Mingyuan Xu
Learning Sequential Decisions from Multiple Sources via Group-Robust Markov Decision Processes
Feb 02, 2026Abstract:We often collect data from multiple sites (e.g., hospitals) that share common structure but also exhibit heterogeneity. This paper aims to learn robust sequential decision-making policies from such offline, multi-site datasets. To model cross-site uncertainty, we study distributionally robust MDPs with a group-linear structure: all sites share a common feature map, and both the transition kernels and expected reward functions are linear in these shared features. We introduce feature-wise (d-rectangular) uncertainty sets, which preserve tractable robust Bellman recursions while maintaining key cross-site structure. Building on this, we then develop an offline algorithm based on pessimistic value iteration that includes: (i) per-site ridge regression for Bellman targets, (ii) feature-wise worst-case (row-wise minimization) aggregation, and (iii) a data-dependent pessimism penalty computed from the diagonals of the inverse design matrices. We further propose a cluster-level extension that pools similar sites to improve sample efficiency, guided by prior knowledge of site similarity. Under a robust partial coverage assumption, we prove a suboptimality bound for the resulting policy. Overall, our framework addresses multi-site learning with heterogeneous data sources and provides a principled approach to robust planning without relying on strong state-action rectangularity assumptions.
A Judge-Aware Ranking Framework for Evaluating Large Language Models without Ground Truth
Jan 29, 2026Abstract:Evaluating large language models (LLMs) on open-ended tasks without ground-truth labels is increasingly done via the LLM-as-a-judge paradigm. A critical but under-modeled issue is that judge LLMs differ substantially in reliability; treating all judges equally can yield biased leaderboards and misleading uncertainty estimates. More data can make evaluation more confidently wrong under misspecified aggregation. We propose a judge-aware ranking framework that extends the Bradley-Terry-Luce model by introducing judge-specific discrimination parameters, jointly estimating latent model quality and judge reliability from pairwise comparisons without reference labels. We establish identifiability up to natural normalizations and prove consistency and asymptotic normality of the maximum likelihood estimator, enabling confidence intervals for score differences and rank comparisons. Across multiple public benchmarks and a newly collected dataset, our method improves agreement with human preferences, achieves higher data efficiency than unweighted baselines, and produces calibrated uncertainty quantification for LLM rankings.
Optimizing Medical Image Segmentation with Advanced Decoder Design
Oct 05, 2024



Abstract:U-Net is widely used in medical image segmentation due to its simple and flexible architecture design. To address the challenges of scale and complexity in medical tasks, several variants of U-Net have been proposed. In particular, methods based on Vision Transformer (ViT), represented by Swin UNETR, have gained widespread attention in recent years. However, these improvements often focus on the encoder, overlooking the crucial role of the decoder in optimizing segmentation details. This design imbalance limits the potential for further enhancing segmentation performance. To address this issue, we analyze the roles of various decoder components, including upsampling method, skip connection, and feature extraction module, as well as the shortcomings of existing methods. Consequently, we propose Swin DER (i.e., Swin UNETR Decoder Enhanced and Refined) by specifically optimizing the design of these three components. Swin DER performs upsampling using learnable interpolation algorithm called offset coordinate neighborhood weighted up sampling (Onsampling) and replaces traditional skip connection with spatial-channel parallel attention gate (SCP AG). Additionally, Swin DER introduces deformable convolution along with attention mechanism in the feature extraction module of the decoder. Our model design achieves excellent results, surpassing other state-of-the-art methods on both the Synapse and the MSD brain tumor segmentation task. Code is available at: https://github.com/WillBeanYang/Swin-DER
Node-based Knowledge Graph Contrastive Learning for Medical Relationship Prediction
Oct 16, 2023



Abstract:The embedding of Biomedical Knowledge Graphs (BKGs) generates robust representations, valuable for a variety of artificial intelligence applications, including predicting drug combinations and reasoning disease-drug relationships. Meanwhile, contrastive learning (CL) is widely employed to enhance the distinctiveness of these representations. However, constructing suitable contrastive pairs for CL, especially within Knowledge Graphs (KGs), has been challenging. In this paper, we proposed a novel node-based contrastive learning method for knowledge graph embedding, NC-KGE. NC-KGE enhances knowledge extraction in embeddings and speeds up training convergence by constructing appropriate contrastive node pairs on KGs. This scheme can be easily integrated with other knowledge graph embedding (KGE) methods. For downstream task such as biochemical relationship prediction, we have incorporated a relation-aware attention mechanism into NC-KGE, focusing on the semantic relationships and node interactions. Extensive experiments show that NC-KGE performs competitively with state-of-the-art models on public datasets like FB15k-237 and WN18RR. Particularly in biomedical relationship prediction tasks, NC-KGE outperforms all baselines on datasets such as PharmKG8k-28, DRKG17k-21, and BioKG72k-14, especially in predicting drug combination relationships. We release our code at https://github.com/zhi520/NC-KGE.
More complex encoder is not all you need
Sep 21, 2023



Abstract:U-Net and its variants have been widely used in medical image segmentation. However, most current U-Net variants confine their improvement strategies to building more complex encoder, while leaving the decoder unchanged or adopting a simple symmetric structure. These approaches overlook the true functionality of the decoder: receiving low-resolution feature maps from the encoder and restoring feature map resolution and lost information through upsampling. As a result, the decoder, especially its upsampling component, plays a crucial role in enhancing segmentation outcomes. However, in 3D medical image segmentation, the commonly used transposed convolution can result in visual artifacts. This issue stems from the absence of direct relationship between adjacent pixels in the output feature map. Furthermore, plain encoder has already possessed sufficient feature extraction capability because downsampling operation leads to the gradual expansion of the receptive field, but the loss of information during downsampling process is unignorable. To address the gap in relevant research, we extend our focus beyond the encoder and introduce neU-Net (i.e., not complex encoder U-Net), which incorporates a novel Sub-pixel Convolution for upsampling to construct a powerful decoder. Additionally, we introduce multi-scale wavelet inputs module on the encoder side to provide additional information. Our model design achieves excellent results, surpassing other state-of-the-art methods on both the Synapse and ACDC datasets.
Neural Network Based in Silico Simulation of Combustion Reactions
Nov 27, 2019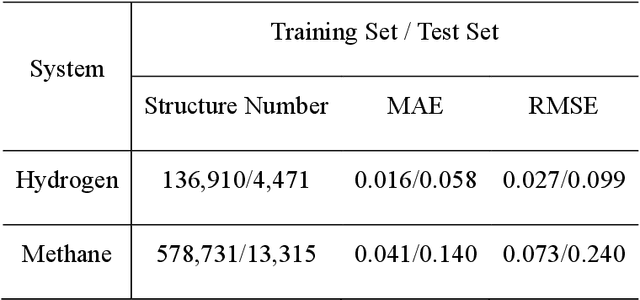
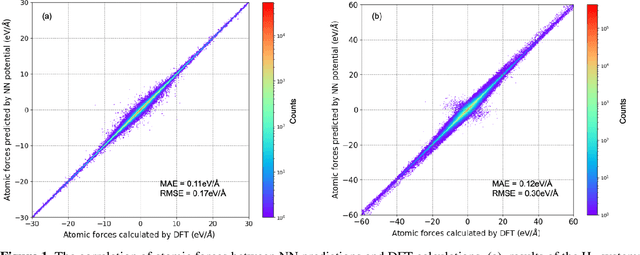
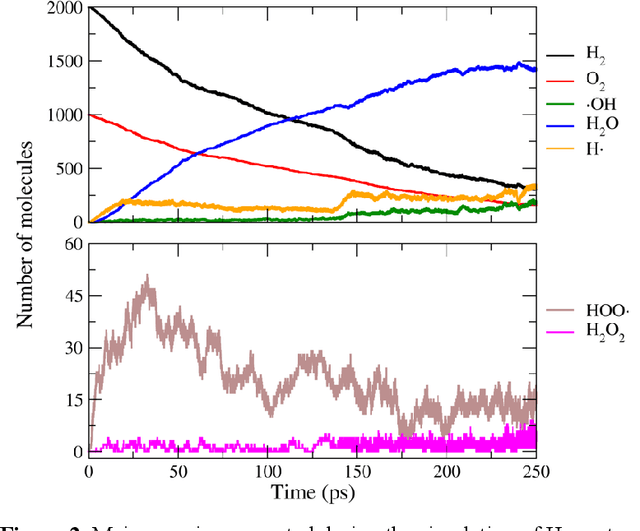
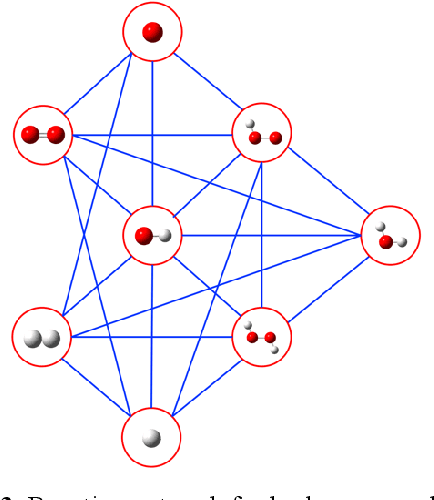
Abstract:Understanding and prediction of the chemical reactions are fundamental demanding in the study of many complex chemical systems. Reactive molecular dynamics (MD) simulation has been widely used for this purpose as it can offer atomic details and can help us better interpret chemical reaction mechanisms. In this study, two reference datasets were constructed and corresponding neural network (NN) potentials were trained based on them. For given large-scale reaction systems, the NN potentials can predict the potential energy and atomic forces of DFT precision, while it is orders of magnitude faster than the conventional DFT calculation. With these two models, reactive MD simulations were performed to explore the combustion mechanisms of hydrogen and methane. Benefit from the high efficiency of the NN model, nanosecond MD trajectories for large-scale systems containing hundreds of atoms were produced and detailed combustion mechanism was obtained. Through further development, the algorithms in this study can be used to explore and discovery reaction mechanisms of many complex reaction systems, such as combustion, synthesis, and heterogeneous catalysis without any predefined reaction coordinates and elementary reaction steps.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge