Marleen de Bruijne
for the ALFA study
Automated Segmentation and Volume Measurement of Intracranial Carotid Artery Calcification on Non-Contrast CT
Jul 20, 2021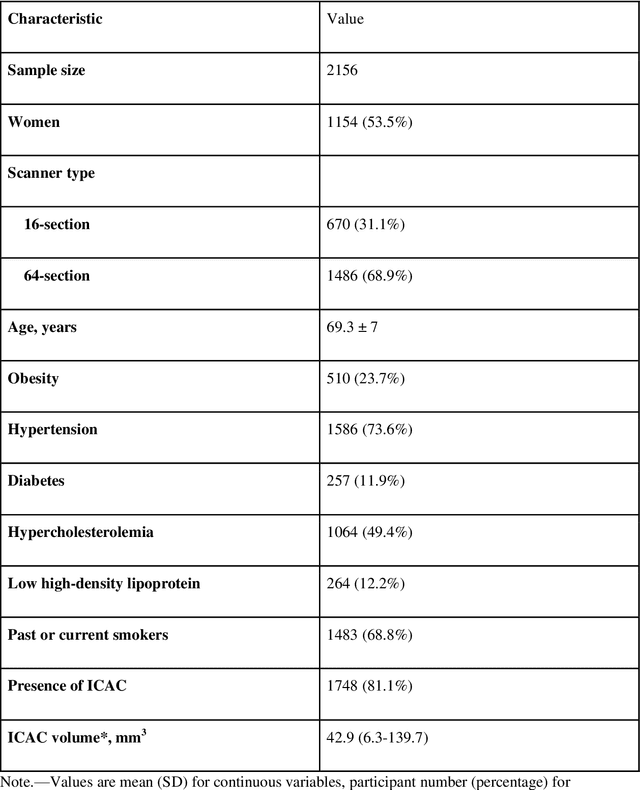
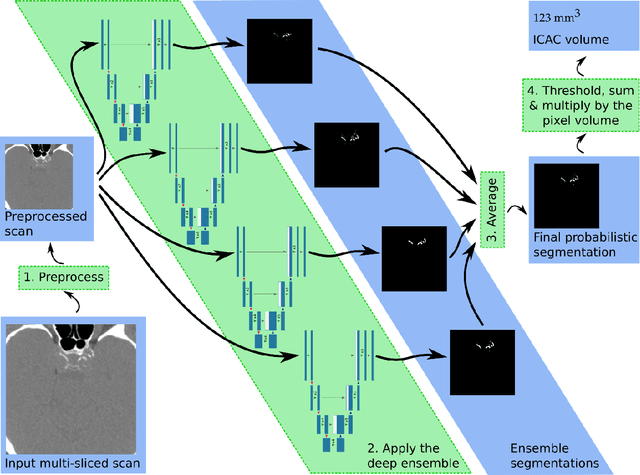
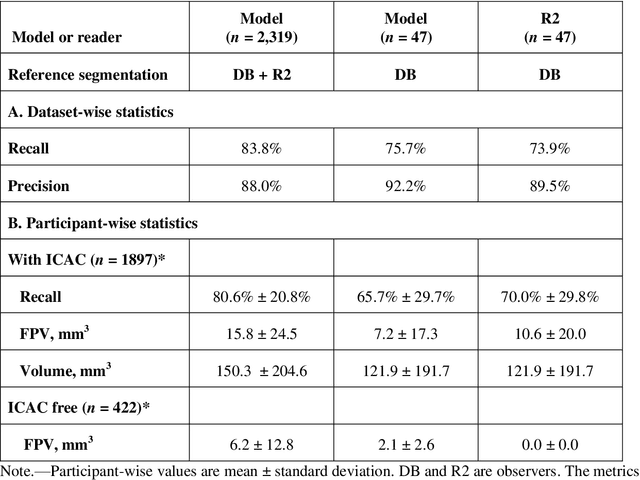
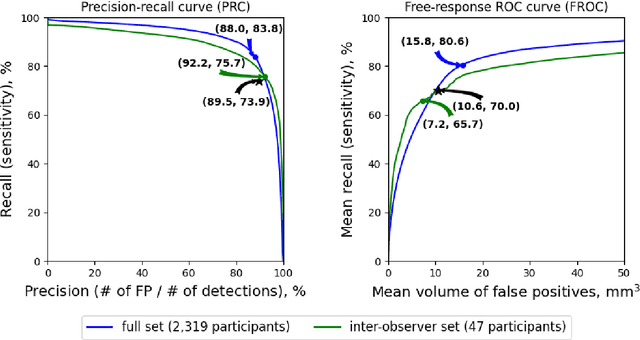
Abstract:Purpose: To evaluate a fully-automated deep-learning-based method for assessment of intracranial carotid artery calcification (ICAC). Methods: Two observers manually delineated ICAC in non-contrast CT scans of 2,319 participants (mean age 69 (SD 7) years; 1154 women) of the Rotterdam Study, prospectively collected between 2003 and 2006. These data were used to retrospectively develop and validate a deep-learning-based method for automated ICAC delineation and volume measurement. To evaluate the method, we compared manual and automatic assessment (computed using ten-fold cross-validation) with respect to 1) the agreement with an independent observer's assessment (available in a random subset of 47 scans); 2) the accuracy in delineating ICAC as judged via blinded visual comparison by an expert; 3) the association with first stroke incidence from the scan date until 2012. All method performance metrics were computed using 10-fold cross-validation. Results: The automated delineation of ICAC reached sensitivity of 83.8% and positive predictive value (PPV) of 88%. The intraclass correlation between automatic and manual ICAC volume measures was 0.98 (95% CI: 0.97, 0.98; computed in the entire dataset). Measured between the assessments of independent observers, sensitivity was 73.9%, PPV was 89.5%, and intraclass correlation was 0.91 (95% CI: 0.84, 0.95; computed in the 47-scan subset). In the blinded visual comparisons, automatic delineations were more accurate than manual ones (p-value = 0.01). The association of ICAC volume with incident stroke was similarly strong for both automated (hazard ratio, 1.38 (95% CI: 1.12, 1.75) and manually measured volumes (hazard ratio, 1.48 (95% CI: 1.20, 1.87)). Conclusions: The developed model was capable of automated segmentation and volume quantification of ICAC with accuracy comparable to human experts.
Adversarial Heart Attack: Neural Networks Fooled to Segment Heart Symbols in Chest X-Ray Images
Apr 07, 2021
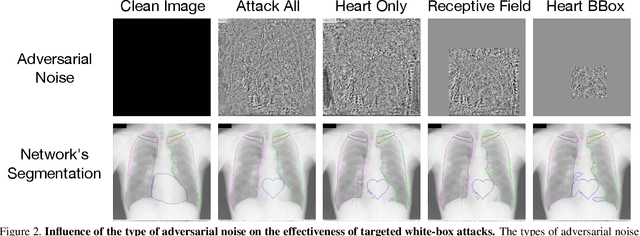
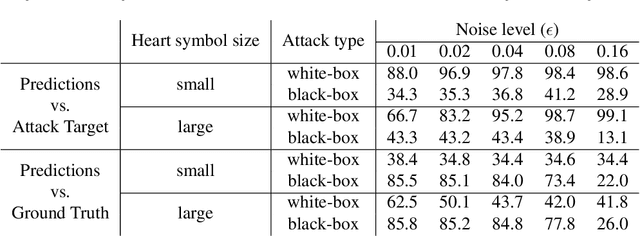
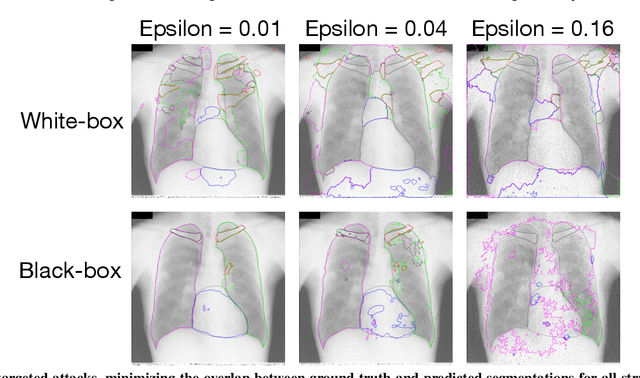
Abstract:Adversarial attacks consist in maliciously changing the input data to mislead the predictions of automated decision systems and are potentially a serious threat for automated medical image analysis. Previous studies have shown that it is possible to adversarially manipulate automated segmentations produced by neural networks in a targeted manner in the white-box attack setting. In this article, we studied the effectiveness of adversarial attacks in targeted modification of segmentations of anatomical structures in chest X-rays. Firstly, we experimented with using anatomically implausible shapes as targets for adversarial manipulation. We showed that, by adding almost imperceptible noise to the image, we can reliably force state-of-the-art neural networks to segment the heart as a heart symbol instead of its real anatomical shape. Moreover, such heart-shaping attack did not appear to require higher adversarial noise level than an untargeted attack based the same attack method. Secondly, we attempted to explore the limits of adversarial manipulation of segmentations. For that, we assessed the effectiveness of shrinking and enlarging segmentation contours for the three anatomical structures. We observed that adversarially extending segmentations of structures into regions with intensity and texture uncharacteristic for them presented a challenge to our attacks, as well as, in some cases, changing segmentations in ways that conflict with class adjacency priors learned by the target network. Additionally, we evaluated performances of the untargeted attacks and targeted heart attacks in the black-box attack scenario, using a surrogate network trained on a different subset of images. In both cases, the attacks were substantially less effective. We believe these findings bring novel insights into the current capabilities and limits of adversarial attacks for semantic segmentation.
Crowdsourcing Airway Annotations in Chest Computed Tomography Images
Nov 20, 2020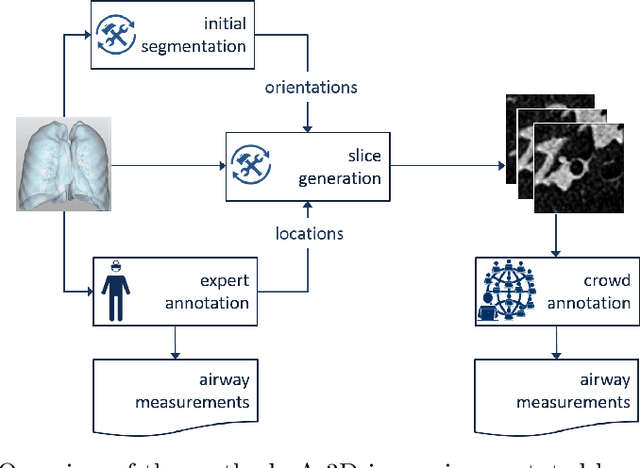
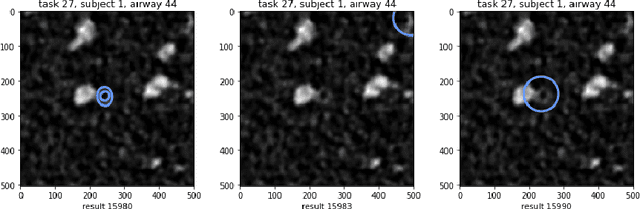
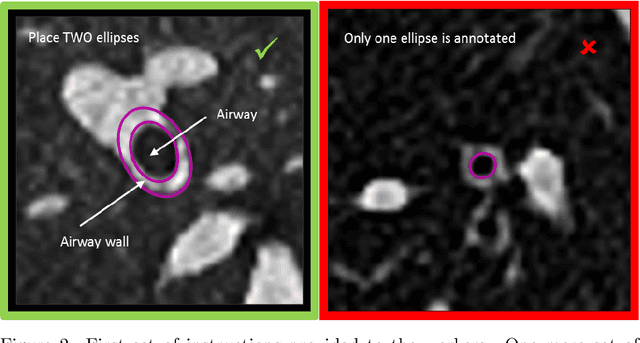
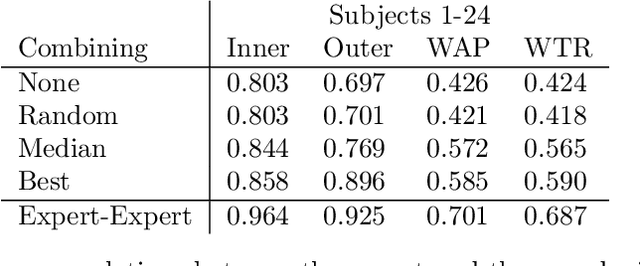

Abstract:Measuring airways in chest computed tomography (CT) scans is important for characterizing diseases such as cystic fibrosis, yet very time-consuming to perform manually. Machine learning algorithms offer an alternative, but need large sets of annotated scans for good performance. We investigate whether crowdsourcing can be used to gather airway annotations. We generate image slices at known locations of airways in 24 subjects and request the crowd workers to outline the airway lumen and airway wall. After combining multiple crowd workers, we compare the measurements to those made by the experts in the original scans. Similar to our preliminary study, a large portion of the annotations were excluded, possibly due to workers misunderstanding the instructions. After excluding such annotations, moderate to strong correlations with the expert can be observed, although these correlations are slightly lower than inter-expert correlations. Furthermore, the results across subjects in this study are quite variable. Although the crowd has potential in annotating airways, further development is needed for it to be robust enough for gathering annotations in practice. For reproducibility, data and code are available online: \url{http://github.com/adriapr/crowdairway.git}.
Medical Imaging with Deep Learning: MIDL 2020 -- Short Paper Track
Jun 29, 2020Abstract:This compendium gathers all the accepted extended abstracts from the Third International Conference on Medical Imaging with Deep Learning (MIDL 2020), held in Montreal, Canada, 6-9 July 2020. Note that only accepted extended abstracts are listed here, the Proceedings of the MIDL 2020 Full Paper Track are published in the Proceedings of Machine Learning Research (PMLR).
Region-of-interest guided Supervoxel Inpainting for Self-supervision
Jun 26, 2020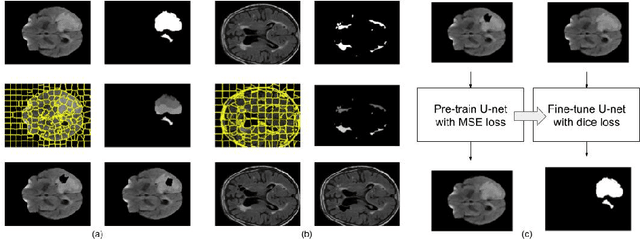
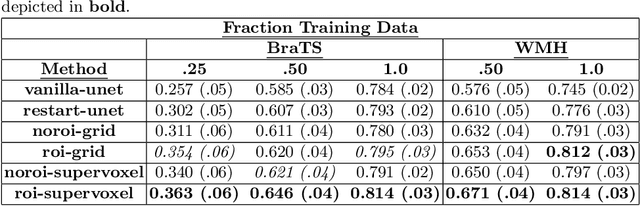
Abstract:Self-supervised learning has proven to be invaluable in making best use of all of the available data in biomedical image segmentation. One particularly simple and effective mechanism to achieve self-supervision is inpainting, the task of predicting arbitrary missing areas based on the rest of an image. In this work, we focus on image inpainting as the self-supervised proxy task, and propose two novel structural changes to further enhance the performance of a deep neural network. We guide the process of generating images to inpaint by using supervoxel-based masking instead of random masking, and also by focusing on the area to be segmented in the primary task, which we term as the region-of-interest. We postulate that these additions force the network to learn semantics that are more attuned to the primary task, and test our hypotheses on two applications: brain tumour and white matter hyperintensities segmentation. We empirically show that our proposed approach consistently outperforms both supervised CNNs, without any self-supervision, and conventional inpainting-based self-supervision methods on both large and small training set sizes.
DS6: Deformation-aware learning for small vessel segmentation with small, imperfectly labeled dataset
Jun 18, 2020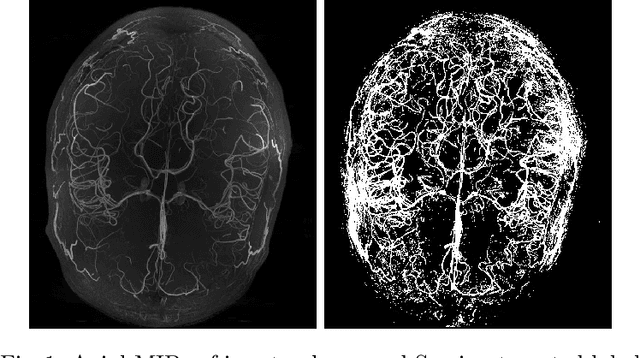
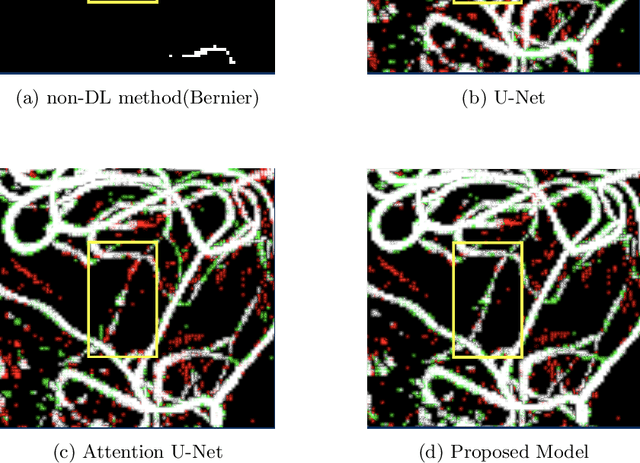
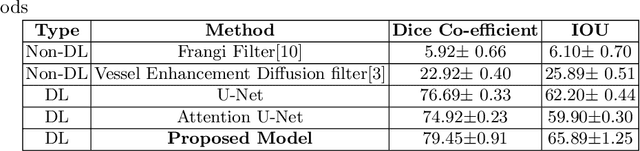


Abstract:Originating from the initial segment of the middle cerebral artery of the human brain, Lenticulostriate Arteries (LSA) are a collection of perforating vessels that supply blood to the basal ganglia region. With the advancement of 7 Tesla scanner, we are able to detect these LSA which are linked to Small Vessel Diseases(SVD) and potentially a cause for neurodegenerative diseases. Segmentation of LSA with traditional approaches like Frangi or semi-automated/manual techniques can depict medium to large vessels but fail to depict the small vessels. Also, semi-automated/manual approaches are time-consuming. In this paper, we put forth a study that incorporates deep learning techniques to automatically segment these LSA using 3D 7 Tesla Time-of-fight Magnetic Resonance Angiogram images. The algorithm is trained and evaluated on a small dataset of 11 volumes. Deep learning models based on Multi-Scale Supervision U-Net accompanied by elastic deformations to give equivariance to the model, were utilized for the vessel segmentation using semi-automated labeled images. We make a qualitative analysis of the output with the original image and also on imperfect semi-manual labels to confirm the presence and continuity of small vessels.
Adversarial Attack Vulnerability of Medical Image Analysis Systems: Unexplored Factors
Jun 12, 2020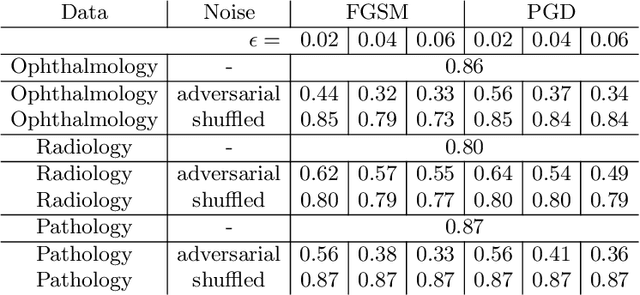
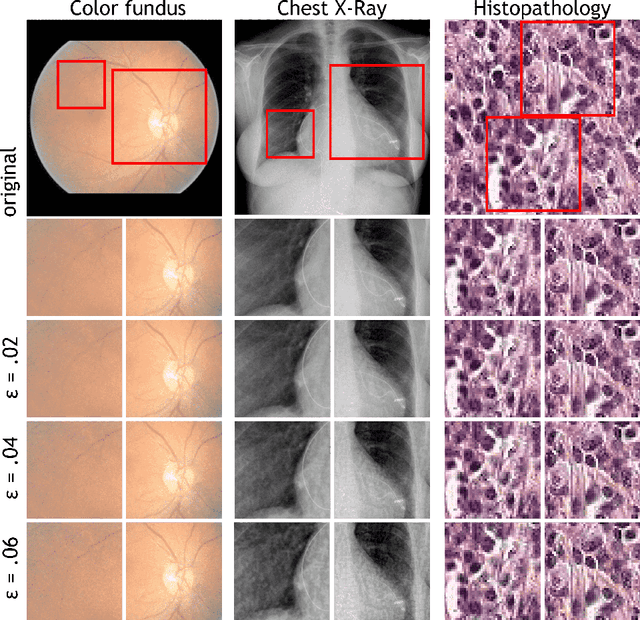
Abstract:Adversarial attacks are considered a potentially serious security threat for machine learning systems. Medical image analysis (MedIA) systems have recently been argued to be particularly vulnerable to adversarial attacks due to strong financial incentives. In this paper, we study several previously unexplored factors affecting adversarial attack vulnerability of deep learning MedIA systems in three medical domains: ophthalmology, radiology and pathology. Firstly, we study the effect of varying the degree of adversarial perturbation on the attack performance and its visual perceptibility. Secondly, we study how pre-training on a public dataset (ImageNet) affects the models' vulnerability to attacks. Thirdly, we study the influence of data and model architecture disparity between target and attacker models. Our experiments show that the degree of perturbation significantly affects both performance and human perceptibility of attacks. Pre-training may dramatically increase the transfer of adversarial examples; the larger the performance gain achieved by pre-training, the larger the transfer. Finally, disparity in data and/or model architecture between target and attacker models substantially decreases the success of attacks. We believe that these factors should be considered when designing cybersecurity-critical MedIA systems, as well as kept in mind when evaluating their vulnerability to adversarial attacks.
Spectral Data Augmentation Techniques to quantify Lung Pathology from CT-images
Apr 24, 2020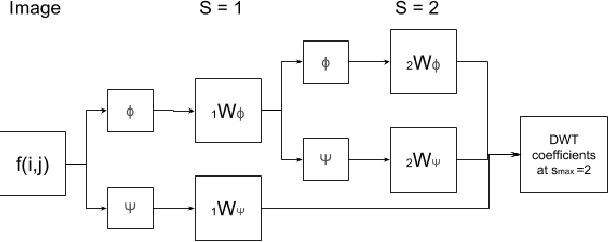
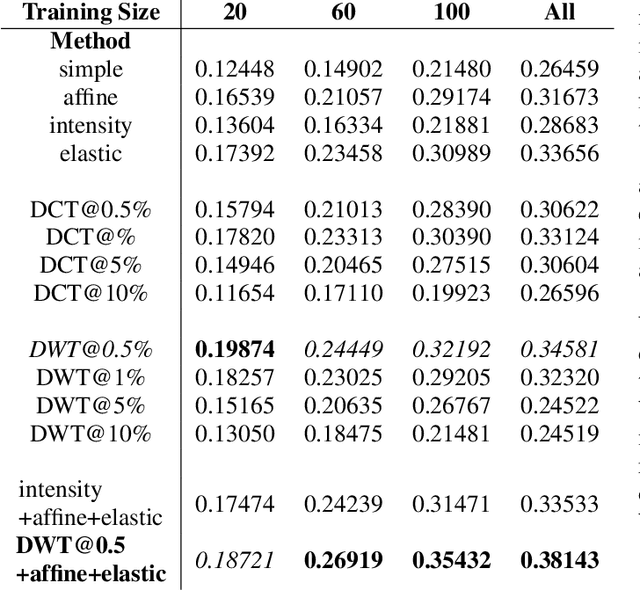
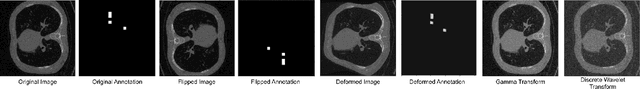
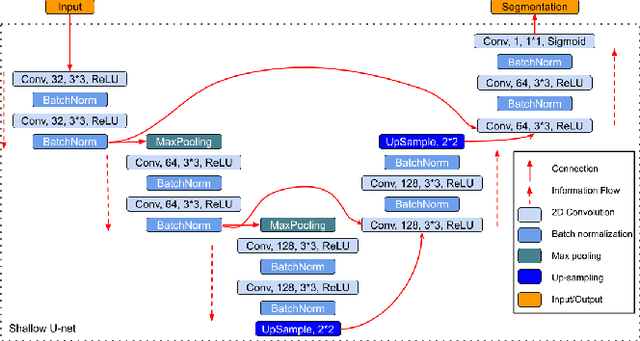
Abstract:Data augmentation is of paramount importance in biomedical image processing tasks, characterized by inadequate amounts of labelled data, to best use all of the data that is present. In-use techniques range from intensity transformations and elastic deformations, to linearly combining existing data points to make new ones. In this work, we propose the use of spectral techniques for data augmentation, using the discrete cosine and wavelet transforms. We empirically evaluate our approaches on a CT texture analysis task to detect abnormal lung-tissue in patients with cystic fibrosis. Empirical experiments show that the proposed spectral methods perform favourably as compared to the existing methods. When used in combination with existing methods, our proposed approach can increase the relative minor class segmentation performance by 44.1% over a simple replication baseline.
When Weak Becomes Strong: Robust Quantification of White Matter Hyperintensities in Brain MRI scans
Apr 12, 2020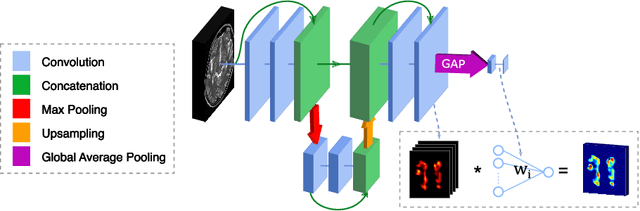
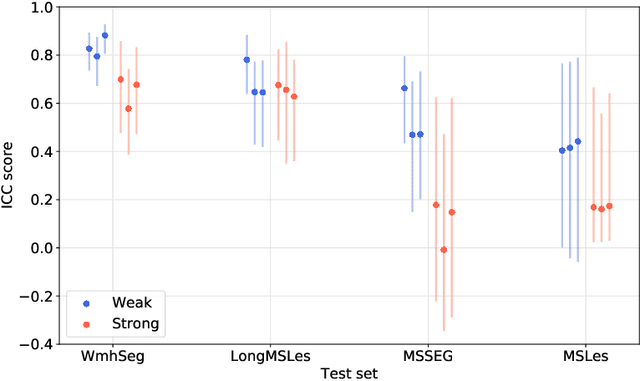
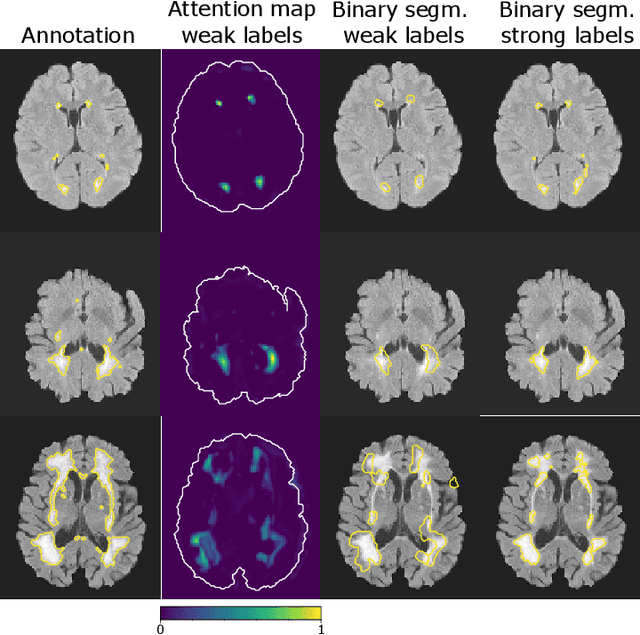
Abstract:To measure the volume of specific image structures, a typical approach is to first segment those structures using a neural network trained on voxel-wise (strong) labels and subsequently compute the volume from the segmentation. A more straightforward approach would be to predict the volume directly using a neural network based regression approach, trained on image-level (weak) labels indicating volume. In this article, we compared networks optimized with weak and strong labels, and study their ability to generalize to other datasets. We experimented with white matter hyperintensity (WMH) volume prediction in brain MRI scans. Neural networks were trained on a large local dataset and their performance was evaluated on four independent public datasets. We showed that networks optimized using only weak labels reflecting WMH volume generalized better for WMH volume prediction than networks optimized with voxel-wise segmentations of WMH. The attention maps of networks trained with weak labels did not seem to delineate WMHs, but highlighted instead areas with smooth contours around or near WMHs. By correcting for possible confounders we showed that networks trained on weak labels may have learnt other meaningful features that are more suited to generalization to unseen data. Our results suggest that for imaging biomarkers that can be derived from segmentations, training networks to predict the biomarker directly may provide more robust results than solving an intermediate segmentation step.
Automated Estimation of the Spinal Curvature via Spine Centerline Extraction with Ensembles of Cascaded Neural Networks
Dec 11, 2019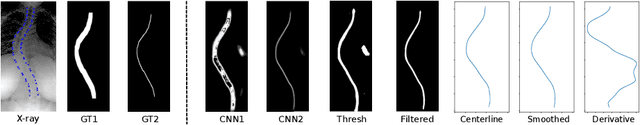

Abstract:Scoliosis is a condition defined by an abnormal spinal curvature. For diagnosis and treatment planning of scoliosis, spinal curvature can be estimated using Cobb angles. We propose an automated method for the estimation of Cobb angles from X-ray scans. First, the centerline of the spine was segmented using a cascade of two convolutional neural networks. After smoothing the centerline, Cobb angles were automatically estimated using the derivative of the centerline. We evaluated the results using the mean absolute error and the average symmetric mean absolute percentage error between the manual assessment by experts and the automated predictions. For optimization, we used 609 X-ray scans from the London Health Sciences Center, and for evaluation, we participated in the international challenge "Accurate Automated Spinal Curvature Estimation, MICCAI 2019" (100 scans). On the challenge's test set, we obtained an average symmetric mean absolute percentage error of 22.96.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge