Rhodri H. Davies
PSIRNet: Deep Learning-based Free-breathing Rapid Acquisition Late Enhancement Imaging
Apr 09, 2026Abstract:Purpose: To develop and evaluate a deep learning (DL) method for free-breathing phase-sensitive inversion recovery (PSIR) late gadolinium enhancement (LGE) cardiac MRI that produces diagnostic-quality images from a single acquisition over two heartbeats, eliminating the need for 8 to 24 motion-corrected (MOCO) signal averages. Materials and Methods: Raw data comprising 800,653 slices from 55,917 patients, acquired on 1.5T and 3T scanners across multiple sites from 2016 to 2024, were used in this retrospective study. Data were split by patient: 640,000 slices (42,822 patients) for training and the remainder for validation and testing, without overlap. The training and testing data were from different institutions. PSIRNet, a physics-guided DL network with 845 million parameters, was trained end-to-end to reconstruct PSIR images with surface coil correction from a single interleaved IR/PD acquisition over two heartbeats. Reconstruction quality was evaluated using SSIM, PSNR, and NRMSE against MOCO PSIR references. Two expert cardiologists performed an independent qualitative assessment, scoring image quality on a 5-point Likert scale across bright blood, dark blood, and wideband LGE variants. Paired superiority and equivalence (margin = 0.25 Likert points) were tested using exact Wilcoxon signed-rank tests at a significance level of 0.05 using R version 4.5.2. Results: Both readers rated single-average PSIRNet reconstructions superior to MOCO PSIR for dark blood LGE (conservative P = .002); for bright blood and wideband, one reader rated it superior and the other confirmed equivalence (all P < .001). Inference required approximately 100 msec per slice versus more than 5 sec for MOCO PSIR. Conclusion: PSIRNet produces diagnostic-quality free-breathing PSIR LGE images from a single acquisition, enabling 8- to 24-fold reduction in acquisition time.
Imaging Transformer for MRI Denoising: a Scalable Model Architecture that enables SNR << 1 Imaging
Apr 13, 2025



Abstract:Purpose: To propose a flexible and scalable imaging transformer (IT) architecture with three attention modules for multi-dimensional imaging data and apply it to MRI denoising with very low input SNR. Methods: Three independent attention modules were developed: spatial local, spatial global, and frame attentions. They capture long-range signal correlation and bring back the locality of information in images. An attention-cell-block design processes 5D tensors ([B, C, F, H, W]) for 2D, 2D+T, and 3D image data. A High Resolution (HRNet) backbone was built to hold IT blocks. Training dataset consists of 206,677 cine series and test datasets had 7,267 series. Ten input SNR levels from 0.05 to 8.0 were tested. IT models were compared to seven convolutional and transformer baselines. To test scalability, four IT models 27m to 218m parameters were trained. Two senior cardiologists reviewed IT model outputs from which the EF was measured and compared against the ground-truth. Results: IT models significantly outperformed other models over the tested SNR levels. The performance gap was most prominent at low SNR levels. The IT-218m model had the highest SSIM and PSNR, restoring good image quality and anatomical details even at SNR 0.2. Two experts agreed at this SNR or above, the IT model output gave the same clinical interpretation as the ground-truth. The model produced images that had accurate EF measurements compared to ground-truth values. Conclusions: Imaging transformer model offers strong performance, scalability, and versatility for MR denoising. It recovers image quality suitable for confident clinical reading and accurate EF measurement, even at very low input SNR of 0.2.
A 3D deep learning classifier and its explainability when assessing coronary artery disease
Jul 29, 2023Abstract:Early detection and diagnosis of coronary artery disease (CAD) could save lives and reduce healthcare costs. In this study, we propose a 3D Resnet-50 deep learning model to directly classify normal subjects and CAD patients on computed tomography coronary angiography images. Our proposed method outperforms a 2D Resnet-50 model by 23.65%. Explainability is also provided by using a Grad-GAM. Furthermore, we link the 3D CAD classification to a 2D two-class semantic segmentation for improved explainability and accurate abnormality localisation.
Improving the generalizability of convolutional neural network-based segmentation on CMR images
Jul 03, 2019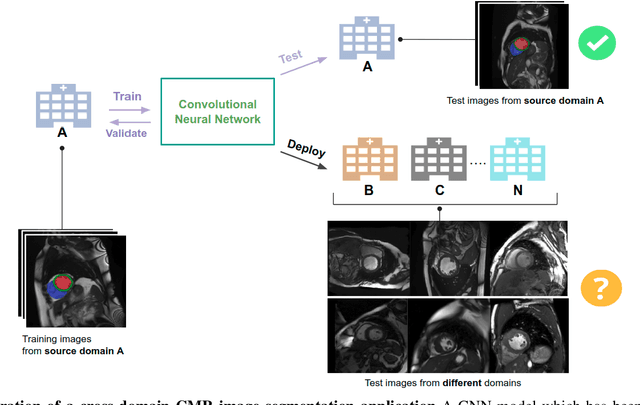
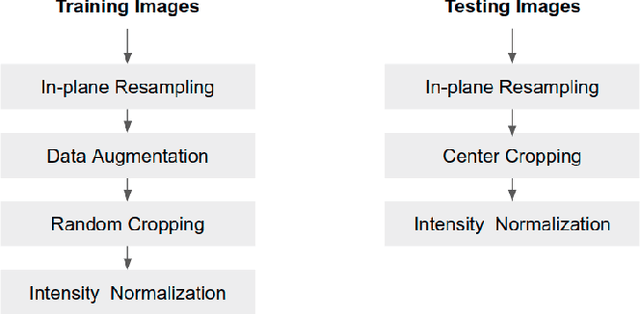
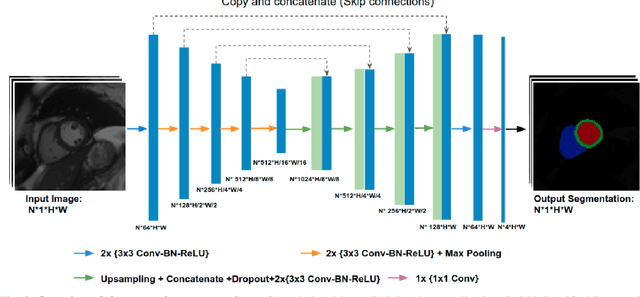

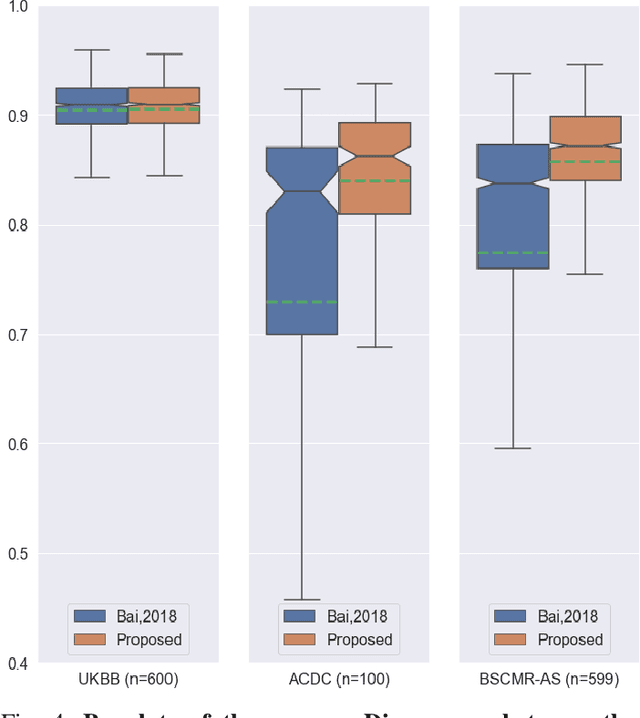
Abstract:Convolutional neural network (CNN) based segmentation methods provide an efficient and automated way for clinicians to assess the structure and function of the heart in cardiac MR images. While CNNs can generally perform the segmentation tasks with high accuracy when training and test images come from the same domain (e.g. same scanner or site), their performance often degrades dramatically on images from different scanners or clinical sites. We propose a simple yet effective way for improving the network generalization ability by carefully designing data normalization and augmentation strategies to accommodate common scenarios in multi-site, multi-scanner clinical imaging data sets. We demonstrate that a neural network trained on a single-site single-scanner dataset from the UK Biobank can be successfully applied to segmenting cardiac MR images across different sites and different scanners without substantial loss of accuracy. Specifically, the method was trained on a large set of 3,975 subjects from the UK Biobank. It was then directly tested on 600 different subjects from the UK Biobank for intra-domain testing and two other sets for cross-domain testing: the ACDC dataset (100 subjects, 1 site, 2 scanners) and the BSCMR-AS dataset (599 subjects, 6 sites, 9 scanners). The proposed method produces promising segmentation results on the UK Biobank test set which are comparable to previously reported values in the literature, while also performing well on cross-domain test sets, achieving a mean Dice metric of 0.90 for the left ventricle, 0.81 for the myocardium and 0.82 for the right ventricle on the ACDC dataset; and 0.89 for the left ventricle, 0.83 for the myocardium on the BSCMR-AS dataset. The proposed method offers a potential solution to improve CNN-based model generalizability for the cross-scanner and cross-site cardiac MR image segmentation task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge