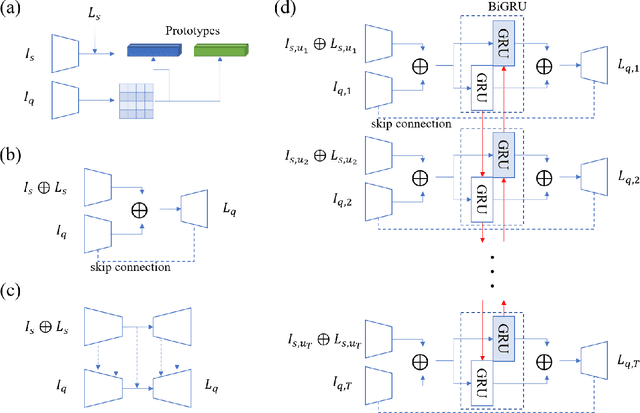
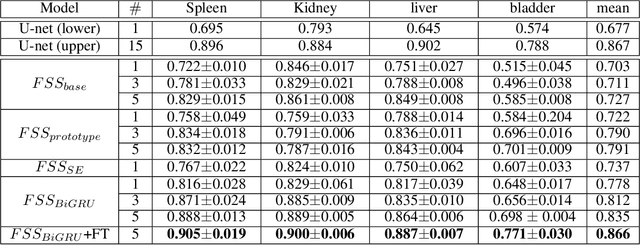
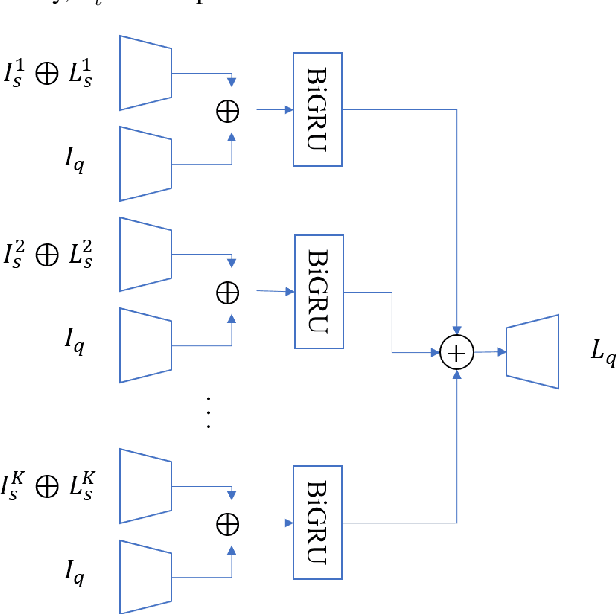
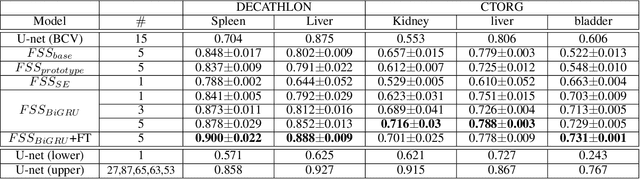
Abstract:Segmentation of organs of interest in 3D medical images is necessary for accurate diagnosis and longitudinal studies. Though recent advances using deep learning have shown success for many segmentation tasks, large datasets are required for high performance and the annotation process is both time consuming and labor intensive. In this paper, we propose a 3D few shot segmentation framework for accurate organ segmentation using limited training samples of the target organ annotation. To achieve this, a U-Net like network is designed to predict segmentation by learning the relationship between 2D slices of support data and a query image, including a bidirectional gated recurrent unit (GRU) that learns consistency of encoded features between adjacent slices. Also, we introduce a transfer learning method to adapt the characteristics of the target image and organ by updating the model before testing with arbitrary support and query data sampled from the support data. We evaluate our proposed model using three 3D CT datasets with annotations of different organs. Our model yielded significantly improved performance over state-of-the-art few shot segmentation models and was comparable to a fully supervised model trained with more target training data.
Few-Shot Relation Learning with Attention for EEG-based Motor Imagery Classification
Mar 03, 2020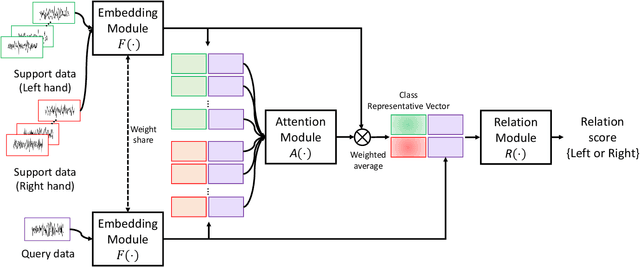
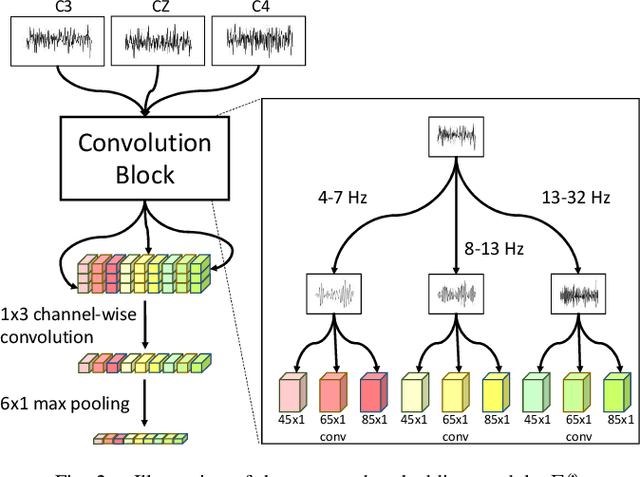
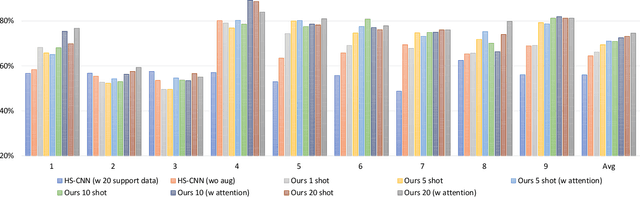
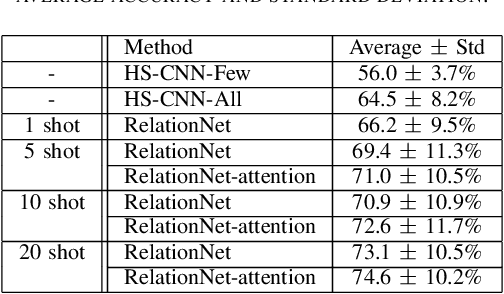
Abstract:Brain-Computer Interfaces (BCI) based on Electroencephalography (EEG) signals, in particular motor imagery (MI) data have received a lot of attention and show the potential towards the design of key technologies both in healthcare and other industries. MI data is generated when a subject imagines movement of limbs and can be used to aid rehabilitation as well as in autonomous driving scenarios. Thus, classification of MI signals is vital for EEG-based BCI systems. Recently, MI EEG classification techniques using deep learning have shown improved performance over conventional techniques. However, due to inter-subject variability, the scarcity of unseen subject data, and low signal-to-noise ratio, extracting robust features and improving accuracy is still challenging. In this context, we propose a novel two-way few shot network that is able to efficiently learn how to learn representative features of unseen subject categories and how to classify them with limited MI EEG data. The pipeline includes an embedding module that learns feature representations from a set of samples, an attention mechanism for key signal feature discovery, and a relation module for final classification based on relation scores between a support set and a query signal. In addition to the unified learning of feature similarity and a few shot classifier, our method leads to emphasize informative features in support data relevant to the query data, which generalizes better on unseen subjects. For evaluation, we used the BCI competition IV 2b dataset and achieved an 9.3% accuracy improvement in the 20-shot classification task with state-of-the-art performance. Experimental results demonstrate the effectiveness of employing attention and the overall generality of our method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge