Nikola Simidjievski
Tabular Foundation Model for Generative Modelling
May 10, 2026Abstract:Generative modelling is a demanding test of foundation models, because it requires robust, holistic representation learning for a given data modality, rather than optimisation for a supervised prediction target alone. While recent work on tabular foundation models has achieved remarkable progress in predictive modelling, generative tabular foundation models remain underexplored. Existing tabular foundation generators, in particular, have not yet consistently matched strong dataset-specific generators in synthetic data quality. A key reason is their misalignment with the distinctive causal structural prior of heterogeneous tabular data. In this paper, we address this gap by introducing a novel tabular foundation model, \textbf{TabFORGE}, built on pretrained \textbf{Tab}ular \textbf{FO}undational \textbf{R}epresentations for \textbf{GE}neration. TabFORGE is designed to utilise the implicitly learned causal information underlying diverse tabular datasets in a unified latent space induced by a pretrained causality-aware feature encoder. It further decouples latent modelling from decoding through a two-stage design: we first pretrain a score-based diffusion transformer, and then pretrain a denoising-aligned decoder using the denoised latent embeddings. This design elegantly mitigates the distribution shifts in latent embeddings that typically arise between training and inference. We evaluate TabFORGE comprehensively against 22 benchmark methods on 45 real-world datasets. Our results show that TabFORGE effectively learns and leverages generalisable tabular representations, enabling efficient generation of high-quality synthetic tabular data, particularly with strong structural fidelity.
Towards Spatial Transcriptomics-driven Pathology Foundation Models
Feb 15, 2026Abstract:Spatial transcriptomics (ST) provides spatially resolved measurements of gene expression, enabling characterization of the molecular landscape of human tissue beyond histological assessment as well as localized readouts that can be aligned with morphology. Concurrently, the success of multimodal foundation models that integrate vision with complementary modalities suggests that morphomolecular coupling between local expression and morphology can be systematically used to improve histological representations themselves. We introduce Spatial Expression-Aligned Learning (SEAL), a vision-omics self-supervised learning framework that infuses localized molecular information into pathology vision encoders. Rather than training new encoders from scratch, SEAL is designed as a parameter-efficient vision-omics finetuning method that can be flexibly applied to widely used pathology foundation models. We instantiate SEAL by training on over 700,000 paired gene expression spot-tissue region examples spanning tumor and normal samples from 14 organs. Tested across 38 slide-level and 15 patch-level downstream tasks, SEAL provides a drop-in replacement for pathology foundation models that consistently improves performance over widely used vision-only and ST prediction baselines on slide-level molecular status, pathway activity, and treatment response prediction, as well as patch-level gene expression prediction tasks. Additionally, SEAL encoders exhibit robust domain generalization on out-of-distribution evaluations and enable new cross-modal capabilities such as gene-to-image retrieval. Our work proposes a general framework for ST-guided finetuning of pathology foundation models, showing that augmenting existing models with localized molecular supervision is an effective and practical step for improving visual representations and expanding their cross-modal utility.
RO-FIGS: Efficient and Expressive Tree-Based Ensembles for Tabular Data
Apr 09, 2025



Abstract:Tree-based models are often robust to uninformative features and can accurately capture non-smooth, complex decision boundaries. Consequently, they often outperform neural network-based models on tabular datasets at a significantly lower computational cost. Nevertheless, the capability of traditional tree-based ensembles to express complex relationships efficiently is limited by using a single feature to make splits. To improve the efficiency and expressiveness of tree-based methods, we propose Random Oblique Fast Interpretable Greedy-Tree Sums (RO-FIGS). RO-FIGS builds on Fast Interpretable Greedy-Tree Sums, and extends it by learning trees with oblique or multivariate splits, where each split consists of a linear combination learnt from random subsets of features. This helps uncover interactions between features and improves performance. The proposed method is suitable for tabular datasets with both numerical and categorical features. We evaluate RO-FIGS on 22 real-world tabular datasets, demonstrating superior performance and much smaller models over other tree- and neural network-based methods. Additionally, we analyse their splits to reveal valuable insights into feature interactions, enriching the information learnt from SHAP summary plots, and thereby demonstrating the enhanced interpretability of RO-FIGS models. The proposed method is well-suited for applications, where balance between accuracy and interpretability is essential.
How Well Does Your Tabular Generator Learn the Structure of Tabular Data?
Mar 12, 2025Abstract:Heterogeneous tabular data poses unique challenges in generative modelling due to its fundamentally different underlying data structure compared to homogeneous modalities, such as images and text. Although previous research has sought to adapt the successes of generative modelling in homogeneous modalities to the tabular domain, defining an effective generator for tabular data remains an open problem. One major reason is that the evaluation criteria inherited from other modalities often fail to adequately assess whether tabular generative models effectively capture or utilise the unique structural information encoded in tabular data. In this paper, we carefully examine the limitations of the prevailing evaluation framework and introduce $\textbf{TabStruct}$, a novel evaluation benchmark that positions structural fidelity as a core evaluation dimension. Specifically, TabStruct evaluates the alignment of causal structures in real and synthetic data, providing a direct measure of how effectively tabular generative models learn the structure of tabular data. Through extensive experiments using generators from eight categories on seven datasets with expert-validated causal graphical structures, we show that structural fidelity offers a task-independent, domain-agnostic evaluation dimension. Our findings highlight the importance of tabular data structure and offer practical guidance for developing more effective and robust tabular generative models. Code is available at https://github.com/SilenceX12138/TabStruct.
LLM Embeddings for Deep Learning on Tabular Data
Feb 17, 2025Abstract:Tabular deep-learning methods require embedding numerical and categorical input features into high-dimensional spaces before processing them. Existing methods deal with this heterogeneous nature of tabular data by employing separate type-specific encoding approaches. This limits the cross-table transfer potential and the exploitation of pre-trained knowledge. We propose a novel approach that first transforms tabular data into text, and then leverages pre-trained representations from LLMs to encode this data, resulting in a plug-and-play solution to improv ing deep-learning tabular methods. We demonstrate that our approach improves accuracy over competitive models, such as MLP, ResNet and FT-Transformer, by validating on seven classification datasets.
Measuring Cross-Modal Interactions in Multimodal Models
Dec 20, 2024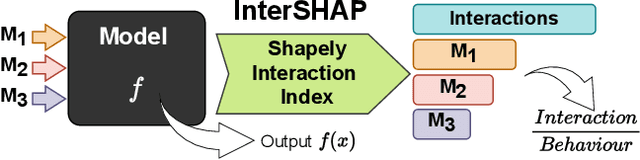
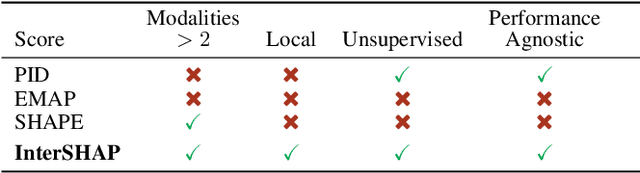
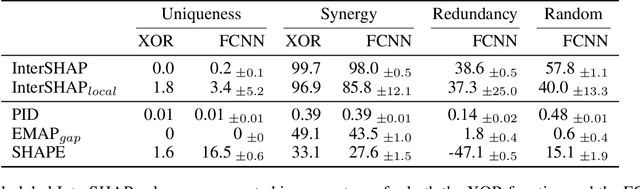
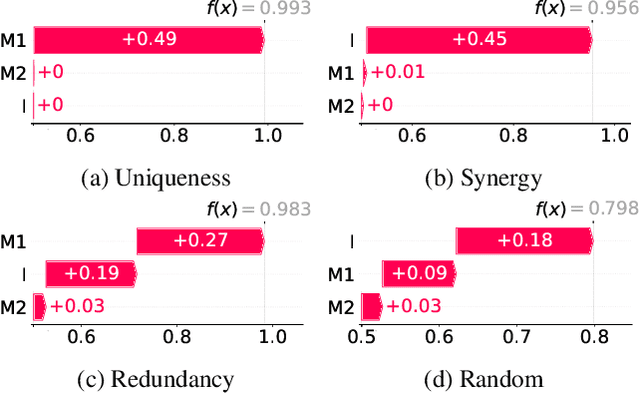
Abstract:Integrating AI in healthcare can greatly improve patient care and system efficiency. However, the lack of explainability in AI systems (XAI) hinders their clinical adoption, especially in multimodal settings that use increasingly complex model architectures. Most existing XAI methods focus on unimodal models, which fail to capture cross-modal interactions crucial for understanding the combined impact of multiple data sources. Existing methods for quantifying cross-modal interactions are limited to two modalities, rely on labelled data, and depend on model performance. This is problematic in healthcare, where XAI must handle multiple data sources and provide individualised explanations. This paper introduces InterSHAP, a cross-modal interaction score that addresses the limitations of existing approaches. InterSHAP uses the Shapley interaction index to precisely separate and quantify the contributions of the individual modalities and their interactions without approximations. By integrating an open-source implementation with the SHAP package, we enhance reproducibility and ease of use. We show that InterSHAP accurately measures the presence of cross-modal interactions, can handle multiple modalities, and provides detailed explanations at a local level for individual samples. Furthermore, we apply InterSHAP to multimodal medical datasets and demonstrate its applicability for individualised explanations.
PATHS: A Hierarchical Transformer for Efficient Whole Slide Image Analysis
Nov 27, 2024Abstract:Computational analysis of whole slide images (WSIs) has seen significant research progress in recent years, with applications ranging across important diagnostic and prognostic tasks such as survival or cancer subtype prediction. Many state-of-the-art models process the entire slide - which may be as large as $150,000 \times 150,000$ pixels - as a bag of many patches, the size of which necessitates computationally cheap feature aggregation methods. However, a large proportion of these patches are uninformative, such as those containing only healthy or adipose tissue, adding significant noise and size to the bag. We propose Pathology Transformer with Hierarchical Selection (PATHS), a novel top-down method for hierarchical weakly supervised representation learning on slide-level tasks in computational pathology. PATHS is inspired by the cross-magnification manner in which a human pathologist examines a slide, recursively filtering patches at each magnification level to a small subset relevant to the diagnosis. Our method overcomes the complications of processing the entire slide, enabling quadratic self-attention and providing a simple interpretable measure of region importance. We apply PATHS to five datasets of The Cancer Genome Atlas (TCGA), and achieve superior performance on slide-level prediction tasks when compared to previous methods, despite processing only a small proportion of the slide.
TabEBM: A Tabular Data Augmentation Method with Distinct Class-Specific Energy-Based Models
Sep 24, 2024



Abstract:Data collection is often difficult in critical fields such as medicine, physics, and chemistry. As a result, classification methods usually perform poorly with these small datasets, leading to weak predictive performance. Increasing the training set with additional synthetic data, similar to data augmentation in images, is commonly believed to improve downstream classification performance. However, current tabular generative methods that learn either the joint distribution $ p(\mathbf{x}, y) $ or the class-conditional distribution $ p(\mathbf{x} \mid y) $ often overfit on small datasets, resulting in poor-quality synthetic data, usually worsening classification performance compared to using real data alone. To solve these challenges, we introduce TabEBM, a novel class-conditional generative method using Energy-Based Models (EBMs). Unlike existing methods that use a shared model to approximate all class-conditional densities, our key innovation is to create distinct EBM generative models for each class, each modelling its class-specific data distribution individually. This approach creates robust energy landscapes, even in ambiguous class distributions. Our experiments show that TabEBM generates synthetic data with higher quality and better statistical fidelity than existing methods. When used for data augmentation, our synthetic data consistently improves the classification performance across diverse datasets of various sizes, especially small ones.
TabMDA: Tabular Manifold Data Augmentation for Any Classifier using Transformers with In-context Subsetting
Jun 03, 2024



Abstract:Tabular data is prevalent in many critical domains, yet it is often challenging to acquire in large quantities. This scarcity usually results in poor performance of machine learning models on such data. Data augmentation, a common strategy for performance improvement in vision and language tasks, typically underperforms for tabular data due to the lack of explicit symmetries in the input space. To overcome this challenge, we introduce TabMDA, a novel method for manifold data augmentation on tabular data. This method utilises a pre-trained in-context model, such as TabPFN, to map the data into a manifold space. TabMDA performs label-invariant transformations by encoding the data multiple times with varied contexts. This process explores the manifold of the underlying in-context models, thereby enlarging the training dataset. TabMDA is a training-free method, making it applicable to any classifier. We evaluate TabMDA on five standard classifiers and observe significant performance improvements across various tabular datasets. Our results demonstrate that TabMDA provides an effective way to leverage information from pre-trained in-context models to enhance the performance of downstream classifiers.
MM-Lego: Modular Biomedical Multimodal Models with Minimal Fine-Tuning
May 30, 2024



Abstract:Learning holistic computational representations in physical, chemical or biological systems requires the ability to process information from different distributions and modalities within the same model. Thus, the demand for multimodal machine learning models has sharply risen for modalities that go beyond vision and language, such as sequences, graphs, time series, or tabular data. While there are many available multimodal fusion and alignment approaches, most of them require end-to-end training, scale quadratically with the number of modalities, cannot handle cases of high modality imbalance in the training set, or are highly topology-specific, making them too restrictive for many biomedical learning tasks. This paper presents Multimodal Lego (MM-Lego), a modular and general-purpose fusion and model merging framework to turn any set of encoders into a competitive multimodal model with no or minimal fine-tuning. We achieve this by introducing a wrapper for unimodal encoders that enforces lightweight dimensionality assumptions between modalities and harmonises their representations by learning features in the frequency domain to enable model merging with little signal interference. We show that MM-Lego 1) can be used as a model merging method which achieves competitive performance with end-to-end fusion models without any fine-tuning, 2) can operate on any unimodal encoder, and 3) is a model fusion method that, with minimal fine-tuning, achieves state-of-the-art results on six benchmarked multimodal biomedical tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge