Rajesh Ranganath
New York University
Causal Estimation with Functional Confounders
Feb 17, 2021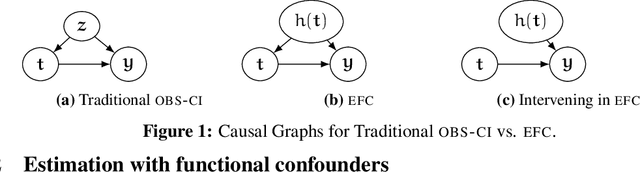
Abstract:Causal inference relies on two fundamental assumptions: ignorability and positivity. We study causal inference when the true confounder value can be expressed as a function of the observed data; we call this setting estimation with functional confounders (EFC). In this setting, ignorability is satisfied, however positivity is violated, and causal inference is impossible in general. We consider two scenarios where causal effects are estimable. First, we discuss interventions on a part of the treatment called functional interventions and a sufficient condition for effect estimation of these interventions called functional positivity. Second, we develop conditions for nonparametric effect estimation based on the gradient fields of the functional confounder and the true outcome function. To estimate effects under these conditions, we develop Level-set Orthogonal Descent Estimation (LODE). Further, we prove error bounds on LODE's effect estimates, evaluate our methods on simulated and real data, and empirically demonstrate the value of EFC.
X-CAL: Explicit Calibration for Survival Analysis
Jan 13, 2021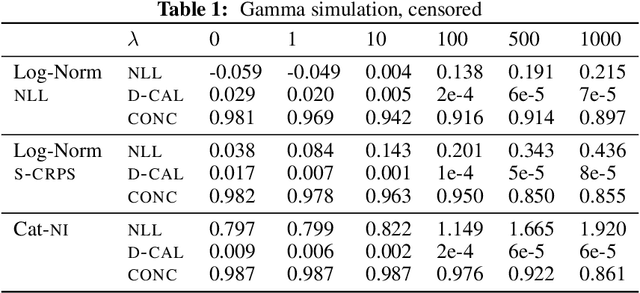
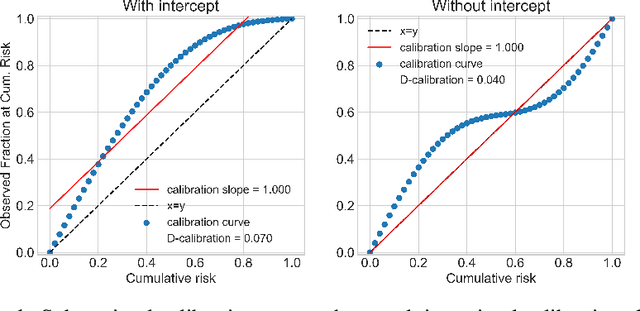
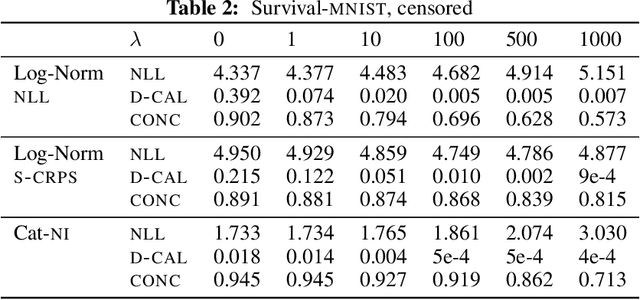
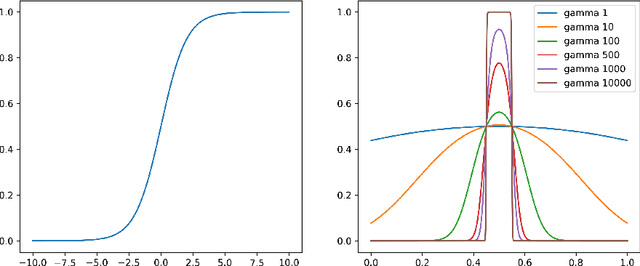
Abstract:Survival analysis models the distribution of time until an event of interest, such as discharge from the hospital or admission to the ICU. When a model's predicted number of events within any time interval is similar to the observed number, it is called well-calibrated. A survival model's calibration can be measured using, for instance, distributional calibration (D-CALIBRATION) [Haider et al., 2020] which computes the squared difference between the observed and predicted number of events within different time intervals. Classically, calibration is addressed in post-training analysis. We develop explicit calibration (X-CAL), which turns D-CALIBRATION into a differentiable objective that can be used in survival modeling alongside maximum likelihood estimation and other objectives. X-CAL allows practitioners to directly optimize calibration and strike a desired balance between predictive power and calibration. In our experiments, we fit a variety of shallow and deep models on simulated data, a survival dataset based on MNIST, on length-of-stay prediction using MIMIC-III data, and on brain cancer data from The Cancer Genome Atlas. We show that the models we study can be miscalibrated. We give experimental evidence on these datasets that X-CAL improves D-CALIBRATION without a large decrease in concordance or likelihood.
Probabilistic Machine Learning for Healthcare
Sep 23, 2020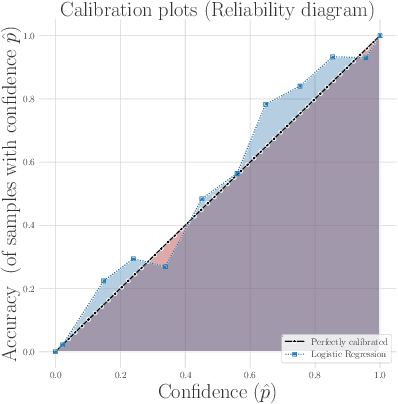
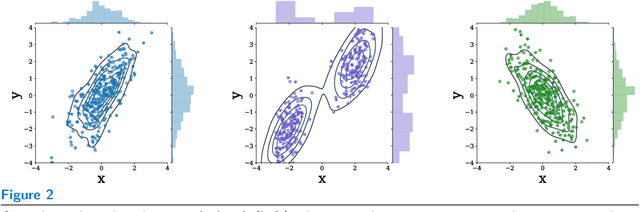
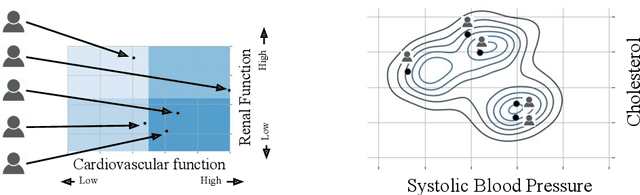
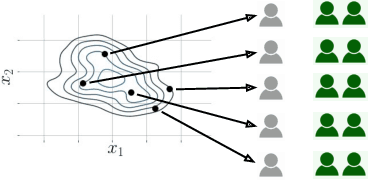
Abstract:Machine learning can be used to make sense of healthcare data. Probabilistic machine learning models help provide a complete picture of observed data in healthcare. In this review, we examine how probabilistic machine learning can advance healthcare. We consider challenges in the predictive model building pipeline where probabilistic models can be beneficial including calibration and missing data. Beyond predictive models, we also investigate the utility of probabilistic machine learning models in phenotyping, in generative models for clinical use cases, and in reinforcement learning.
Deep Direct Likelihood Knockoffs
Jul 31, 2020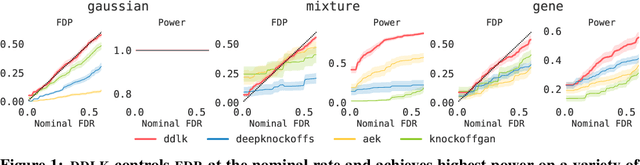
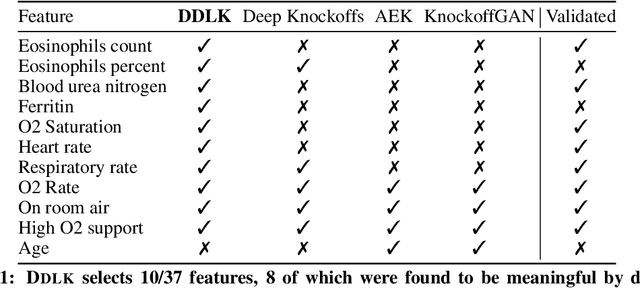
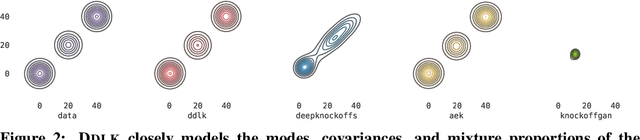
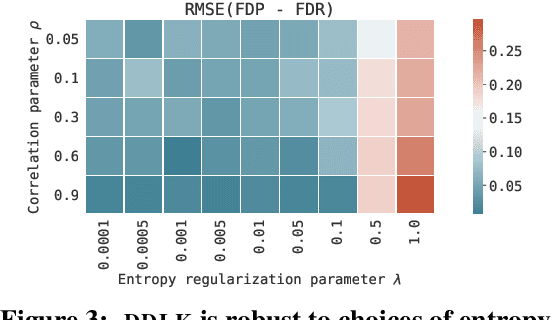
Abstract:Predictive modeling often uses black box machine learning methods, such as deep neural networks, to achieve state-of-the-art performance. In scientific domains, the scientist often wishes to discover which features are actually important for making the predictions. These discoveries may lead to costly follow-up experiments and as such it is important that the error rate on discoveries is not too high. Model-X knockoffs enable important features to be discovered with control of the FDR. However, knockoffs require rich generative models capable of accurately modeling the knockoff features while ensuring they obey the so-called "swap" property. We develop Deep Direct Likelihood Knockoffs (DDLK), which directly minimizes the KL divergence implied by the knockoff swap property. DDLK consists of two stages: it first maximizes the explicit likelihood of the features, then minimizes the KL divergence between the joint distribution of features and knockoffs and any swap between them. To ensure that the generated knockoffs are valid under any possible swap, DDLK uses the Gumbel-Softmax trick to optimize the knockoff generator under the worst-case swap. We find DDLK has higher power than baselines while controlling the false discovery rate on a variety of synthetic and real benchmarks including a task involving a large dataset from one of the epicenters of COVID-19.
Overfitting and Optimization in Offline Policy Learning
Jun 27, 2020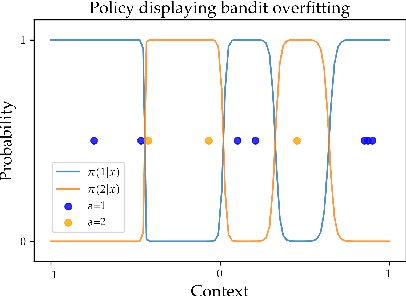
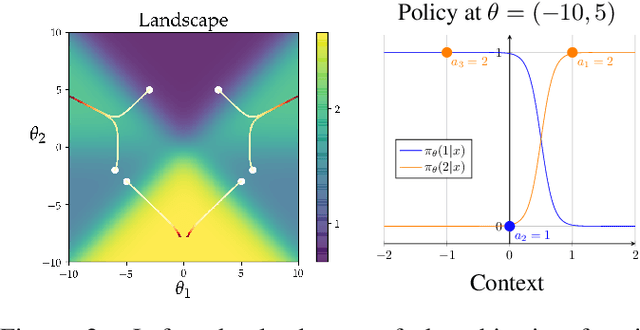
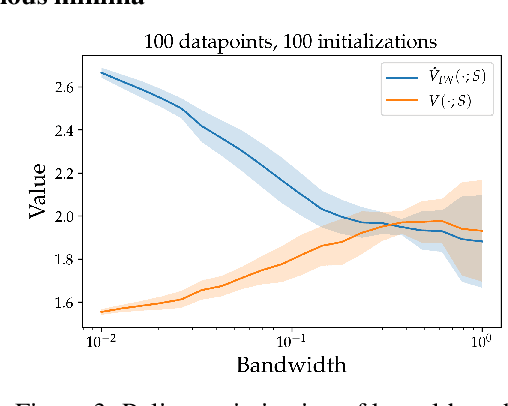
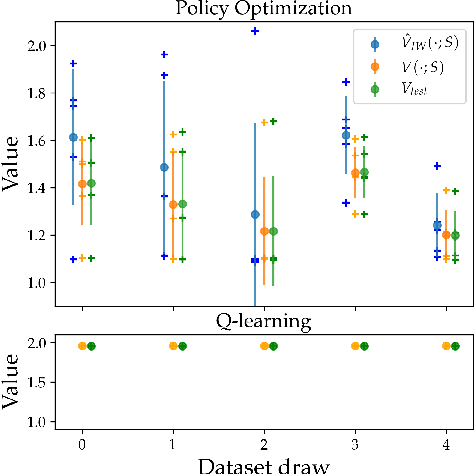
Abstract:We consider the task of policy learning from an offline dataset generated by some behavior policy. We analyze the two most prominent families of algorithms for this task: policy optimization and Q-learning. We demonstrate that policy optimization suffers from two problems, overfitting and spurious minima, that do not appear in Q-learning or full-feedback problems (i.e. cost-sensitive classification). Specifically, we describe the phenomenon of ``bandit overfitting'' in which an algorithm overfits based on the actions observed in the dataset, and show that it affects policy optimization but not Q-learning. Moreover, we show that the policy optimization objective suffers from spurious minima even with linear policies, whereas the Q-learning objective is convex for linear models. We empirically verify the existence of both problems in realistic datasets with neural network models.
The Counterfactual $χ$-GAN
Jan 09, 2020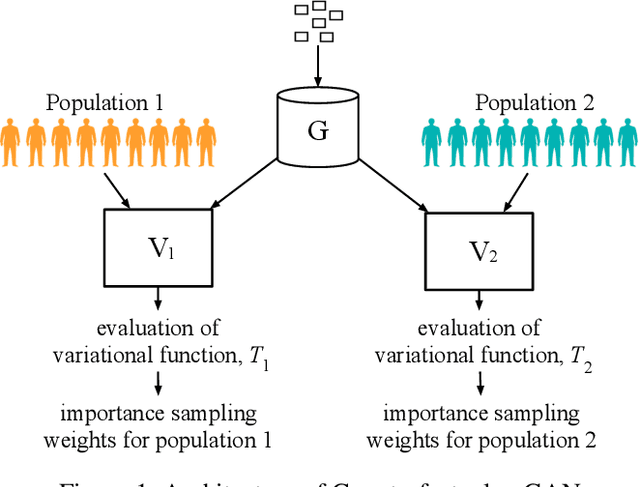
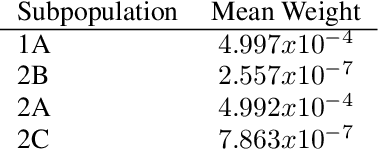
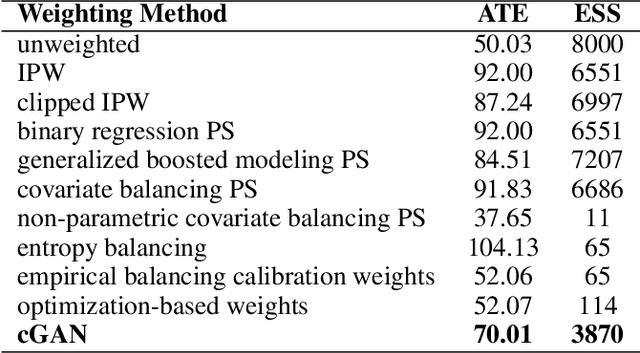
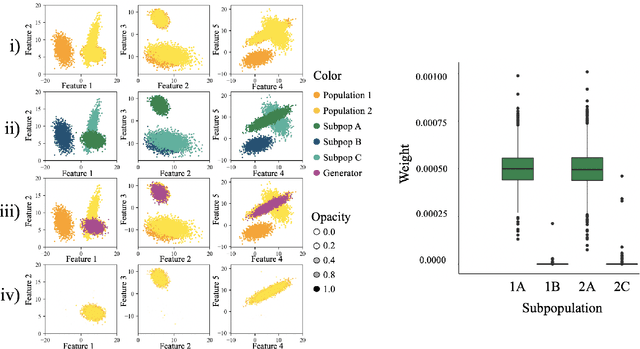
Abstract:Causal inference often relies on the counterfactual framework, which requires that treatment assignment is independent of the outcome, known as strong ignorability. Approaches to enforcing strong ignorability in causal analyses of observational data include weighting and matching methods. Effect estimates, such as the average treatment effect (ATE), are then estimated as expectations under the reweighted or matched distribution, P . The choice of P is important and can impact the interpretation of the effect estimate and the variance of effect estimates. In this work, instead of specifying P, we learn a distribution that simultaneously maximizes coverage and minimizes variance of ATE estimates. In order to learn this distribution, this research proposes a generative adversarial network (GAN)-based model called the Counterfactual $\chi$-GAN (cGAN), which also learns feature-balancing weights and supports unbiased causal estimation in the absence of unobserved confounding. Our model minimizes the Pearson $\chi^2$ divergence, which we show simultaneously maximizes coverage and minimizes the variance of importance sampling estimates. To our knowledge, this is the first such application of the Pearson $\chi^2$ divergence. We demonstrate the effectiveness of cGAN in achieving feature balance relative to established weighting methods in simulation and with real-world medical data.
Energy-Inspired Models: Learning with Sampler-Induced Distributions
Oct 31, 2019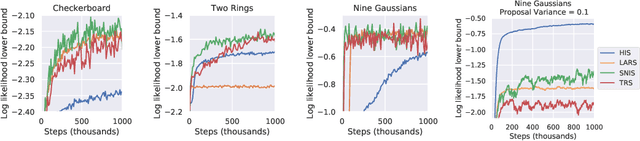
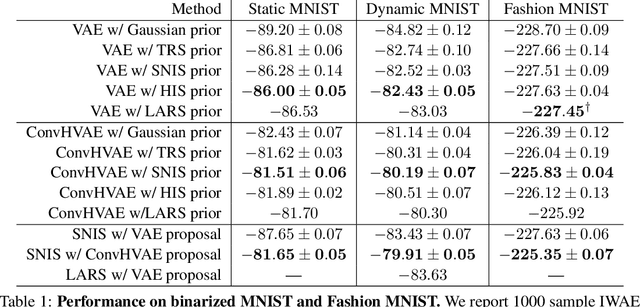
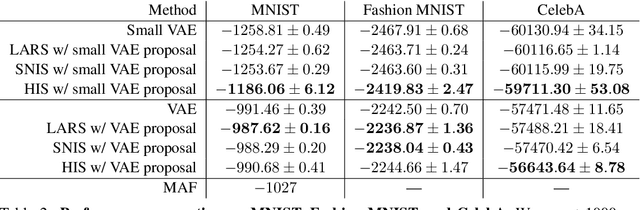
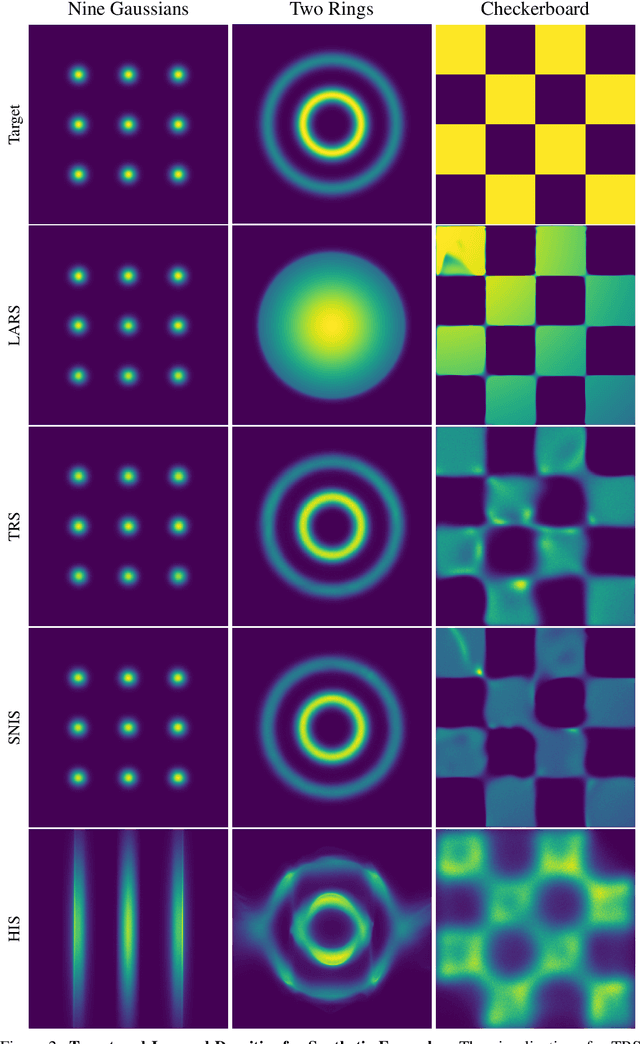
Abstract:Energy-based models (EBMs) are powerful probabilistic models, but suffer from intractable sampling and density evaluation due to the partition function. As a result, inference in EBMs relies on approximate sampling algorithms, leading to a mismatch between the model and inference. Motivated by this, we consider the sampler-induced distribution as the model of interest and maximize the likelihood of this model. This yields a class of energy-inspired models (EIMs) that incorporate learned energy functions while still providing exact samples and tractable log-likelihood lower bounds. We describe and evaluate three instantiations of such models based on truncated rejection sampling, self-normalized importance sampling, and Hamiltonian importance sampling. These models outperform or perform comparably to the recently proposed Learned Accept/Reject Sampling algorithm and provide new insights on ranking Noise Contrastive Estimation and Contrastive Predictive Coding. Moreover, EIMs allow us to generalize a recent connection between multi-sample variational lower bounds and auxiliary variable variational inference. We show how recent variational bounds can be unified with EIMs as the variational family.
Population Predictive Checks
Aug 08, 2019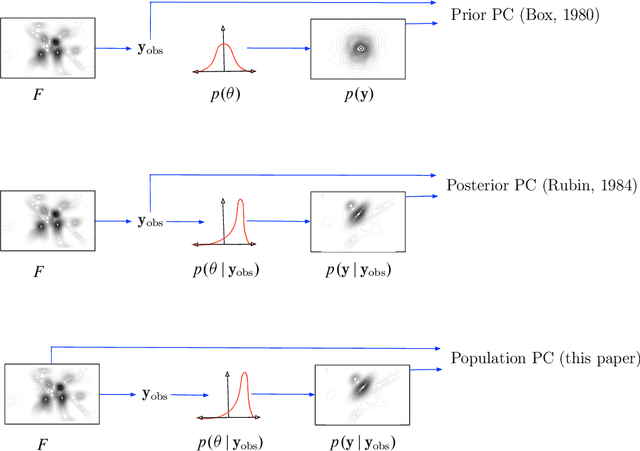
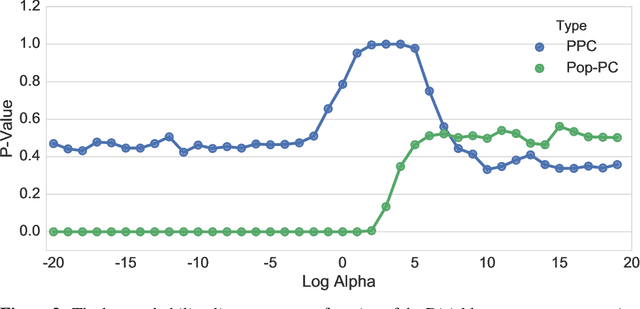
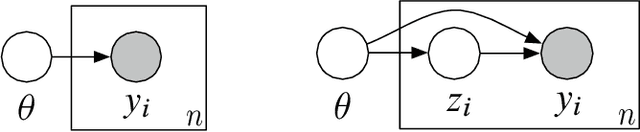
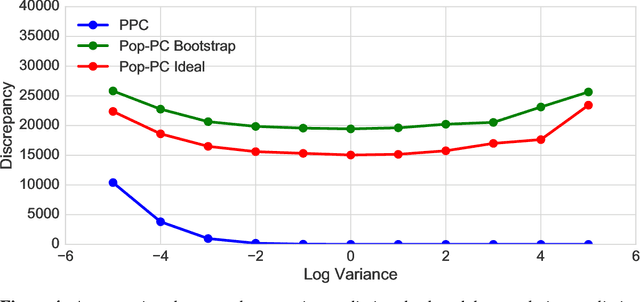
Abstract:Bayesian modeling has become a staple for researchers analyzing data. Thanks to recent developments in approximate posterior inference, modern researchers can easily build, use, and revise complicated Bayesian models for large and rich data. These new abilities, however, bring into focus the problem of model assessment. Researchers need tools to diagnose the fitness of their models, to understand where a model falls short, and to guide its revision. In this paper we develop a new method for Bayesian model checking, the population predictive check (Pop-PC). Pop-PCs are built on posterior predictive checks (PPC), a seminal method that checks a model by assessing the posterior predictive distribution on the observed data. Though powerful, PPCs use the data twice---both to calculate the posterior predictive and to evaluate it---which can lead to overconfident assessments. Pop-PCs, in contrast, compare the posterior predictive distribution to the population distribution of the data. This strategy blends Bayesian modeling with frequentist assessment, leading to a robust check that validates the model on its generalization. Of course the population distribution is not usually available; thus we use tools like the bootstrap and cross validation to estimate the Pop-PC. Further, we extend Pop-PCs to hierarchical models. We study Pop-PCs on classical regression and a hierarchical model of text. We show that Pop-PCs are robust to overfitting and can be easily deployed on a broad family of models.
Generalized Control Functions via Variational Decoupling
Jul 08, 2019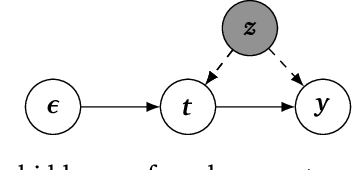
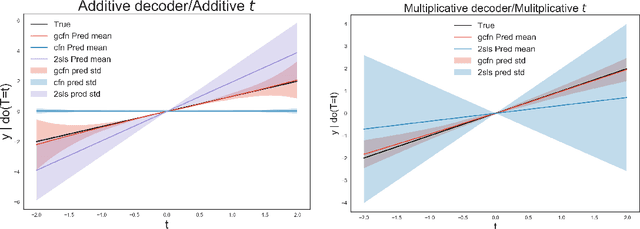
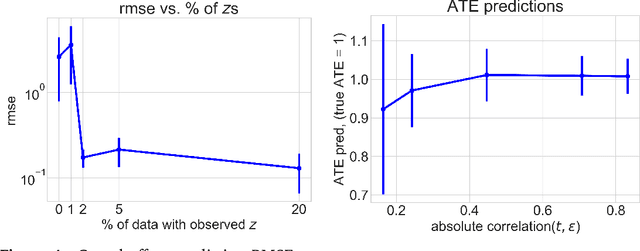

Abstract:Causal estimation relies on separating the variation in the outcome due to the confounders from that due to the treatment. To achieve this separation, practitioners can use external sources of randomness that only influence the treatment called instrumental variables (IVs). Traditional IV-methods rely on structural assumptions that limit the effect that the confounders can have on both outcome and treatment. To relax these assumptions we develop a new estimator called the generalized control-function method (GCFN). GCFN's first stage called variational decoupling (VDE) recovers the residual variation in the treatment given the IV. In the second stage, GCFN regresses the outcome on the treatment and residual variation to compute the causal effect. We evaluate GCFN on simulated data and on recovering the causal effect of slave export on community trust. We show how VDE can help unify IV-estimators and non-IV-estimators.
Reproducibility in Machine Learning for Health
Jul 02, 2019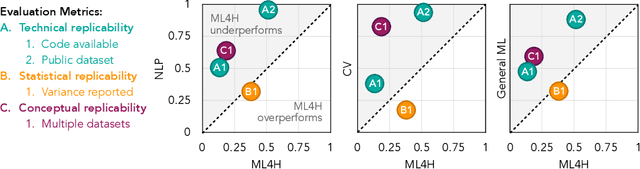
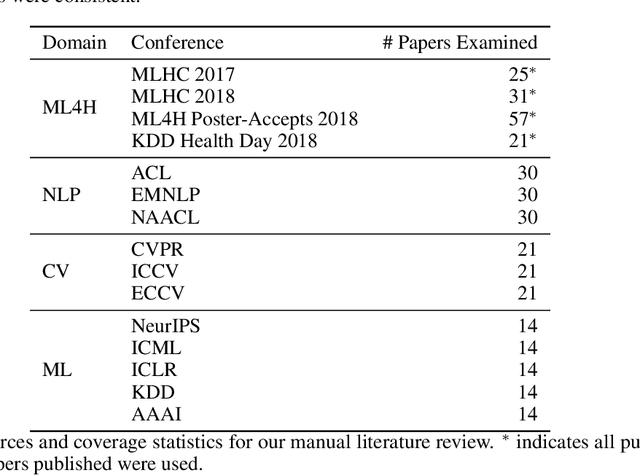
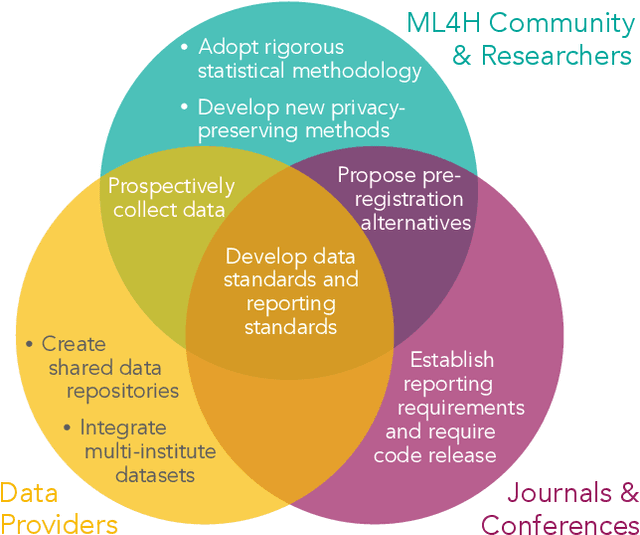
Abstract:Machine learning algorithms designed to characterize, monitor, and intervene on human health (ML4H) are expected to perform safely and reliably when operating at scale, potentially outside strict human supervision. This requirement warrants a stricter attention to issues of reproducibility than other fields of machine learning. In this work, we conduct a systematic evaluation of over 100 recently published ML4H research papers along several dimensions related to reproducibility. We find that the field of ML4H compares poorly to more established machine learning fields, particularly concerning data and code accessibility. Finally, drawing from success in other fields of science, we propose recommendations to data providers, academic publishers, and the ML4H research community in order to promote reproducible research moving forward.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge