Prithwish Chakraborty
G-Net: A Deep Learning Approach to G-computation for Counterfactual Outcome Prediction Under Dynamic Treatment Regimes
Mar 23, 2020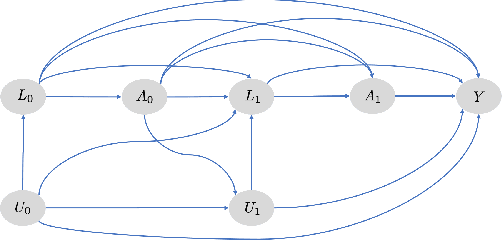
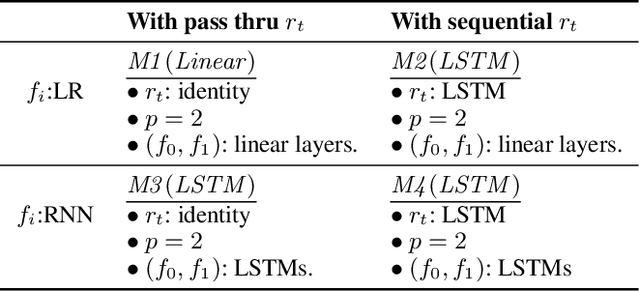

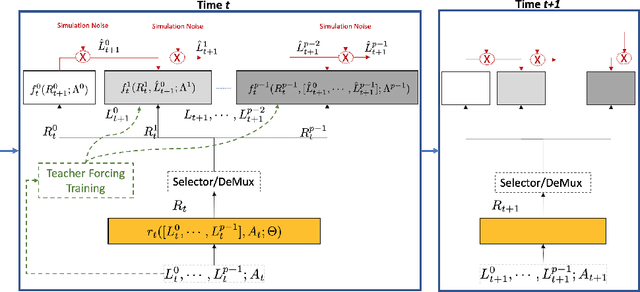
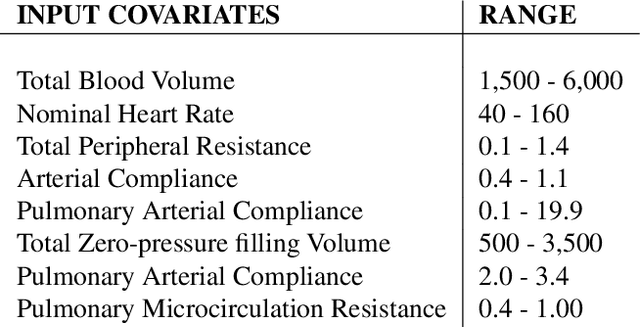
Abstract:Counterfactual prediction is a fundamental task in decision-making. G-computation is a method for estimating expected counterfactual outcomes under dynamic time-varying treatment strategies. Existing G-computation implementations have mostly employed classical regression models with limited capacity to capture complex temporal and nonlinear dependence structures. This paper introduces G-Net, a novel sequential deep learning framework for G-computation that can handle complex time series data while imposing minimal modeling assumptions and provide estimates of individual or population-level time varying treatment effects. We evaluate alternative G-Net implementations using realistically complex temporal simulated data obtained from CVSim, a mechanistic model of the cardiovascular system.
Explicit-Blurred Memory Network for Analyzing Patient Electronic Health Records
Nov 15, 2019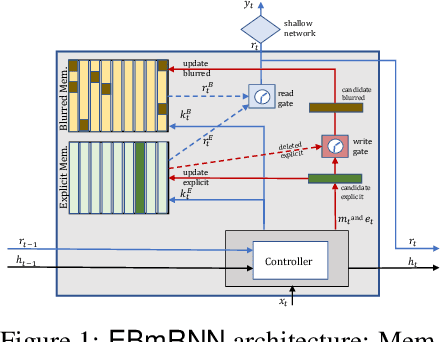
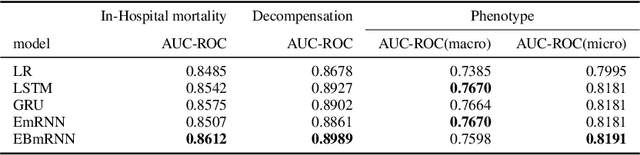
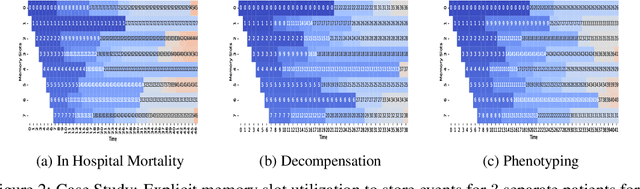
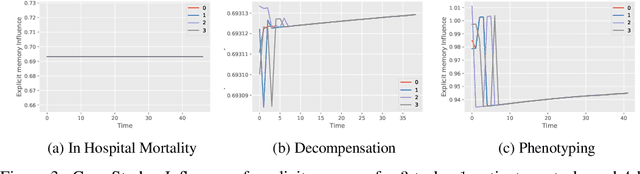
Abstract:In recent years, we have witnessed an increased interest in temporal modeling of patient records from large scale Electronic Health Records (EHR). While simpler RNN models have been used for such problems, memory networks, which in other domains were found to generalize well, are underutilized. Traditional memory networks involve diffused and non-linear operations where influence of past events on outputs are not readily quantifiable. We posit that this lack of interpretability makes such networks not applicable for EHR analysis. While networks with explicit memory have been proposed recently, the discontinuities imposed by the discrete operations make such networks harder to train and require more supervision. The problem is further exacerbated in the limited data setting of EHR studies. In this paper, we propose a novel memory architecture that is more interpretable than traditional memory networks while being easier to train than explicit memory banks. Inspired by well-known models of human cognition, we propose partitioning the external memory space into (a) a primary explicit memory block to store exact replicas of recent events to support interpretations, followed by (b) a secondary blurred memory block that accumulates salient aspects of past events dropped from the explicit block as higher level abstractions and allow training with less supervision by stabilize the gradients. We apply the model for 3 learning problems on ICU records from the MIMIC III database spanning millions of data points. Our model performs comparably to the state-of the art while also, crucially, enabling ready interpretation of the results.
A Novel Data-Driven Framework for Risk Characterization and Prediction from Electronic Medical Records: A Case Study of Renal Failure
Nov 29, 2017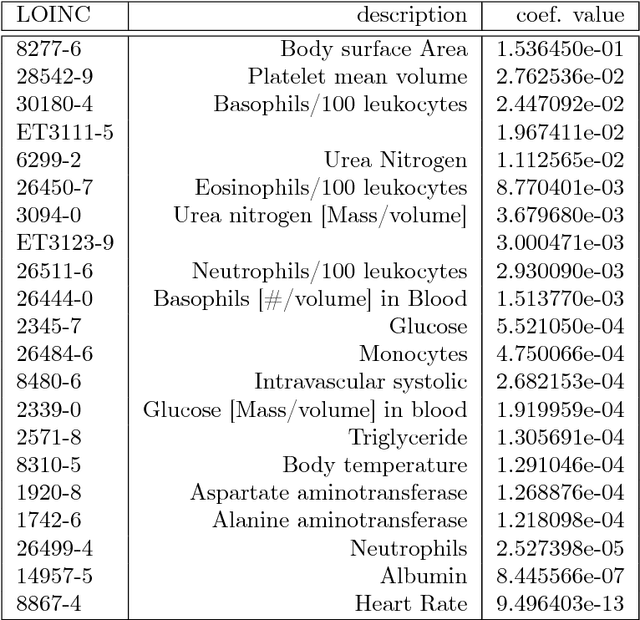
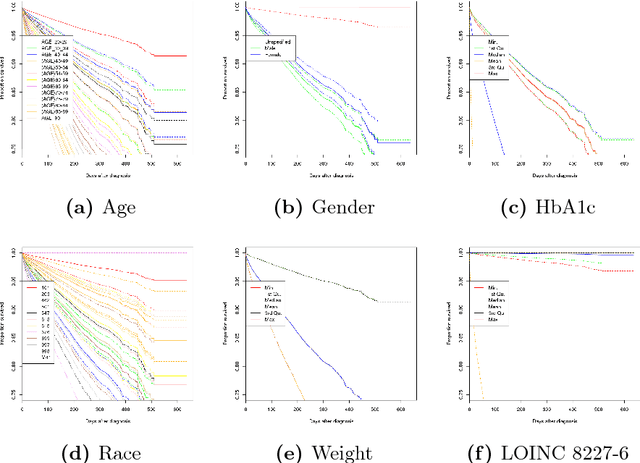
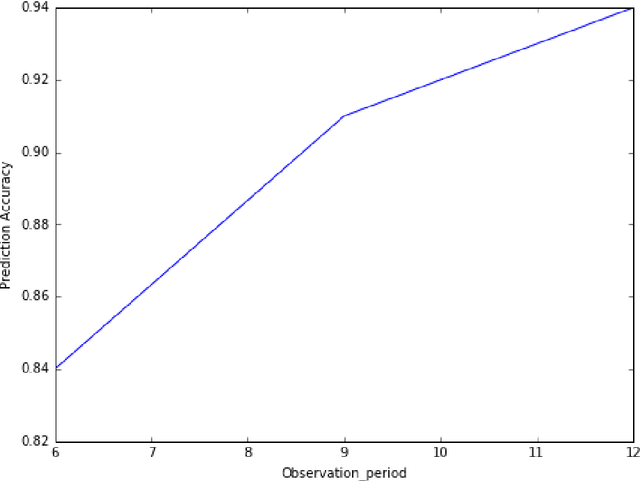
Abstract:Electronic medical records (EMR) contain longitudinal information about patients that can be used to analyze outcomes. Typically, studies on EMR data have worked with established variables that have already been acknowledged to be associated with certain outcomes. However, EMR data may also contain hitherto unrecognized factors for risk association and prediction of outcomes for a disease. In this paper, we present a scalable data-driven framework to analyze EMR data corpus in a disease agnostic way that systematically uncovers important factors influencing outcomes in patients, as supported by data and without expert guidance. We validate the importance of such factors by using the framework to predict for the relevant outcomes. Specifically, we analyze EMR data covering approximately 47 million unique patients to characterize renal failure (RF) among type 2 diabetic (T2DM) patients. We propose a specialized L1 regularized Cox Proportional Hazards (CoxPH) survival model to identify the important factors from those available from patient encounter history. To validate the identified factors, we use a specialized generalized linear model (GLM) to predict the probability of renal failure for individual patients within a specified time window. Our experiments indicate that the factors identified via our data-driven method overlap with the patient characteristics recognized by experts. Our approach allows for scalable, repeatable and efficient utilization of data available in EMRs, confirms prior medical knowledge and can generate new hypothesis without expert supervision.
Guided Deep List: Automating the Generation of Epidemiological Line Lists from Open Sources
Feb 22, 2017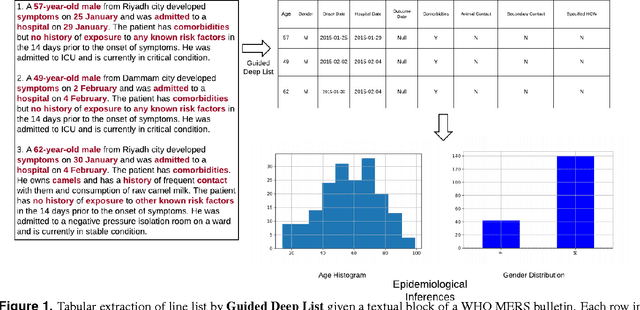
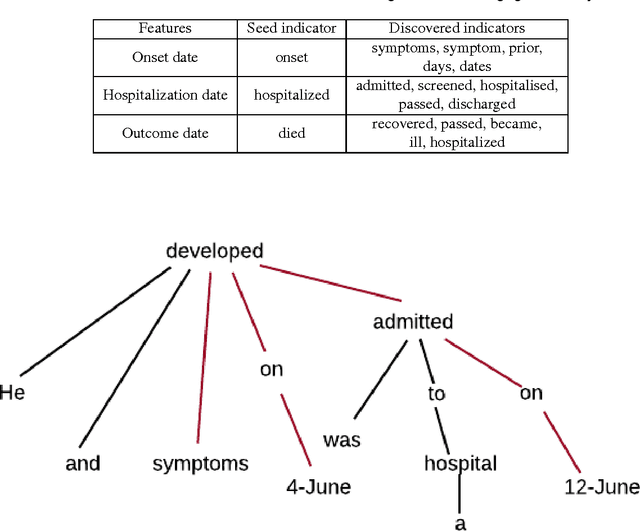
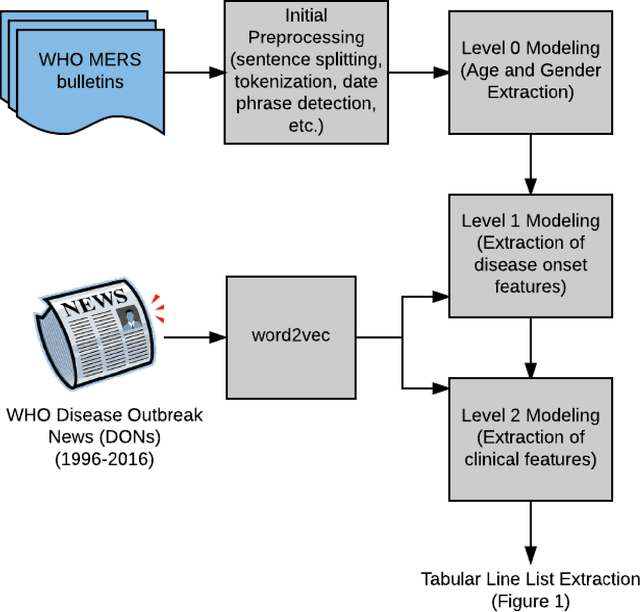
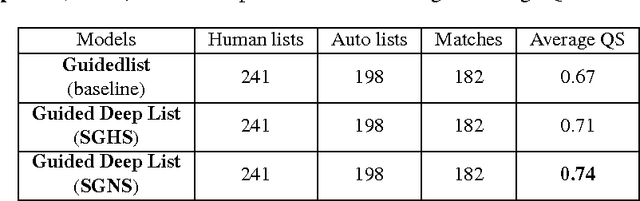
Abstract:Real-time monitoring and responses to emerging public health threats rely on the availability of timely surveillance data. During the early stages of an epidemic, the ready availability of line lists with detailed tabular information about laboratory-confirmed cases can assist epidemiologists in making reliable inferences and forecasts. Such inferences are crucial to understand the epidemiology of a specific disease early enough to stop or control the outbreak. However, construction of such line lists requires considerable human supervision and therefore, difficult to generate in real-time. In this paper, we motivate Guided Deep List, the first tool for building automated line lists (in near real-time) from open source reports of emerging disease outbreaks. Specifically, we focus on deriving epidemiological characteristics of an emerging disease and the affected population from reports of illness. Guided Deep List uses distributed vector representations (ala word2vec) to discover a set of indicators for each line list feature. This discovery of indicators is followed by the use of dependency parsing based techniques for final extraction in tabular form. We evaluate the performance of Guided Deep List against a human annotated line list provided by HealthMap corresponding to MERS outbreaks in Saudi Arabia. We demonstrate that Guided Deep List extracts line list features with increased accuracy compared to a baseline method. We further show how these automatically extracted line list features can be used for making epidemiological inferences, such as inferring demographics and symptoms-to-hospitalization period of affected individuals.
Characterizing Diseases from Unstructured Text: A Vocabulary Driven Word2vec Approach
Jun 03, 2016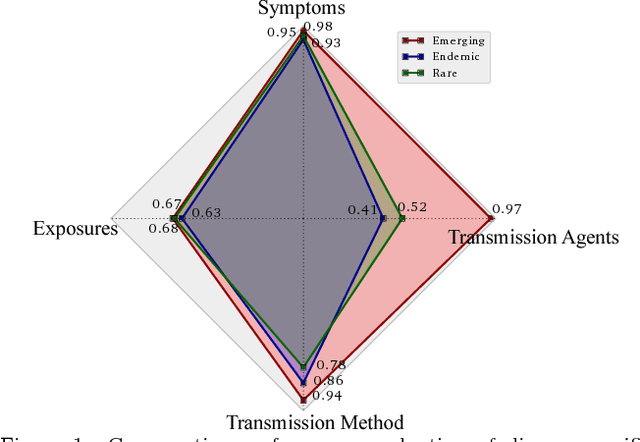
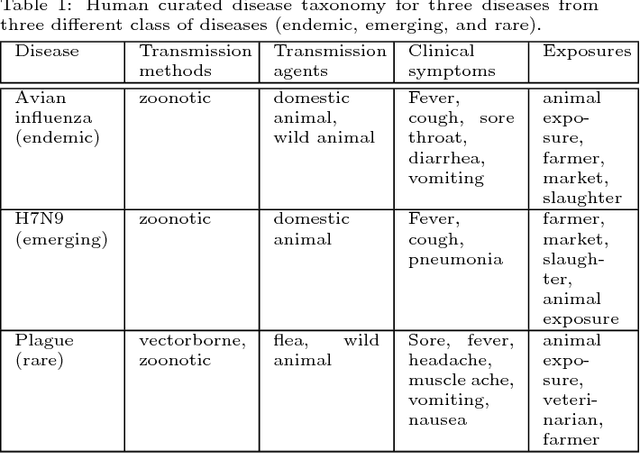
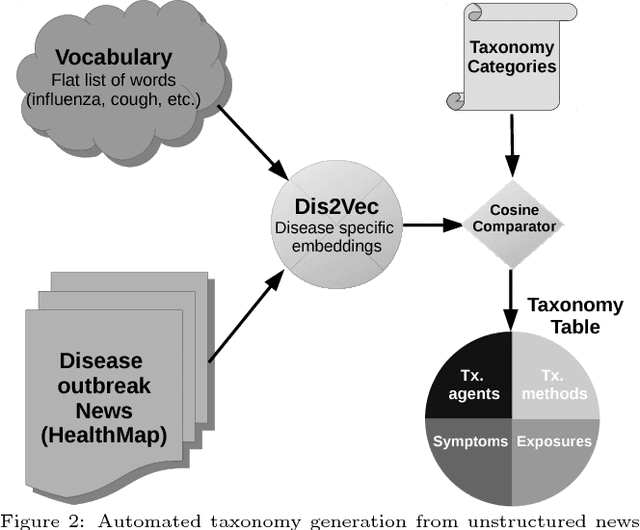
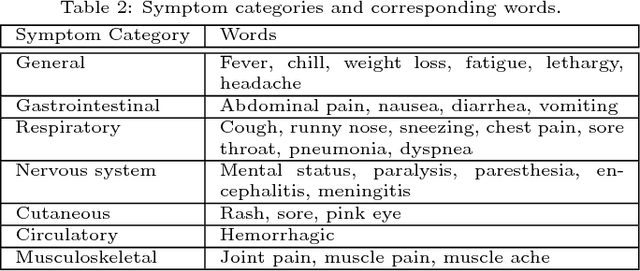
Abstract:Traditional disease surveillance can be augmented with a wide variety of real-time sources such as, news and social media. However, these sources are in general unstructured and, construction of surveillance tools such as taxonomical correlations and trace mapping involves considerable human supervision. In this paper, we motivate a disease vocabulary driven word2vec model (Dis2Vec) to model diseases and constituent attributes as word embeddings from the HealthMap news corpus. We use these word embeddings to automatically create disease taxonomies and evaluate our model against corresponding human annotated taxonomies. We compare our model accuracies against several state-of-the art word2vec methods. Our results demonstrate that Dis2Vec outperforms traditional distributed vector representations in its ability to faithfully capture taxonomical attributes across different class of diseases such as endemic, emerging and rare.
Temporal Topic Modeling to Assess Associations between News Trends and Infectious Disease Outbreaks
Jun 01, 2016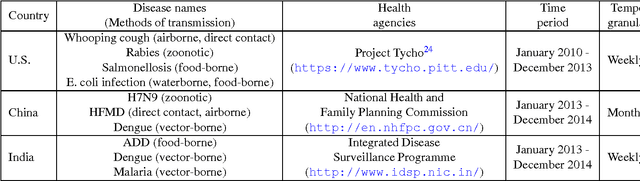
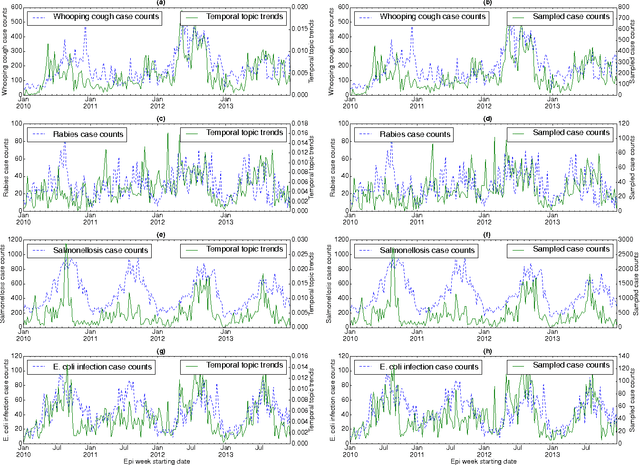

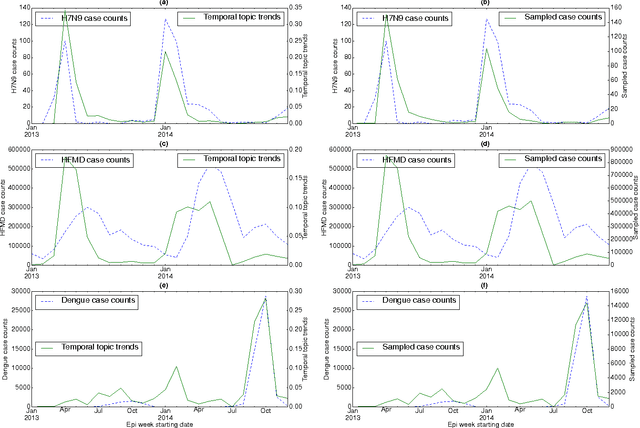
Abstract:In retrospective assessments, internet news reports have been shown to capture early reports of unknown infectious disease transmission prior to official laboratory confirmation. In general, media interest and reporting peaks and wanes during the course of an outbreak. In this study, we quantify the extent to which media interest during infectious disease outbreaks is indicative of trends of reported incidence. We introduce an approach that uses supervised temporal topic models to transform large corpora of news articles into temporal topic trends. The key advantages of this approach include, applicability to a wide range of diseases, and ability to capture disease dynamics - including seasonality, abrupt peaks and troughs. We evaluated the method using data from multiple infectious disease outbreaks reported in the United States of America (U.S.), China and India. We noted that temporal topic trends extracted from disease-related news reports successfully captured the dynamics of multiple outbreaks such as whooping cough in U.S. (2012), dengue outbreaks in India (2013) and China (2014). Our observations also suggest that efficient modeling of temporal topic trends using time-series regression techniques can estimate disease case counts with increased precision before official reports by health organizations.
Hierarchical Quickest Change Detection via Surrogates
Mar 31, 2016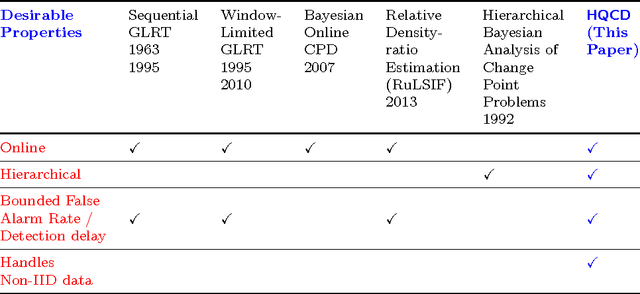
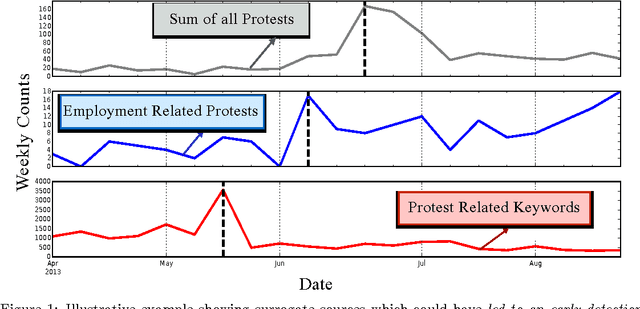
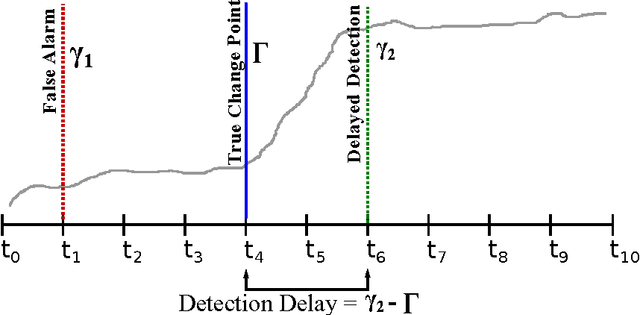
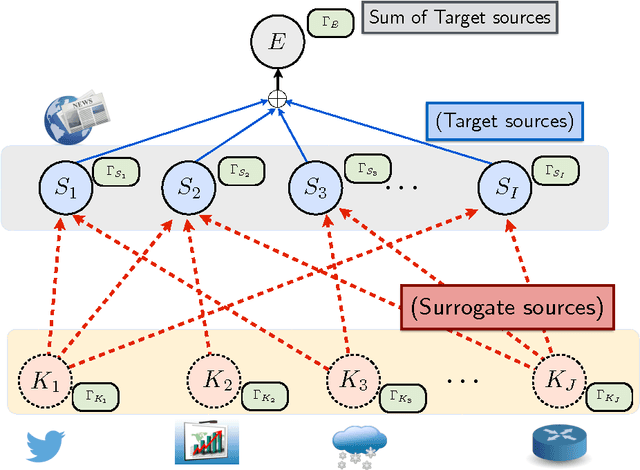
Abstract:Change detection (CD) in time series data is a critical problem as it reveal changes in the underlying generative processes driving the time series. Despite having received significant attention, one important unexplored aspect is how to efficiently utilize additional correlated information to improve the detection and the understanding of changepoints. We propose hierarchical quickest change detection (HQCD), a framework that formalizes the process of incorporating additional correlated sources for early changepoint detection. The core ideas behind HQCD are rooted in the theory of quickest detection and HQCD can be regarded as its novel generalization to a hierarchical setting. The sources are classified into targets and surrogates, and HQCD leverages this structure to systematically assimilate observed data to update changepoint statistics across layers. The decision on actual changepoints are provided by minimizing the delay while still maintaining reliability bounds. In addition, HQCD also uncovers interesting relations between changes at targets from changes across surrogates. We validate HQCD for reliability and performance against several state-of-the-art methods for both synthetic dataset (known changepoints) and several real-life examples (unknown changepoints). Our experiments indicate that we gain significant robustness without loss of detection delay through HQCD. Our real-life experiments also showcase the usefulness of the hierarchical setting by connecting the surrogate sources (such as Twitter chatter) to target sources (such as Employment related protests that ultimately lead to major uprisings).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge