Mauricio Reyes
Institute for Surgical Technology and Biomechanics, University of Bern, Switzerland
Assessing Reliability and Challenges of Uncertainty Estimations for Medical Image Segmentation
Jul 07, 2019
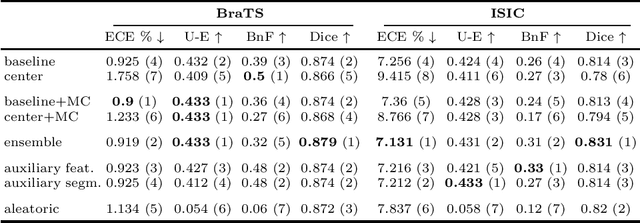
Abstract:Despite the recent improvements in overall accuracy, deep learning systems still exhibit low levels of robustness. Detecting possible failures is critical for a successful clinical integration of these systems, where each data point corresponds to an individual patient. Uncertainty measures are a promising direction to improve failure detection since they provide a measure of a system's confidence. Although many uncertainty estimation methods have been proposed for deep learning, little is known on their benefits and current challenges for medical image segmentation. Therefore, we report results of evaluating common voxel-wise uncertainty measures with respect to their reliability, and limitations on two medical image segmentation datasets. Results show that current uncertainty methods perform similarly and although they are well-calibrated at the dataset level, they tend to be miscalibrated at subject-level. Therefore, the reliability of uncertainty estimates is compromised, highlighting the importance of developing subject-wise uncertainty estimations. Additionally, among the benchmarked methods, we found auxiliary networks to be a valid alternative to common uncertainty methods since they can be applied to any previously trained segmentation model.
Learning Shape Representation on Sparse Point Clouds for Volumetric Image Segmentation
Jun 05, 2019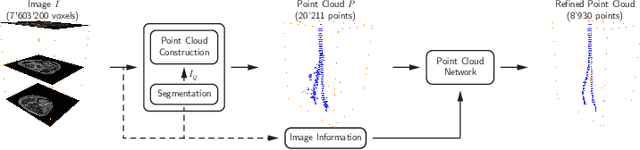
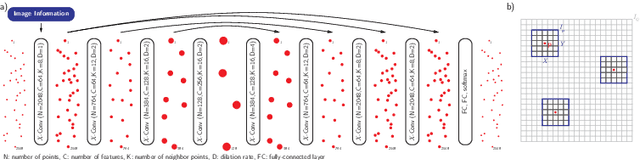
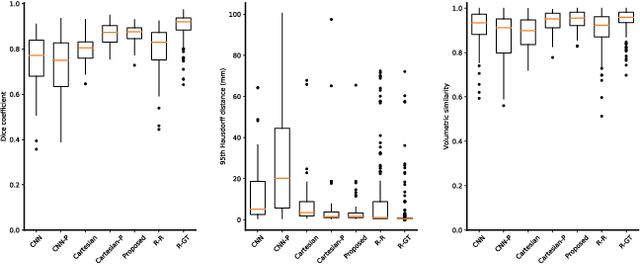
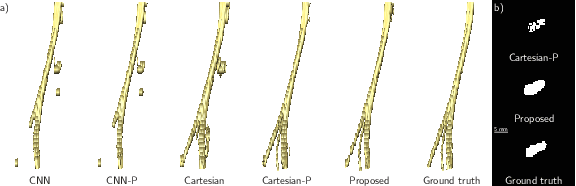
Abstract:Volumetric image segmentation with convolutional neural networks (CNNs) encounters several challenges, which are specific to medical images. Among these challenges are large volumes of interest, high class imbalances, and difficulties in learning shape representations. To tackle these challenges, we propose to improve over traditional CNN-based volumetric image segmentation through point-wise classification of point clouds. The sparsity of point clouds allows processing of entire image volumes, balancing highly imbalanced segmentation problems, and explicitly learning an anatomical shape. We build upon PointCNN, a neural network proposed to process point clouds, and propose here to jointly encode shape and volumetric information within the point cloud in a compact and computationally effective manner. We demonstrate how this approach can then be used to refine CNN-based segmentation, which yields significantly improved results in our experiments on the difficult task of peripheral nerve segmentation from magnetic resonance neurography images. By synthetic experiments, we further show the capability of our approach in learning an explicit anatomical shape representation.
Informative sample generation using class aware generative adversarial networks for classification of chest Xrays
Apr 30, 2019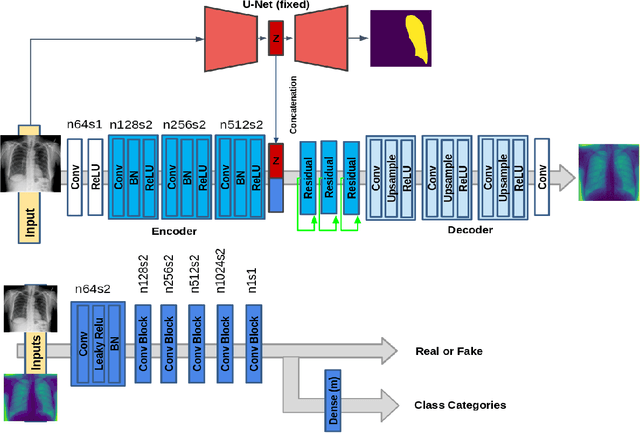
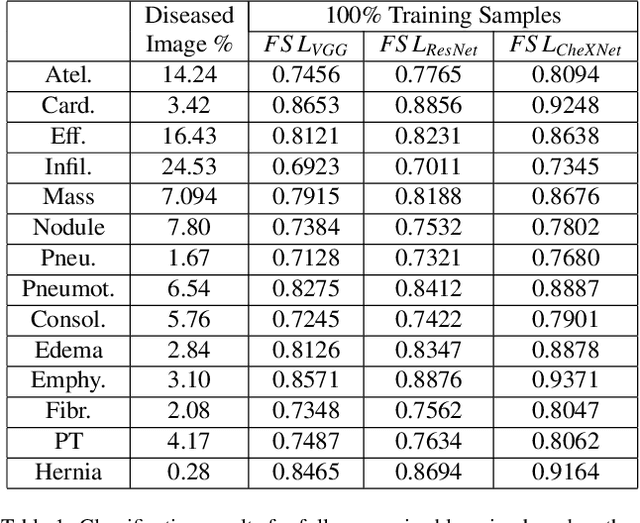
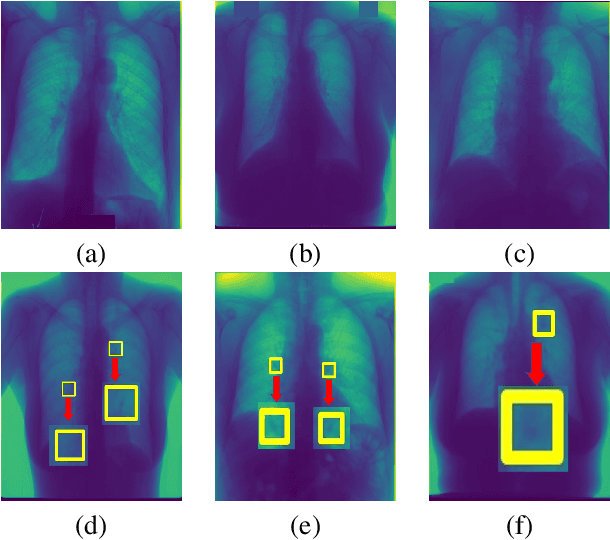
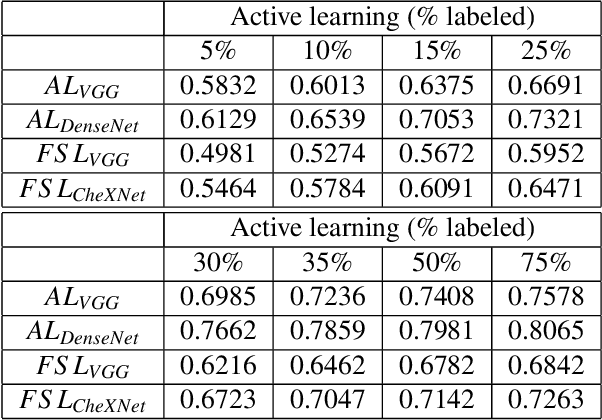
Abstract:Training robust deep learning (DL) systems for disease detection from medical images is challenging due to limited images covering different disease types and severity. The problem is especially acute, where there is a severe class imbalance. We propose an active learning (AL) framework to select most informative samples for training our model using a Bayesian neural network. Informative samples are then used within a novel class aware generative adversarial network (CAGAN) to generate realistic chest xray images for data augmentation by transferring characteristics from one class label to another. Experiments show our proposed AL framework is able to achieve state-of-the-art performance by using about $35\%$ of the full dataset, thus saving significant time and effort over conventional methods.
Automatic detection of lesion load change in Multiple Sclerosis using convolutional neural networks with segmentation confidence
Apr 05, 2019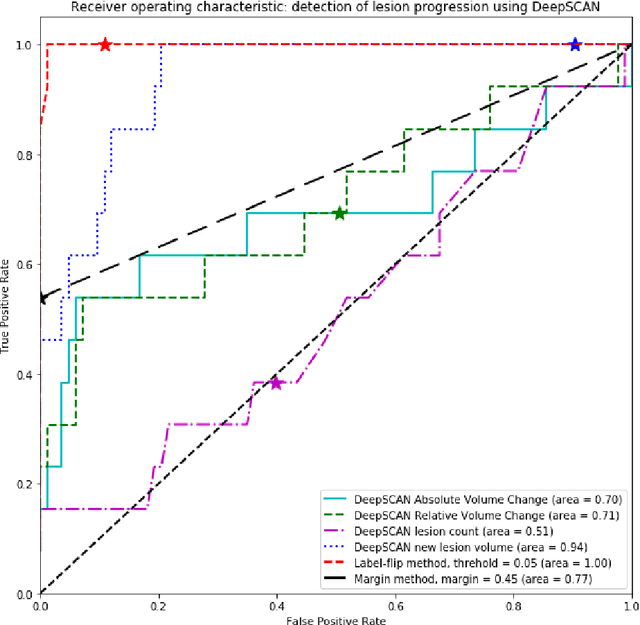
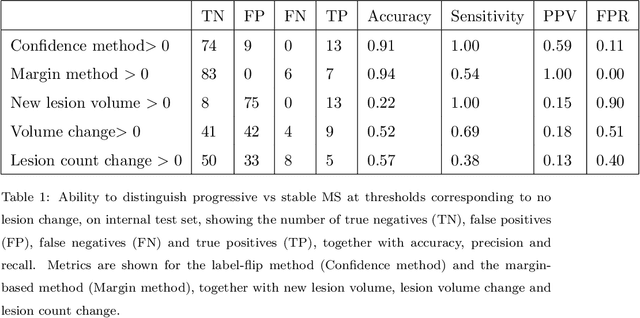
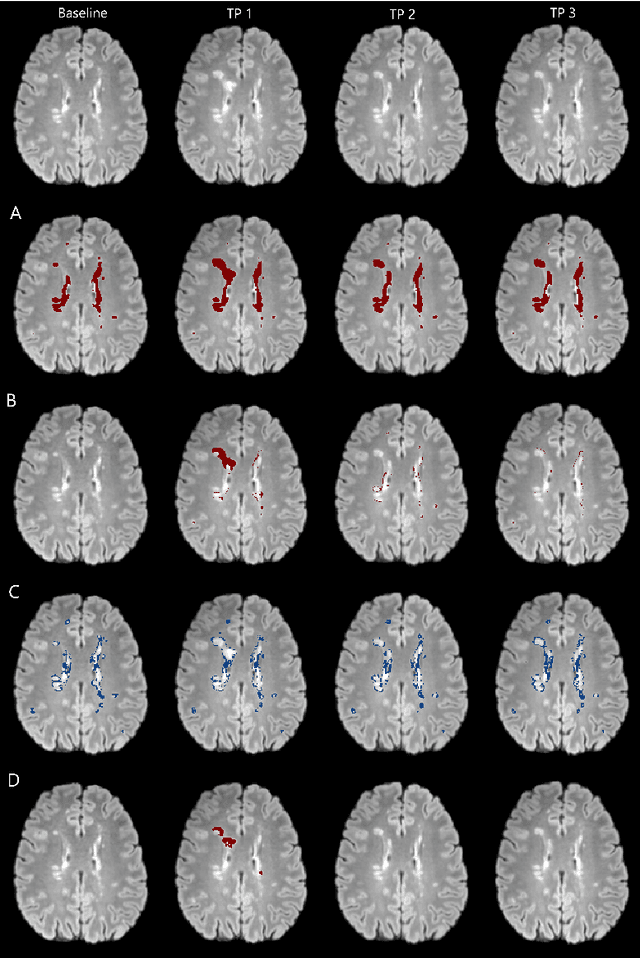
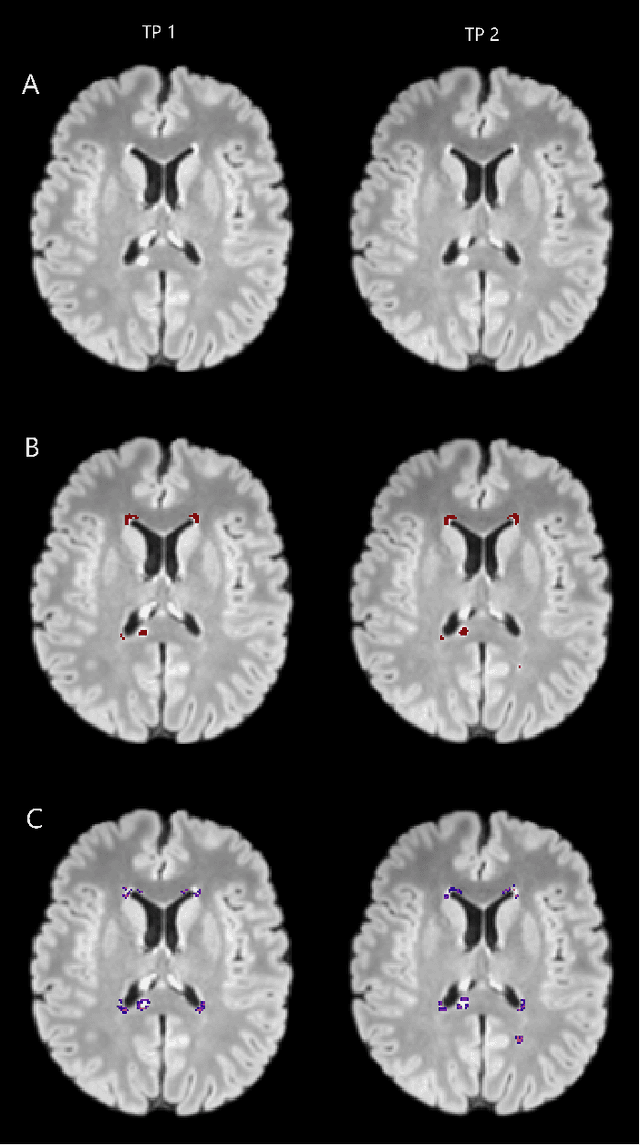
Abstract:The detection of new or enlarged white-matter lesions in multiple sclerosis is a vital task in the monitoring of patients undergoing disease-modifying treatment for multiple sclerosis. However, the definition of 'new or enlarged' is not fixed, and it is known that lesion-counting is highly subjective, with high degree of inter- and intra-rater variability. Automated methods for lesion quantification hold the potential to make the detection of new and enlarged lesions consistent and repeatable. However, the majority of lesion segmentation algorithms are not evaluated for their ability to separate progressive from stable patients, despite this being a pressing clinical use-case. In this paper we show that change in volumetric measurements of lesion load alone is not a good method for performing this separation, even for highly performing segmentation methods. Instead, we propose a method for identifying lesion changes of high certainty, and establish on a dataset of longitudinal multiple sclerosis cases that this method is able to separate progressive from stable timepoints with a very high level of discrimination (AUC = 0.99), while changes in lesion volume are much less able to perform this separation (AUC = 0.71). Validation of the method on a second external dataset confirms that the method is able to generalize beyond the setting in which it was trained, achieving an accuracy of 83% in separating stable and progressive timepoints. Both lesion volume and count have previously been shown to be strong predictors of disease course across a population. However, we demonstrate that for individual patients, changes in these measures are not an adequate means of establishing no evidence of disease activity. Meanwhile, directly detecting tissue which changes, with high confidence, from non-lesion to lesion is a feasible methodology for identifying radiologically active patients.
Few-shot brain segmentation from weakly labeled data with deep heteroscedastic multi-task networks
Apr 04, 2019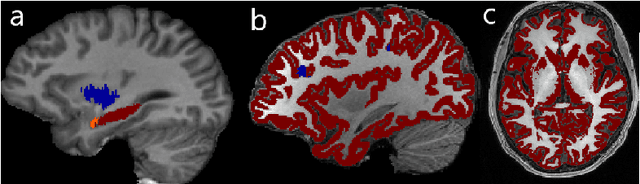
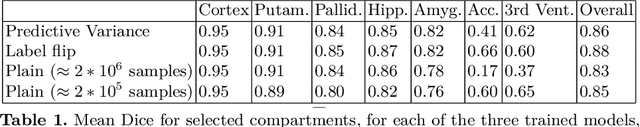
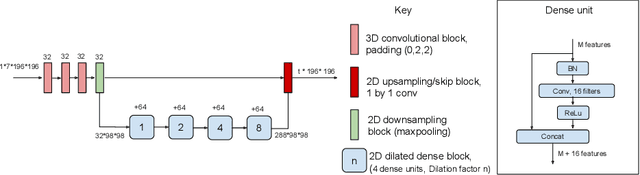
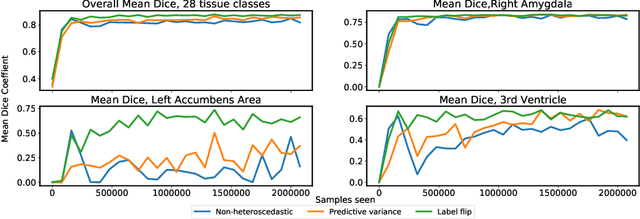
Abstract:In applications of supervised learning applied to medical image segmentation, the need for large amounts of labeled data typically goes unquestioned. In particular, in the case of brain anatomy segmentation, hundreds or thousands of weakly-labeled volumes are often used as training data. In this paper, we first observe that for many brain structures, a small number of training examples, (n=9), weakly labeled using Freesurfer 6.0, plus simple data augmentation, suffice as training data to achieve high performance, achieving an overall mean Dice coefficient of $0.84 \pm 0.12$ compared to Freesurfer over 28 brain structures in T1-weighted images of $\approx 4000$ 9-10 year-olds from the Adolescent Brain Cognitive Development study. We then examine two varieties of heteroscedastic network as a method for improving classification results. An existing proposal by Kendall and Gal, which uses Monte-Carlo inference to learn to predict the variance of each prediction, yields an overall mean Dice of $0.85 \pm 0.14$ and showed statistically significant improvements over 25 brain structures. Meanwhile a novel heteroscedastic network which directly learns the probability that an example has been mislabeled yielded an overall mean Dice of $0.87 \pm 0.11$ and showed statistically significant improvements over all but one of the brain structures considered. The loss function associated to this network can be interpreted as performing a form of learned label smoothing, where labels are only smoothed where they are judged to be uncertain.
Simultaneous lesion and neuroanatomy segmentation in Multiple Sclerosis using deep neural networks
Jan 22, 2019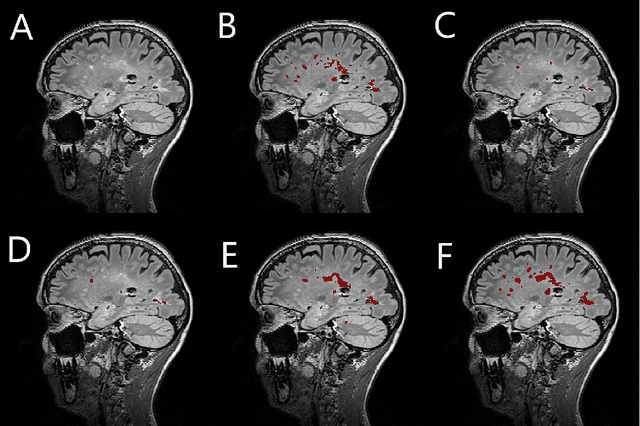
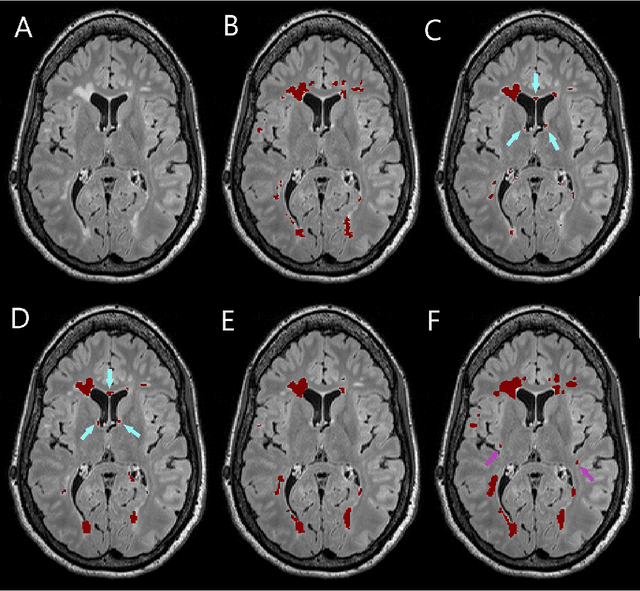
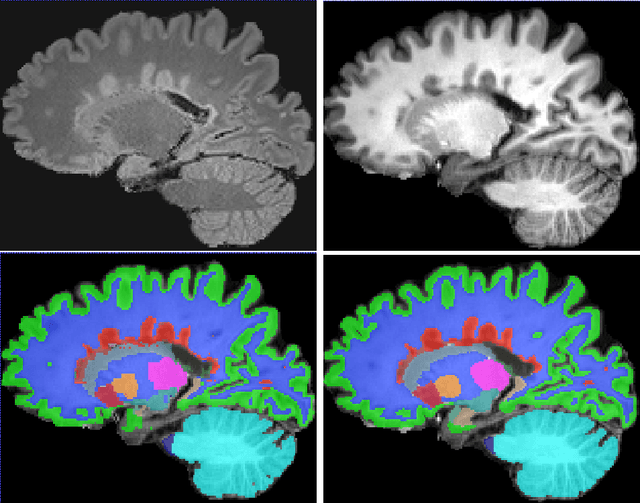
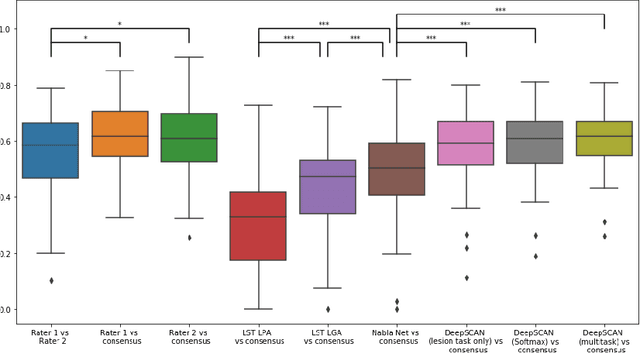
Abstract:Segmentation of both white matter lesions and deep grey matter structures is an important task in the quantification of magnetic resonance imaging in multiple sclerosis. Typically these tasks are performed separately: in this paper we present a single CNN-based segmentation solution for providing fast, reliable segmentations of multimodal MR imagies into lesion classes and healthy-appearing grey- and white-matter structures. We show substantial, statistically significant improvements in both Dice coefficient and in lesion-wise specificity and sensitivity, compared to previous approaches, and agreement with individual human raters in the range of human inter-rater variability. The method is trained on data gathered from a single centre: nonetheless, it performs well on data from centres, scanners and field-strengths not represented in the training dataset. A retrospective study found that the classifier successfully identified lesions missed by the human raters. Lesion labels were provided by human raters, while weak labels for other brain structures (including CSF, cortical grey matter, cortical white matter, cerebellum, amygdala, hippocampus, subcortical GM structures and choroid plexus) were provided by Freesurfer 5.3. The segmentations of these structures compared well, not only with Freesurfer 5.3, but also with FSL-First and Freesurfer 6.1.
Deep Learning versus Classical Regression for Brain Tumor Patient Survival Prediction
Nov 12, 2018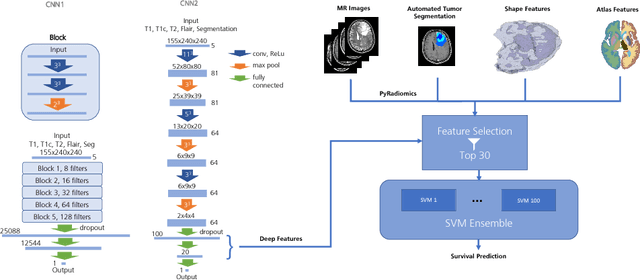
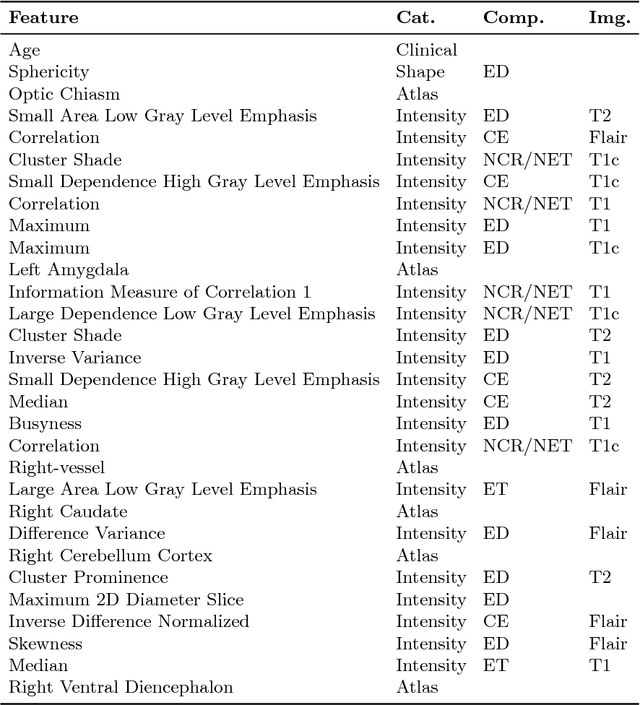
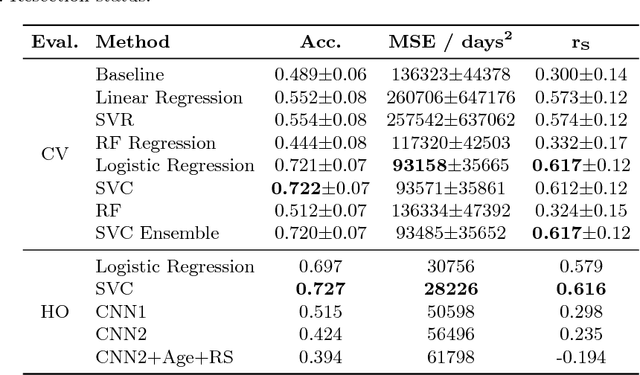
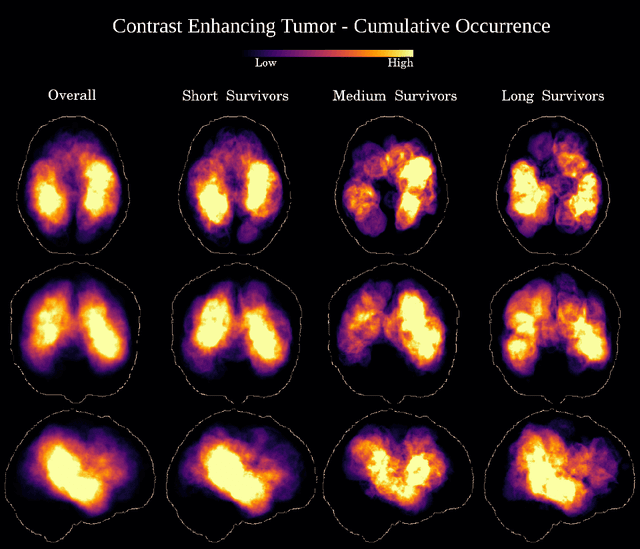
Abstract:Deep learning for regression tasks on medical imaging data has shown promising results. However, compared to other approaches, their power is strongly linked to the dataset size. In this study, we evaluate 3D-convolutional neural networks (CNNs) and classical regression methods with hand-crafted features for survival time regression of patients with high grade brain tumors. The tested CNNs for regression showed promising but unstable results. The best performing deep learning approach reached an accuracy of 51.5% on held-out samples of the training set. All tested deep learning experiments were outperformed by a Support Vector Classifier (SVC) using 30 radiomic features. The investigated features included intensity, shape, location and deep features. The submitted method to the BraTS 2018 survival prediction challenge is an ensemble of SVCs, which reached a cross-validated accuracy of 72.2% on the BraTS 2018 training set, 57.1% on the validation set, and 42.9% on the testing set. The results suggest that more training data is necessary for a stable performance of a CNN model for direct regression from magnetic resonance images, and that non-imaging clinical patient information is crucial along with imaging information.
Automatic brain tumor grading from MRI data using convolutional neural networks and quality assessment
Sep 25, 2018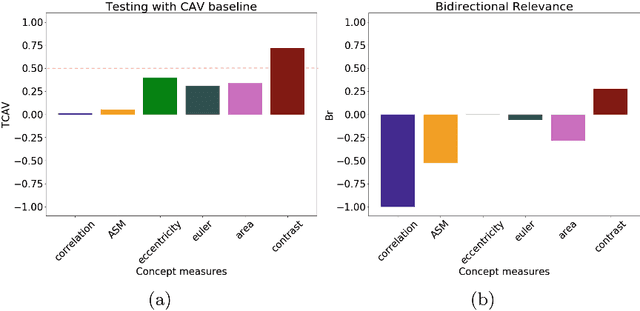
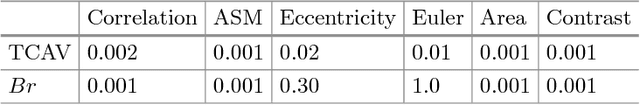
Abstract:Glioblastoma Multiforme is a high grade, very aggressive, brain tumor, with patients having a poor prognosis. Lower grade gliomas are less aggressive, but they can evolve into higher grade tumors over time. Patient management and treatment can vary considerably with tumor grade, ranging from tumor resection followed by a combined radio- and chemotherapy to a "wait and see" approach. Hence, tumor grading is important for adequate treatment planning and monitoring. The gold standard for tumor grading relies on histopathological diagnosis of biopsy specimens. However, this procedure is invasive, time consuming, and prone to sampling error. Given these disadvantages, automatic tumor grading from widely used MRI protocols would be clinically important, as a way to expedite treatment planning and assessment of tumor evolution. In this paper, we propose to use Convolutional Neural Networks for predicting tumor grade directly from imaging data. In this way, we overcome the need for expert annotations of regions of interest. We evaluate two prediction approaches: from the whole brain, and from an automatically defined tumor region. Finally, we employ interpretability methodologies as a quality assurance stage to check if the method is using image regions indicative of tumor grade for classification.
Magnetic Resonance Fingerprinting Reconstruction via Spatiotemporal Convolutional Neural Networks
Jul 24, 2018
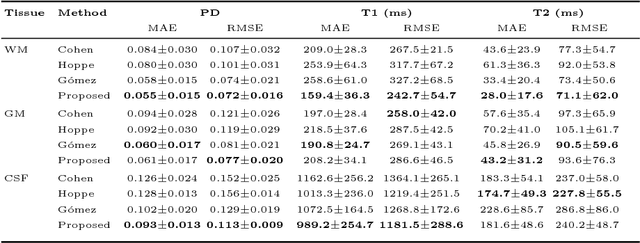
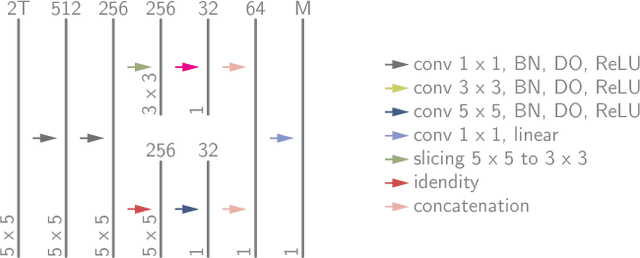
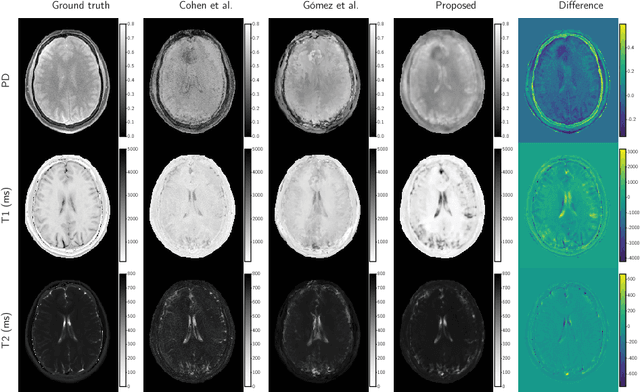
Abstract:Magnetic resonance fingerprinting (MRF) quantifies multiple nuclear magnetic resonance parameters in a single and fast acquisition. Standard MRF reconstructs parametric maps using dictionary matching, which lacks scalability due to computational inefficiency. We propose to perform MRF map reconstruction using a spatiotemporal convolutional neural network, which exploits the relationship between neighboring MRF signal evolutions to replace the dictionary matching. We evaluate our method on multiparametric brain scans and compare it to three recent MRF reconstruction approaches. Our method achieves state-of-the-art reconstruction accuracy and yields qualitatively more appealing maps compared to other reconstruction methods. In addition, the reconstruction time is significantly reduced compared to a dictionary-based approach.
Efficient Active Learning for Image Classification and Segmentation using a Sample Selection and Conditional Generative Adversarial Network
Jun 22, 2018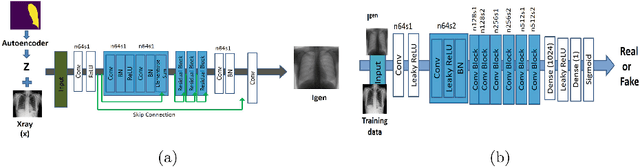
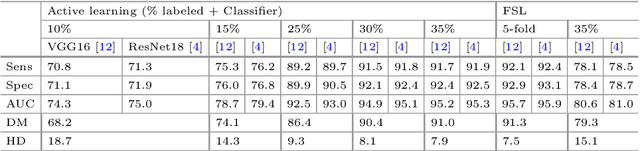
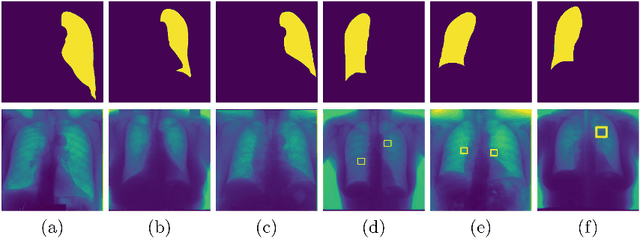
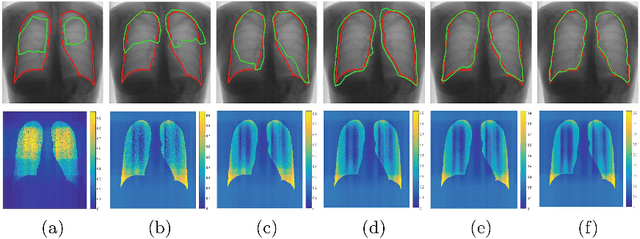
Abstract:Training robust deep learning (DL) systems for medical image classification or segmentation is challenging due to limited images covering different disease types and severity. We propose an active learning (AL) framework to select most informative samples and add to the training data. We use conditional generative adversarial networks (cGANs) to generate realistic chest xray images with different disease characteristics by conditioning its generation on a real image sample. Informative samples to add to the training set are identified using a Bayesian neural network. Experiments show our proposed AL framework is able to achieve state of the art performance by using about 35% of the full dataset, thus saving significant time and effort over conventional methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge