Paul Büschl
VesselTok: Tokenizing Vessel-like 3D Biomedical Graph Representations for Reconstruction and Generation
Mar 19, 2026Abstract:Spatial graphs provide a lightweight and elegant representation of curvilinear anatomical structures such as blood vessels, lung airways, and neuronal networks. Accurately modeling these graphs is crucial in clinical and (bio-)medical research. However, the high spatial resolution of large networks drastically increases their complexity, resulting in significant computational challenges. In this work, we aim to tackle these challenges by proposing VesselTok, a framework that approaches spatially dense graphs from a parametric shape perspective to learn latent representations (tokens). VesselTok leverages centerline points with a pseudo radius to effectively encode tubular geometry. Specifically, we learn a novel latent representation conditioned on centerline points to encode neural implicit representations of vessel-like, tubular structures. We demonstrate VesselTok's performance across diverse anatomies, including lung airways, lung vessels, and brain vessels, highlighting its ability to robustly encode complex topologies. To prove the effectiveness of VesselTok's learnt latent representations, we show that they (i) generalize to unseen anatomies, (ii) support generative modeling of plausible anatomical graphs, and (iii) transfer effectively to downstream inverse problems, such as link prediction.
Whole Brain Vessel Graphs: A Dataset and Benchmark for Graph Learning and Neuroscience
Aug 30, 2021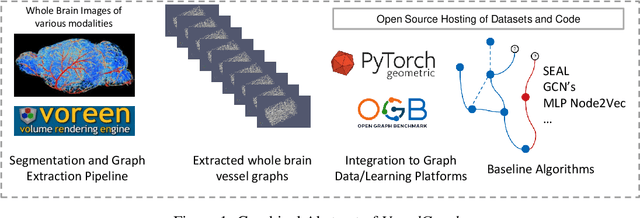
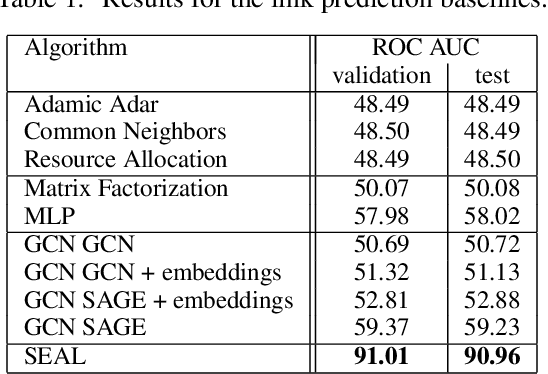
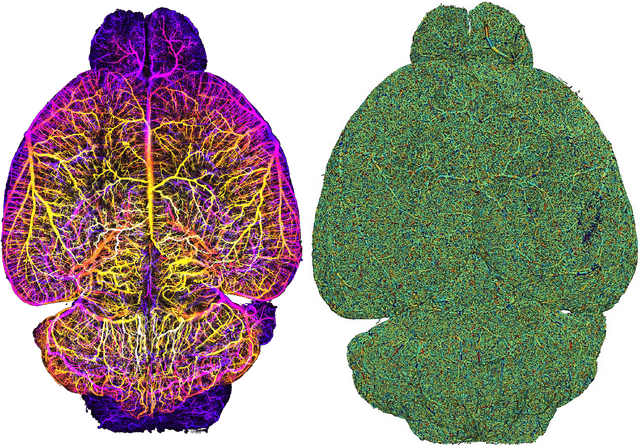
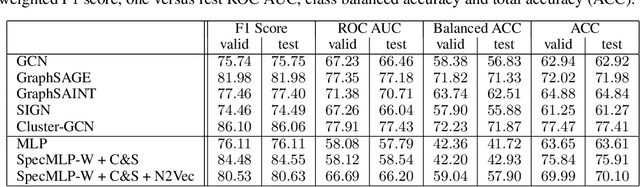
Abstract:Biological neural networks define the brain function and intelligence of humans and other mammals, and form ultra-large, spatial, structured graphs. Their neuronal organization is closely interconnected with the spatial organization of the brain's microvasculature, which supplies oxygen to the neurons and builds a complementary spatial graph. This vasculature (or the vessel structure) plays an important role in neuroscience; for example, the organization of (and changes to) vessel structure can represent early signs of various pathologies, e.g. Alzheimer's disease or stroke. Recently, advances in tissue clearing have enabled whole brain imaging and segmentation of the entirety of the mouse brain's vasculature. Building on these advances in imaging, we are presenting an extendable dataset of whole-brain vessel graphs based on specific imaging protocols. Specifically, we extract vascular graphs using a refined graph extraction scheme leveraging the volume rendering engine Voreen and provide them in an accessible and adaptable form through the OGB and PyTorch Geometric dataloaders. Moreover, we benchmark numerous state-of-the-art graph learning algorithms on the biologically relevant tasks of vessel prediction and vessel classification using the introduced vessel graph dataset. Our work paves a path towards advancing graph learning research into the field of neuroscience. Complementarily, the presented dataset raises challenging graph learning research questions for the machine learning community, in terms of incorporating biological priors into learning algorithms, or in scaling these algorithms to handle sparse,spatial graphs with millions of nodes and edges. All datasets and code are available for download at https://github.com/jocpae/VesselGraph .
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge