Max A. Viergever
Standardized Assessment of Automatic Segmentation of White Matter Hyperintensities and Results of the WMH Segmentation Challenge
Apr 01, 2019
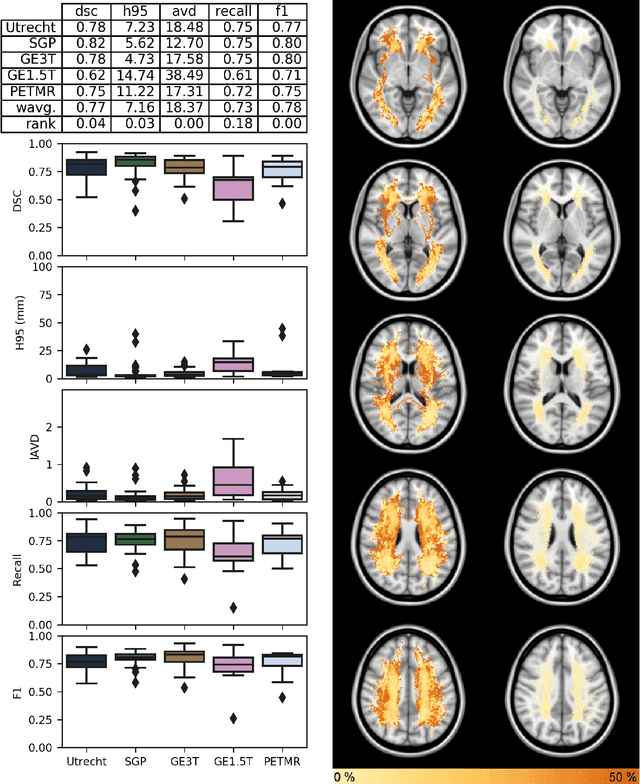
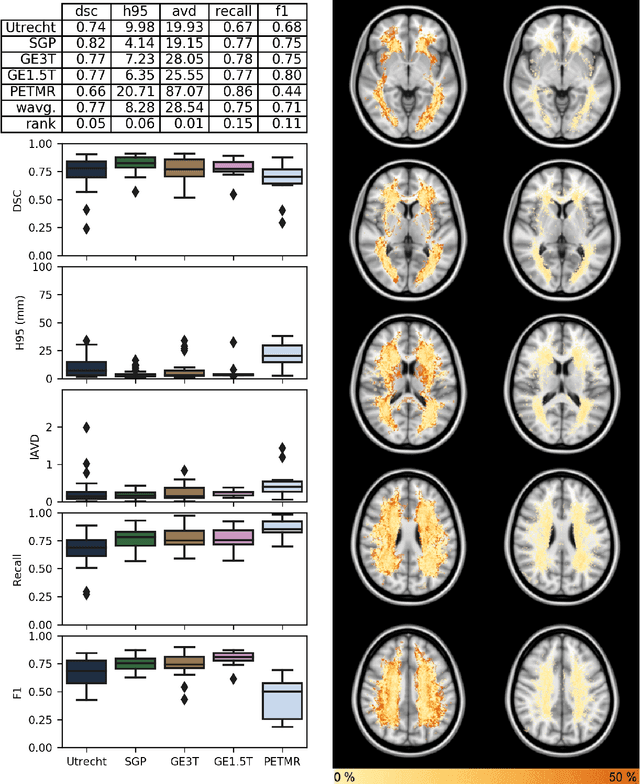
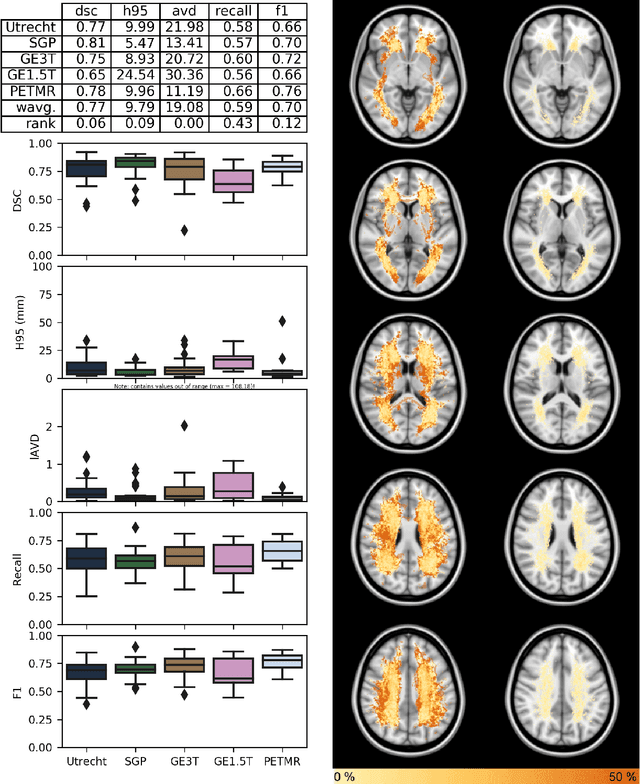
Abstract:Quantification of cerebral white matter hyperintensities (WMH) of presumed vascular origin is of key importance in many neurological research studies. Currently, measurements are often still obtained from manual segmentations on brain MR images, which is a laborious procedure. Automatic WMH segmentation methods exist, but a standardized comparison of the performance of such methods is lacking. We organized a scientific challenge, in which developers could evaluate their method on a standardized multi-center/-scanner image dataset, giving an objective comparison: the WMH Segmentation Challenge (https://wmh.isi.uu.nl/). Sixty T1+FLAIR images from three MR scanners were released with manual WMH segmentations for training. A test set of 110 images from five MR scanners was used for evaluation. Segmentation methods had to be containerized and submitted to the challenge organizers. Five evaluation metrics were used to rank the methods: (1) Dice similarity coefficient, (2) modified Hausdorff distance (95th percentile), (3) absolute log-transformed volume difference, (4) sensitivity for detecting individual lesions, and (5) F1-score for individual lesions. Additionally, methods were ranked on their inter-scanner robustness. Twenty participants submitted their method for evaluation. This paper provides a detailed analysis of the results. In brief, there is a cluster of four methods that rank significantly better than the other methods, with one clear winner. The inter-scanner robustness ranking shows that not all methods generalize to unseen scanners. The challenge remains open for future submissions and provides a public platform for method evaluation.
Coronary Artery Centerline Extraction in Cardiac CT Angiography Using a CNN-Based Orientation Classifier
Oct 24, 2018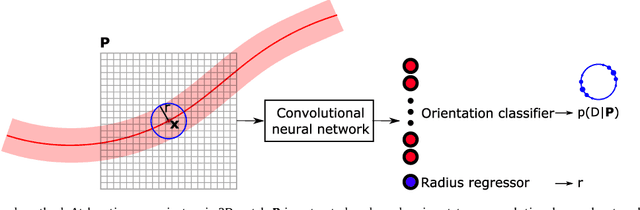
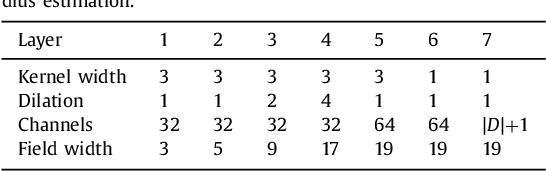
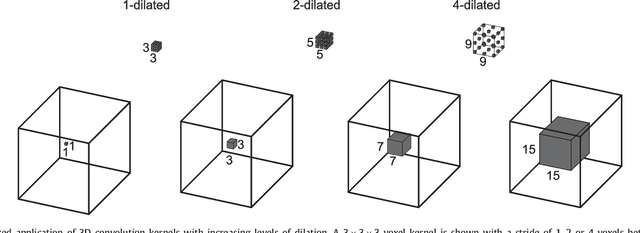
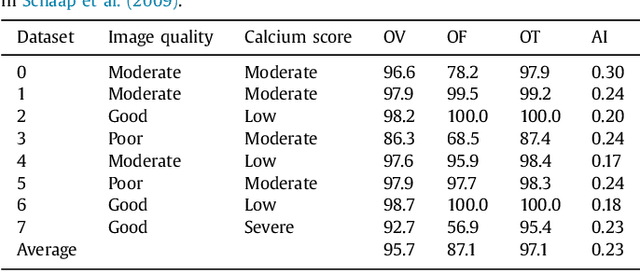
Abstract:Coronary artery centerline extraction in cardiac CT angiography (CCTA) images is a prerequisite for evaluation of stenoses and atherosclerotic plaque. We propose an algorithm that extracts coronary artery centerlines in CCTA using a convolutional neural network (CNN). A 3D dilated CNN is trained to predict the most likely direction and radius of an artery at any given point in a CCTA image based on a local image patch. Starting from a single seed point placed manually or automatically anywhere in a coronary artery, a tracker follows the vessel centerline in two directions using the predictions of the CNN. Tracking is terminated when no direction can be identified with high certainty. The CNN was trained using 32 manually annotated centerlines in a training set consisting of 8 CCTA images provided in the MICCAI 2008 Coronary Artery Tracking Challenge (CAT08). Evaluation using 24 test images of the CAT08 challenge showed that extracted centerlines had an average overlap of 93.7% with 96 manually annotated reference centerlines. Extracted centerline points were highly accurate, with an average distance of 0.21 mm to reference centerline points. In a second test set consisting of 50 CCTA scans, 5,448 markers in the coronary arteries were used as seed points to extract single centerlines. This showed strong correspondence between extracted centerlines and manually placed markers. In a third test set containing 36 CCTA scans, fully automatic seeding and centerline extraction led to extraction of on average 92% of clinically relevant coronary artery segments. The proposed method is able to accurately and efficiently determine the direction and radius of coronary arteries. The method can be trained with limited training data, and once trained allows fast automatic or interactive extraction of coronary artery trees from CCTA images.
Automatic, fast and robust characterization of noise distributions for diffusion MRI
Oct 02, 2018
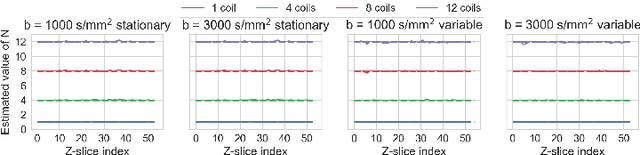
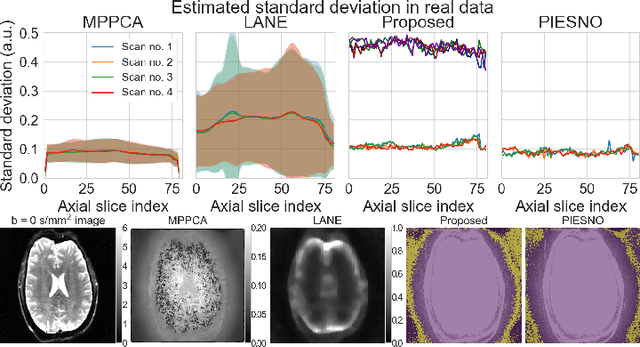
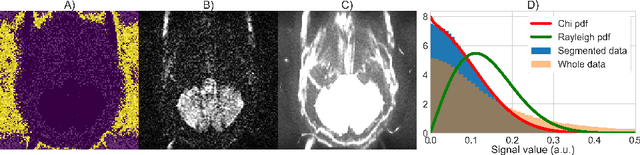
Abstract:Knowledge of the noise distribution in magnitude diffusion MRI images is the centerpiece to quantify uncertainties arising from the acquisition process. The use of parallel imaging methods, the number of receiver coils and imaging filters applied by the scanner, amongst other factors, dictate the resulting signal distribution. Accurate estimation beyond textbook Rician or noncentral chi distributions often requires information about the acquisition process (e.g. coils sensitivity maps or reconstruction coefficients), which is not usually available. We introduce a new method where a change of variable naturally gives rise to a particular form of the gamma distribution for background signals. The first moments and maximum likelihood estimators of this gamma distribution explicitly depend on the number of coils, making it possible to estimate all unknown parameters using only the magnitude data. A rejection step is used to make the method automatic and robust to artifacts. Experiments on synthetic datasets show that the proposed method can reliably estimate both the degrees of freedom and the standard deviation. The worst case errors range from below 2% (spatially uniform noise) to approximately 10% (spatially variable noise). Repeated acquisitions of in vivo datasets show that the estimated parameters are stable and have lower variances than compared methods.
* v2: added publisher DOI statement, fixed text typo in appendix A2
A Deep Learning Framework for Unsupervised Affine and Deformable Image Registration
Sep 17, 2018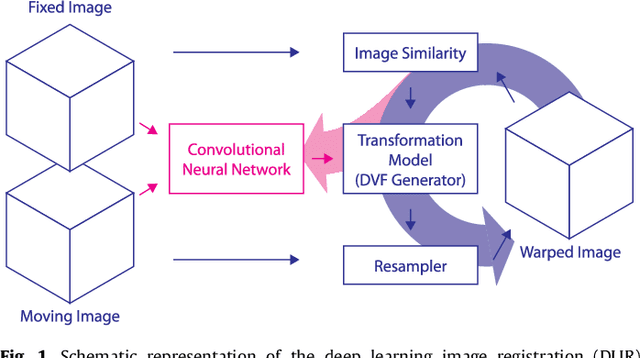
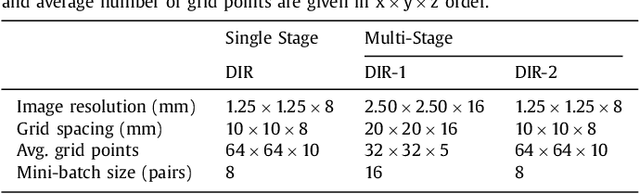
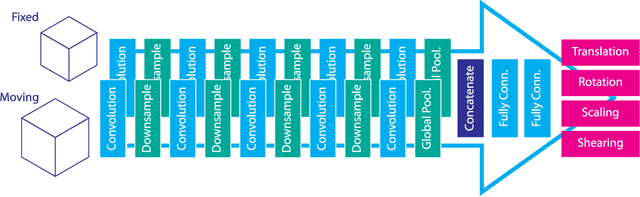
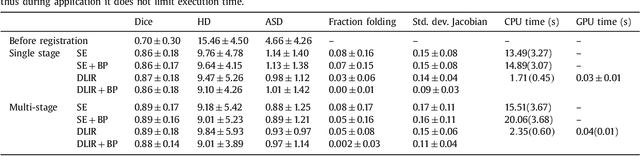
Abstract:Image registration, the process of aligning two or more images, is the core technique of many (semi-)automatic medical image analysis tasks. Recent studies have shown that deep learning methods, notably convolutional neural networks (ConvNets), can be used for image registration. Thus far training of ConvNets for registration was supervised using predefined example registrations. However, obtaining example registrations is not trivial. To circumvent the need for predefined examples, and thereby to increase convenience of training ConvNets for image registration, we propose the Deep Learning Image Registration (DLIR) framework for \textit{unsupervised} affine and deformable image registration. In the DLIR framework ConvNets are trained for image registration by exploiting image similarity analogous to conventional intensity-based image registration. After a ConvNet has been trained with the DLIR framework, it can be used to register pairs of unseen images in one shot. We propose flexible ConvNets designs for affine image registration and for deformable image registration. By stacking multiple of these ConvNets into a larger architecture, we are able to perform coarse-to-fine image registration. We show for registration of cardiac cine MRI and registration of chest CT that performance of the DLIR framework is comparable to conventional image registration while being several orders of magnitude faster.
A Recurrent CNN for Automatic Detection and Classification of Coronary Artery Plaque and Stenosis in Coronary CT Angiography
Aug 20, 2018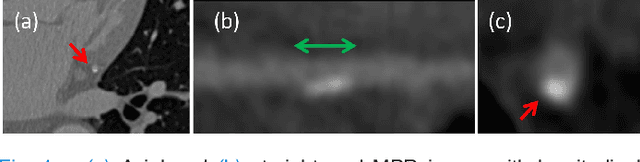
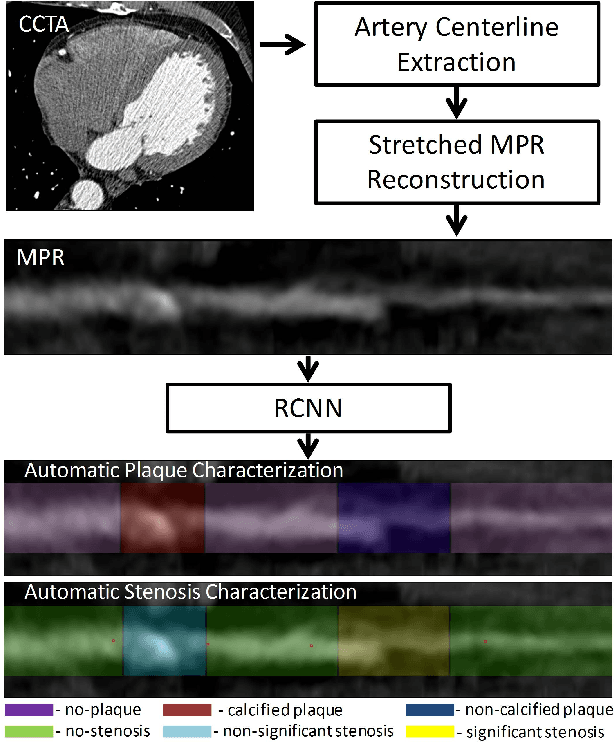
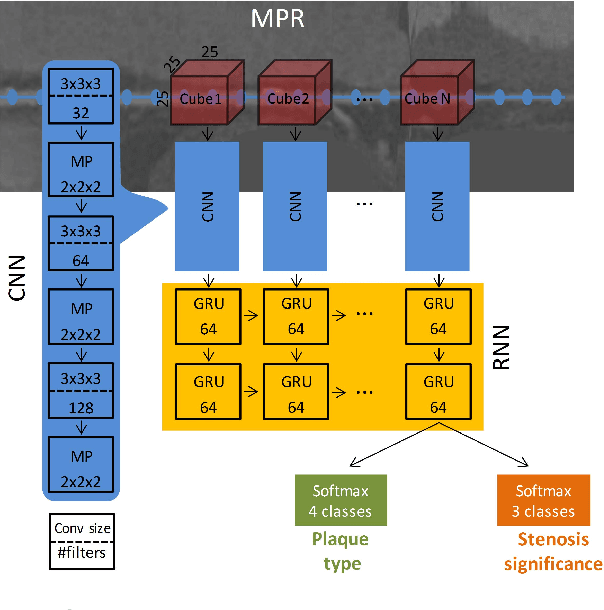
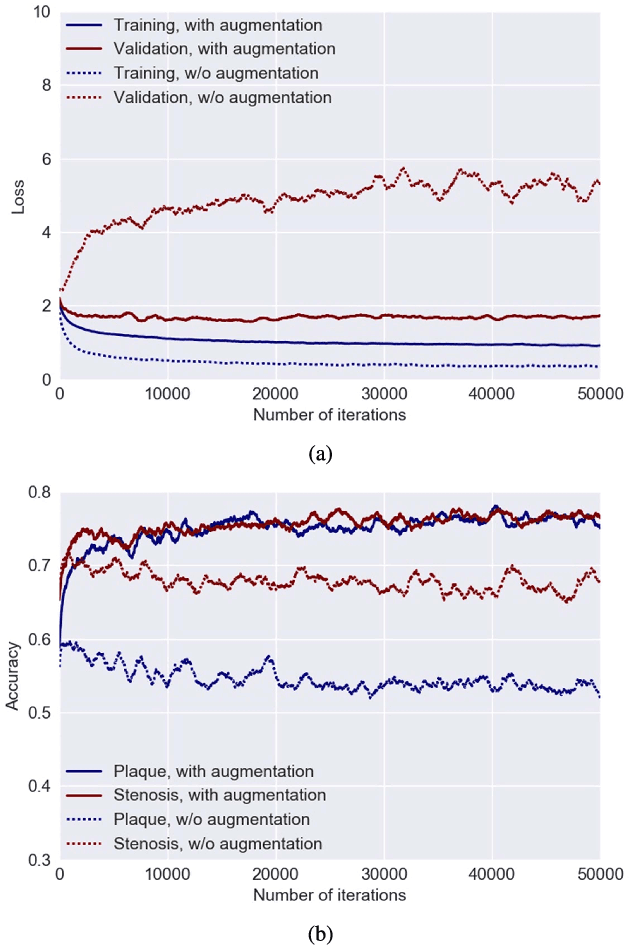
Abstract:Various types of atherosclerotic plaque and varying grades of stenosis could lead to different management of patients with coronary artery disease. Therefore, it is crucial to detect and classify the type of coronary artery plaque, as well as to detect and determine the degree of coronary artery stenosis. This study includes retrospectively collected clinically obtained coronary CT angiography (CCTA) scans of 163 patients. To perform automatic analysis for coronary artery plaque and stenosis classification, a multi-task recurrent convolutional neural network is applied on multi-planar reformatted (MPR) images of the coronary arteries. First, a 3D convolutional neural network is utilized to extract features along the coronary artery. Subsequently, the extracted features are aggregated by a recurrent neural network that performs two simultaneous multi-class classification tasks. In the first task, the network detects and characterizes the type of the coronary artery plaque (no plaque, non-calcified, mixed, calcified). In the second task, the network detects and determines the anatomical significance of the coronary artery stenosis (no stenosis, non-significant i.e. <50% luminal narrowing, significant i.e. >50% luminal narrowing). For detection and classification of coronary plaque, the method achieved an accuracy of 0.77. For detection and classification of stenosis, the method achieved an accuracy of 0.80. The results demonstrate that automatic detection and classification of coronary artery plaque and stenosis are feasible. This may enable automated triage of patients to those without coronary plaque and those with coronary plaque and stenosis in need for further cardiovascular workup.
Automatic calcium scoring in low-dose chest CT using deep neural networks with dilated convolutions
Feb 01, 2018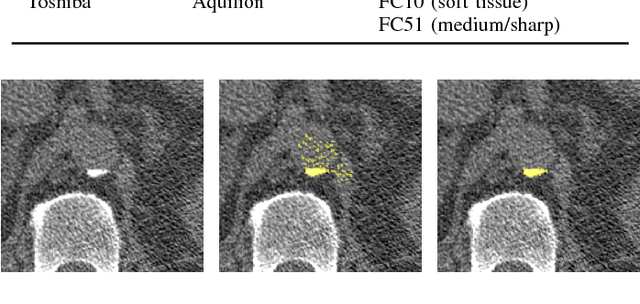
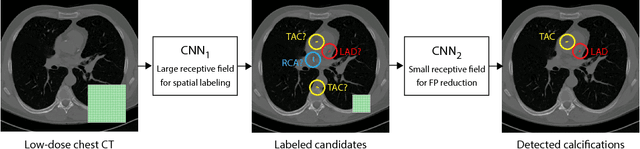
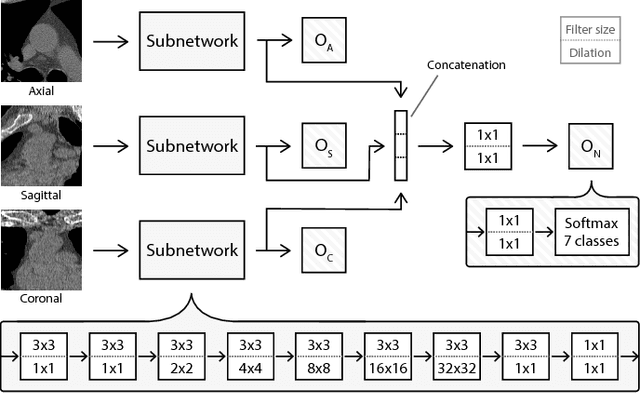
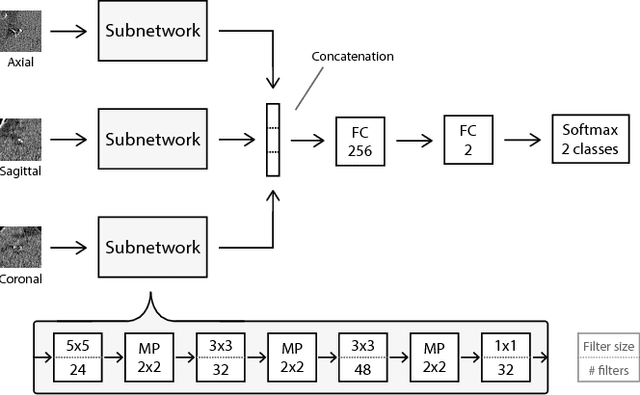
Abstract:Heavy smokers undergoing screening with low-dose chest CT are affected by cardiovascular disease as much as by lung cancer. Low-dose chest CT scans acquired in screening enable quantification of atherosclerotic calcifications and thus enable identification of subjects at increased cardiovascular risk. This paper presents a method for automatic detection of coronary artery, thoracic aorta and cardiac valve calcifications in low-dose chest CT using two consecutive convolutional neural networks. The first network identifies and labels potential calcifications according to their anatomical location and the second network identifies true calcifications among the detected candidates. This method was trained and evaluated on a set of 1744 CT scans from the National Lung Screening Trial. To determine whether any reconstruction or only images reconstructed with soft tissue filters can be used for calcification detection, we evaluated the method on soft and medium/sharp filter reconstructions separately. On soft filter reconstructions, the method achieved F1 scores of 0.89, 0.89, 0.67, and 0.55 for coronary artery, thoracic aorta, aortic valve and mitral valve calcifications, respectively. On sharp filter reconstructions, the F1 scores were 0.84, 0.81, 0.64, and 0.66, respectively. Linearly weighted kappa coefficients for risk category assignment based on per subject coronary artery calcium were 0.91 and 0.90 for soft and sharp filter reconstructions, respectively. These results demonstrate that the presented method enables reliable automatic cardiovascular risk assessment in all low-dose chest CT scans acquired for lung cancer screening.
Direct and Real-Time Cardiovascular Risk Prediction
Dec 08, 2017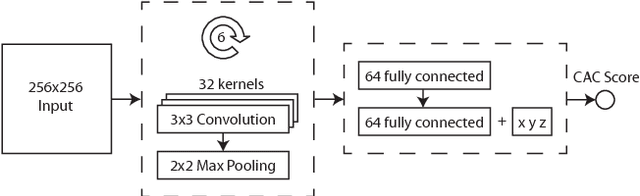
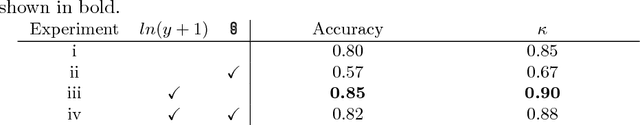
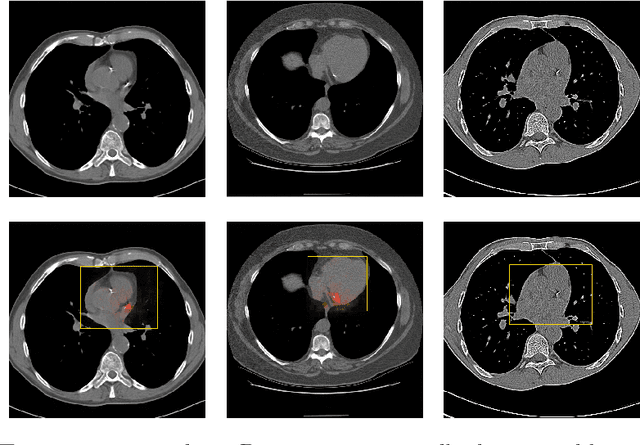
Abstract:Coronary artery calcium (CAC) burden quantified in low-dose chest CT is a predictor of cardiovascular events. We propose an automatic method for CAC quantification, circumventing intermediate segmentation of CAC. The method determines a bounding box around the heart using a ConvNet for localization. Subsequently, a dedicated ConvNet analyzes axial slices within the bounding boxes to determine CAC quantity by regression. A dataset of 1,546 baseline CT scans was used from the National Lung Screening Trial with manually identified CAC. The method achieved an ICC of 0.98 between manual reference and automatically obtained Agatston scores. Stratification of subjects into five cardiovascular risk categories resulted in an accuracy of 85\% and Cohen's linearly weighted $\kappa$ of 0.90. The results demonstrate that real-time quantification of CAC burden in chest CT without the need for segmentation of CAC is possible.
Deep learning analysis of the myocardium in coronary CT angiography for identification of patients with functionally significant coronary artery stenosis
Dec 06, 2017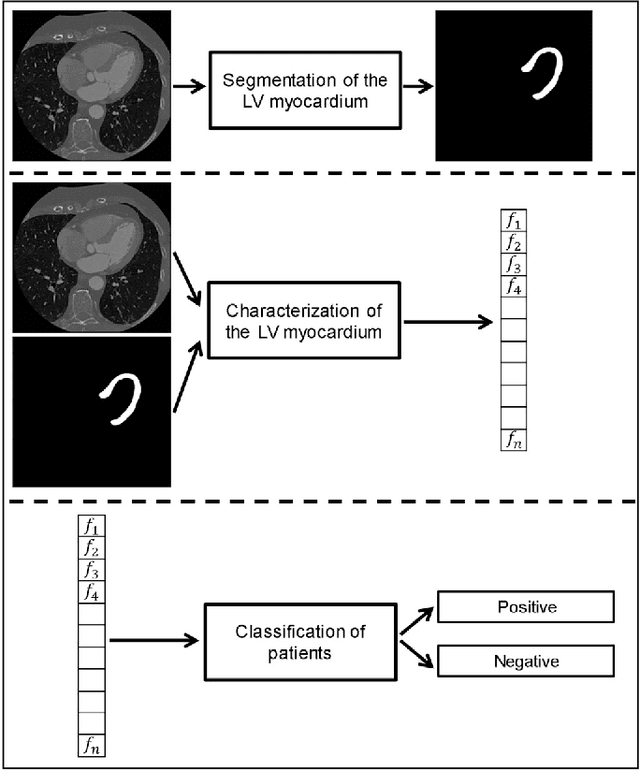
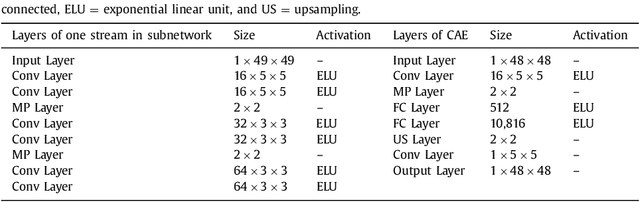
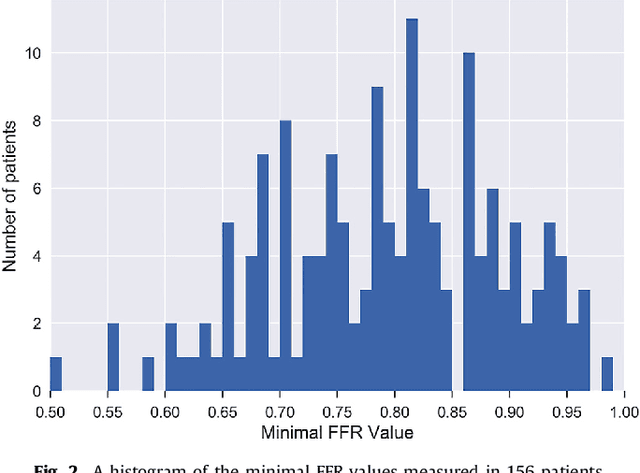
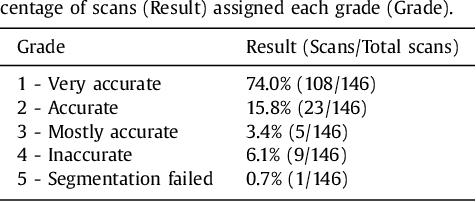
Abstract:In patients with coronary artery stenoses of intermediate severity, the functional significance needs to be determined. Fractional flow reserve (FFR) measurement, performed during invasive coronary angiography (ICA), is most often used in clinical practice. To reduce the number of ICA procedures, we present a method for automatic identification of patients with functionally significant coronary artery stenoses, employing deep learning analysis of the left ventricle (LV) myocardium in rest coronary CT angiography (CCTA). The study includes consecutively acquired CCTA scans of 166 patients with FFR measurements. To identify patients with a functionally significant coronary artery stenosis, analysis is performed in several stages. First, the LV myocardium is segmented using a multiscale convolutional neural network (CNN). To characterize the segmented LV myocardium, it is subsequently encoded using unsupervised convolutional autoencoder (CAE). Thereafter, patients are classified according to the presence of functionally significant stenosis using an SVM classifier based on the extracted and clustered encodings. Quantitative evaluation of LV myocardium segmentation in 20 images resulted in an average Dice coefficient of 0.91 and an average mean absolute distance between the segmented and reference LV boundaries of 0.7 mm. Classification of patients was evaluated in the remaining 126 CCTA scans in 50 10-fold cross-validation experiments and resulted in an area under the receiver operating characteristic curve of 0.74 +- 0.02. At sensitivity levels 0.60, 0.70 and 0.80, the corresponding specificity was 0.77, 0.71 and 0.59, respectively. The results demonstrate that automatic analysis of the LV myocardium in a single CCTA scan acquired at rest, without assessment of the anatomy of the coronary arteries, can be used to identify patients with functionally significant coronary artery stenosis.
Automatic Segmentation and Disease Classification Using Cardiac Cine MR Images
Aug 03, 2017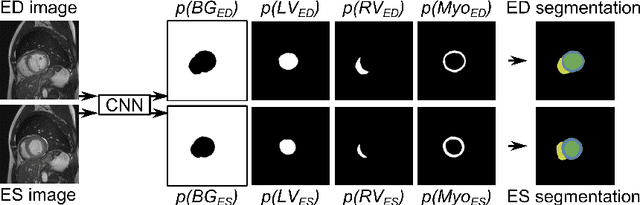
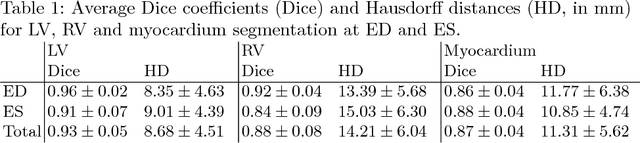

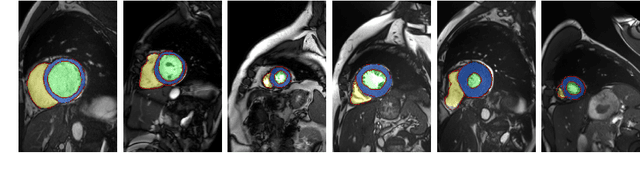
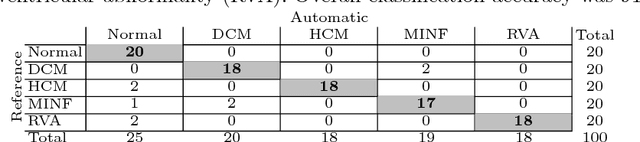
Abstract:Segmentation of the heart in cardiac cine MR is clinically used to quantify cardiac function. We propose a fully automatic method for segmentation and disease classification using cardiac cine MR images. A convolutional neural network (CNN) was designed to simultaneously segment the left ventricle (LV), right ventricle (RV) and myocardium in end-diastole (ED) and end-systole (ES) images. Features derived from the obtained segmentations were used in a Random Forest classifier to label patients as suffering from dilated cardiomyopathy, hypertrophic cardiomyopathy, heart failure following myocardial infarction, right ventricular abnormality, or no cardiac disease. The method was developed and evaluated using a balanced dataset containing images of 100 patients, which was provided in the MICCAI 2017 automated cardiac diagnosis challenge (ACDC). The segmentation and classification pipeline were evaluated in a four-fold stratified cross-validation. Average Dice scores between reference and automatically obtained segmentations were 0.94, 0.88 and 0.87 for the LV, RV and myocardium. The classifier assigned 91% of patients to the correct disease category. Segmentation and disease classification took 5 s per patient. The results of our study suggest that image-based diagnosis using cine MR cardiac scans can be performed automatically with high accuracy.
End-to-End Unsupervised Deformable Image Registration with a Convolutional Neural Network
Apr 20, 2017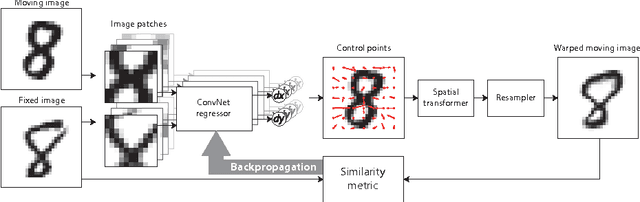
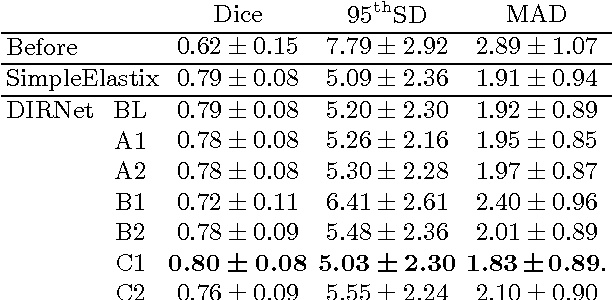

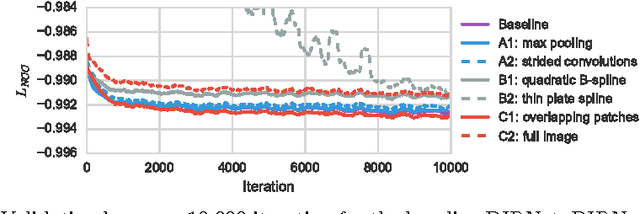
Abstract:In this work we propose a deep learning network for deformable image registration (DIRNet). The DIRNet consists of a convolutional neural network (ConvNet) regressor, a spatial transformer, and a resampler. The ConvNet analyzes a pair of fixed and moving images and outputs parameters for the spatial transformer, which generates the displacement vector field that enables the resampler to warp the moving image to the fixed image. The DIRNet is trained end-to-end by unsupervised optimization of a similarity metric between input image pairs. A trained DIRNet can be applied to perform registration on unseen image pairs in one pass, thus non-iteratively. Evaluation was performed with registration of images of handwritten digits (MNIST) and cardiac cine MR scans (Sunnybrook Cardiac Data). The results demonstrate that registration with DIRNet is as accurate as a conventional deformable image registration method with substantially shorter execution times.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge