Mateja Jamnik
GCI: A (G)raph (C)oncept (I)nterpretation Framework
Feb 09, 2023Abstract:Explainable AI (XAI) underwent a recent surge in research on concept extraction, focusing on extracting human-interpretable concepts from Deep Neural Networks. An important challenge facing concept extraction approaches is the difficulty of interpreting and evaluating discovered concepts, especially for complex tasks such as molecular property prediction. We address this challenge by presenting GCI: a (G)raph (C)oncept (I)nterpretation framework, used for quantitatively measuring alignment between concepts discovered from Graph Neural Networks (GNNs) and their corresponding human interpretations. GCI encodes concept interpretations as functions, which can be used to quantitatively measure the alignment between a given interpretation and concept definition. We demonstrate four applications of GCI: (i) quantitatively evaluating concept extractors, (ii) measuring alignment between concept extractors and human interpretations, (iii) measuring the completeness of interpretations with respect to an end task and (iv) a practical application of GCI to molecular property prediction, in which we demonstrate how to use chemical functional groups to explain GNNs trained on molecular property prediction tasks, and implement interpretations with a 0.76 AUCROC completeness score.
Towards Robust Metrics for Concept Representation Evaluation
Jan 25, 2023



Abstract:Recent work on interpretability has focused on concept-based explanations, where deep learning models are explained in terms of high-level units of information, referred to as concepts. Concept learning models, however, have been shown to be prone to encoding impurities in their representations, failing to fully capture meaningful features of their inputs. While concept learning lacks metrics to measure such phenomena, the field of disentanglement learning has explored the related notion of underlying factors of variation in the data, with plenty of metrics to measure the purity of such factors. In this paper, we show that such metrics are not appropriate for concept learning and propose novel metrics for evaluating the purity of concept representations in both approaches. We show the advantage of these metrics over existing ones and demonstrate their utility in evaluating the robustness of concept representations and interventions performed on them. In addition, we show their utility for benchmarking state-of-the-art methods from both families and find that, contrary to common assumptions, supervision alone may not be sufficient for pure concept representations.
Weight Predictor Network with Feature Selection for Small Sample Tabular Biomedical Data
Nov 28, 2022Abstract:Tabular biomedical data is often high-dimensional but with a very small number of samples. Although recent work showed that well-regularised simple neural networks could outperform more sophisticated architectures on tabular data, they are still prone to overfitting on tiny datasets with many potentially irrelevant features. To combat these issues, we propose Weight Predictor Network with Feature Selection (WPFS) for learning neural networks from high-dimensional and small sample data by reducing the number of learnable parameters and simultaneously performing feature selection. In addition to the classification network, WPFS uses two small auxiliary networks that together output the weights of the first layer of the classification model. We evaluate on nine real-world biomedical datasets and demonstrate that WPFS outperforms other standard as well as more recent methods typically applied to tabular data. Furthermore, we investigate the proposed feature selection mechanism and show that it improves performance while providing useful insights into the learning task.
Discrete Lagrangian Neural Networks with Automatic Symmetry Discovery
Nov 20, 2022Abstract:By one of the most fundamental principles in physics, a dynamical system will exhibit those motions which extremise an action functional. This leads to the formation of the Euler-Lagrange equations, which serve as a model of how the system will behave in time. If the dynamics exhibit additional symmetries, then the motion fulfils additional conservation laws, such as conservation of energy (time invariance), momentum (translation invariance), or angular momentum (rotational invariance). To learn a system representation, one could learn the discrete Euler-Lagrange equations, or alternatively, learn the discrete Lagrangian function $\mathcal{L}_d$ which defines them. Based on ideas from Lie group theory, in this work we introduce a framework to learn a discrete Lagrangian along with its symmetry group from discrete observations of motions and, therefore, identify conserved quantities. The learning process does not restrict the form of the Lagrangian, does not require velocity or momentum observations or predictions and incorporates a cost term which safeguards against unwanted solutions and against potential numerical issues in forward simulations. The learnt discrete quantities are related to their continuous analogues using variational backward error analysis and numerical results demonstrate the improvement such models can have both qualitatively and quantitatively even in the presence of noise.
Explainer Divergence Scores (EDS): Some Post-Hoc Explanations May be Effective for Detecting Unknown Spurious Correlations
Nov 14, 2022Abstract:Recent work has suggested post-hoc explainers might be ineffective for detecting spurious correlations in Deep Neural Networks (DNNs). However, we show there are serious weaknesses with the existing evaluation frameworks for this setting. Previously proposed metrics are extremely difficult to interpret and are not directly comparable between explainer methods. To alleviate these constraints, we propose a new evaluation methodology, Explainer Divergence Scores (EDS), grounded in an information theory approach to evaluate explainers. EDS is easy to interpret and naturally comparable across explainers. We use our methodology to compare the detection performance of three different explainers - feature attribution methods, influential examples and concept extraction, on two different image datasets. We discover post-hoc explainers often contain substantial information about a DNN's dependence on spurious artifacts, but in ways often imperceptible to human users. This suggests the need for new techniques that can use this information to better detect a DNN's reliance on spurious correlations.
Graph-Conditioned MLP for High-Dimensional Tabular Biomedical Data
Nov 11, 2022



Abstract:Genome-wide studies leveraging recent high-throughput sequencing technologies collect high-dimensional data. However, they usually include small cohorts of patients, and the resulting tabular datasets suffer from the "curse of dimensionality". Training neural networks on such datasets is typically unstable, and the models overfit. One problem is that modern weight initialisation strategies make simplistic assumptions unsuitable for small-size datasets. We propose Graph-Conditioned MLP, a novel method to introduce priors on the parameters of an MLP. Instead of randomly initialising the first layer, we condition it directly on the training data. More specifically, we create a graph for each feature in the dataset (e.g., a gene), where each node represents a sample from the same dataset (e.g., a patient). We then use Graph Neural Networks (GNNs) to learn embeddings from these graphs and use the embeddings to initialise the MLP's parameters. Our approach opens the prospect of introducing additional biological knowledge when constructing the graphs. We present early results on 7 classification tasks from gene expression data and show that GC-MLP outperforms an MLP.
Distributed representations of graphs for drug pair scoring
Sep 19, 2022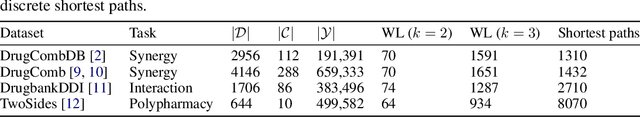
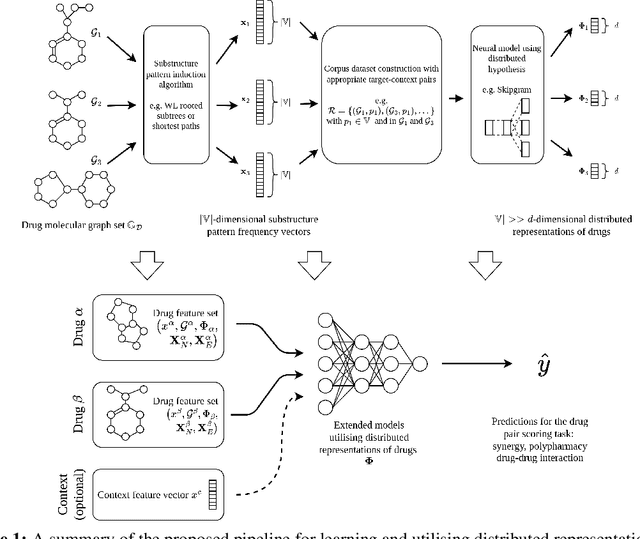
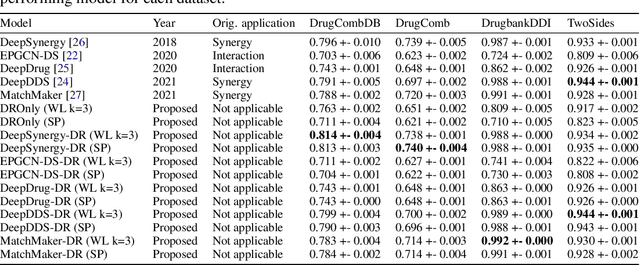
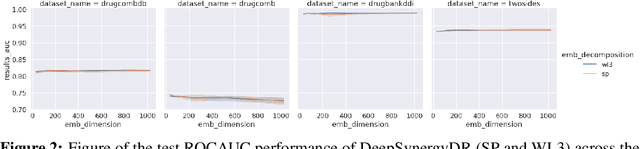
Abstract:In this paper we study the practicality and usefulness of incorporating distributed representations of graphs into models within the context of drug pair scoring. We argue that the real world growth and update cycles of drug pair scoring datasets subvert the limitations of transductive learning associated with distributed representations. Furthermore, we argue that the vocabulary of discrete substructure patterns induced over drug sets is not dramatically large due to the limited set of atom types and constraints on bonding patterns enforced by chemistry. Under this pretext, we explore the effectiveness of distributed representations of the molecular graphs of drugs in drug pair scoring tasks such as drug synergy, polypharmacy, and drug-drug interaction prediction. To achieve this, we present a methodology for learning and incorporating distributed representations of graphs within a unified framework for drug pair scoring. Subsequently, we augment a number of recent and state-of-the-art models to utilise our embeddings. We empirically show that the incorporation of these embeddings improves downstream performance of almost every model across different drug pair scoring tasks, even those the original model was not designed for. We publicly release all of our drug embeddings for the DrugCombDB, DrugComb, DrugbankDDI, and TwoSides datasets.
Concept Embedding Models
Sep 19, 2022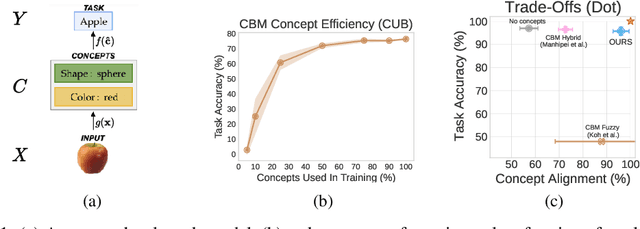
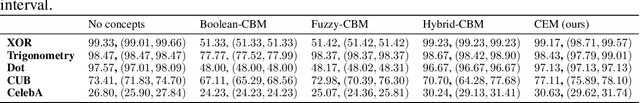
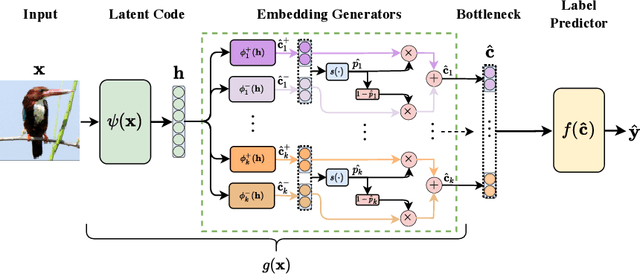
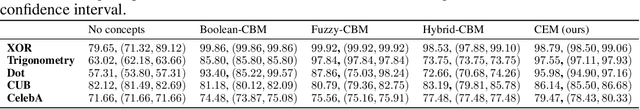
Abstract:Deploying AI-powered systems requires trustworthy models supporting effective human interactions, going beyond raw prediction accuracy. Concept bottleneck models promote trustworthiness by conditioning classification tasks on an intermediate level of human-like concepts. This enables human interventions which can correct mispredicted concepts to improve the model's performance. However, existing concept bottleneck models are unable to find optimal compromises between high task accuracy, robust concept-based explanations, and effective interventions on concepts -- particularly in real-world conditions where complete and accurate concept supervisions are scarce. To address this, we propose Concept Embedding Models, a novel family of concept bottleneck models which goes beyond the current accuracy-vs-interpretability trade-off by learning interpretable high-dimensional concept representations. Our experiments demonstrate that Concept Embedding Models (1) attain better or competitive task accuracy w.r.t. standard neural models without concepts, (2) provide concept representations capturing meaningful semantics including and beyond their ground truth labels, (3) support test-time concept interventions whose effect in test accuracy surpasses that in standard concept bottleneck models, and (4) scale to real-world conditions where complete concept supervisions are scarce.
Encoding Concepts in Graph Neural Networks
Aug 07, 2022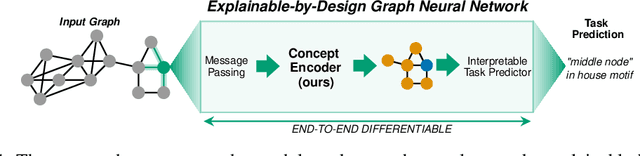
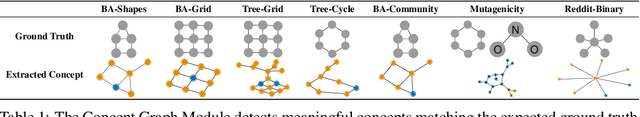
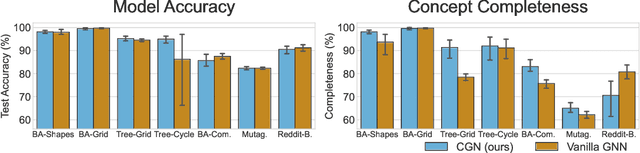
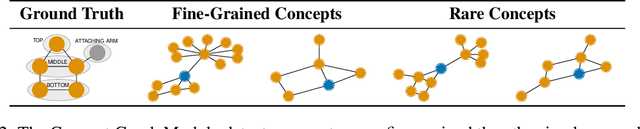
Abstract:The opaque reasoning of Graph Neural Networks induces a lack of human trust. Existing graph network explainers attempt to address this issue by providing post-hoc explanations, however, they fail to make the model itself more interpretable. To fill this gap, we introduce the Concept Encoder Module, the first differentiable concept-discovery approach for graph networks. The proposed approach makes graph networks explainable by design by first discovering graph concepts and then using these to solve the task. Our results demonstrate that this approach allows graph networks to: (i) attain model accuracy comparable with their equivalent vanilla versions, (ii) discover meaningful concepts that achieve high concept completeness and purity scores, (iii) provide high-quality concept-based logic explanations for their prediction, and (iv) support effective interventions at test time: these can increase human trust as well as significantly improve model performance.
Representational Systems Theory: A Unified Approach to Encoding, Analysing and Transforming Representations
Jun 07, 2022Abstract:The study of representations is of fundamental importance to any form of communication, and our ability to exploit them effectively is paramount. This article presents a novel theory -- Representational Systems Theory -- that is designed to abstractly encode a wide variety of representations from three core perspectives: syntax, entailment, and their properties. By introducing the concept of a construction space, we are able to encode each of these core components under a single, unifying paradigm. Using our Representational Systems Theory, it becomes possible to structurally transform representations in one system into representations in another. An intrinsic facet of our structural transformation technique is representation selection based on properties that representations possess, such as their relative cognitive effectiveness or structural complexity. A major theoretical barrier to providing general structural transformation techniques is a lack of terminating algorithms. Representational Systems Theory permits the derivation of partial transformations when no terminating algorithm can produce a full transformation. Since Representational Systems Theory provides a universal approach to encoding representational systems, a further key barrier is eliminated: the need to devise system-specific structural transformation algorithms, that are necessary when different systems adopt different formalisation approaches. Consequently, Representational Systems Theory is the first general framework that provides a unified approach to encoding representations, supports representation selection via structural transformations, and has the potential for widespread practical application.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge