Maria Skoularidou
CountsDiff: A Diffusion Model on the Natural Numbers for Generation and Imputation of Count-Based Data
Apr 04, 2026Abstract:Diffusion models have excelled at generative tasks for both continuous and token-based domains, but their application to discrete ordinal data remains underdeveloped. We present CountsDiff, a diffusion framework designed to natively model distributions on the natural numbers. CountsDiff extends the Blackout diffusion framework by simplifying its formulation through a direct parameterization in terms of a survival probability schedule and an explicit loss weighting. This introduces flexibility through design parameters with direct analogues in existing diffusion modeling frameworks. Beyond this reparameterization, CountsDiff introduces features from modern diffusion models, previously absent in counts-based domains, including continuous-time training, classifier-free guidance, and churn/remasking reverse dynamics that allow non-monotone reverse trajectories. We propose an initial instantiation of CountsDiff and validate it on natural image datasets (CIFAR-10, CelebA), exploring the effects of varying the introduced design parameters in a complex, well-studied, and interpretable data domain. We then highlight biological count assays as a natural use case, evaluating CountsDiff on single-cell RNA-seq imputation in a fetal cell and heart cell atlas. Remarkably, we find that even this simple instantiation matches or surpasses the performance of a state-of-the-art discrete generative model and leading RNA-seq imputation methods, while leaving substantial headroom for further gains through optimized design choices in future work.
Position Paper: Bayesian Deep Learning in the Age of Large-Scale AI
Feb 06, 2024

Abstract:In the current landscape of deep learning research, there is a predominant emphasis on achieving high predictive accuracy in supervised tasks involving large image and language datasets. However, a broader perspective reveals a multitude of overlooked metrics, tasks, and data types, such as uncertainty, active and continual learning, and scientific data, that demand attention. Bayesian deep learning (BDL) constitutes a promising avenue, offering advantages across these diverse settings. This paper posits that BDL can elevate the capabilities of deep learning. It revisits the strengths of BDL, acknowledges existing challenges, and highlights some exciting research avenues aimed at addressing these obstacles. Looking ahead, the discussion focuses on possible ways to combine large-scale foundation models with BDL to unlock their full potential.
Modeling Tabular data using Conditional GAN
Jul 01, 2019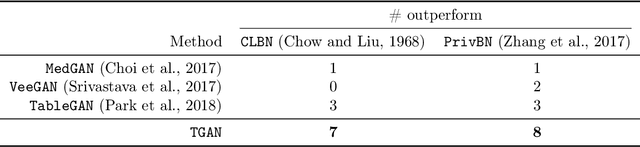
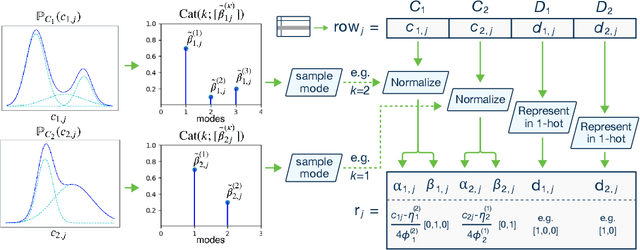
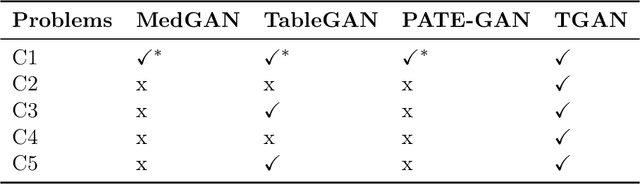
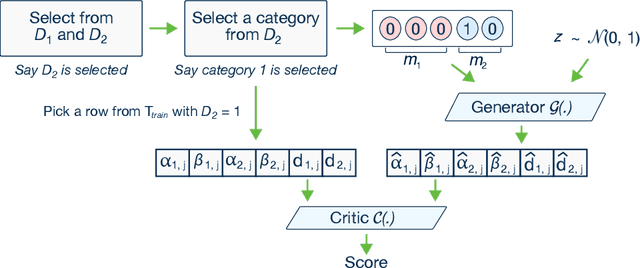
Abstract:Modeling the probability distribution of rows in tabular data and generating realistic synthetic data is a non-trivial task. Tabular data usually contains a mix of discrete and continuous columns. Continuous columns may have multiple modes whereas discrete columns are sometimes imbalanced making the modeling difficult. Existing statistical and deep neural network models fail to properly model this type of data. We design TGAN, which uses a conditional generative adversarial network to address these challenges. To aid in a fair and thorough comparison, we design a benchmark with 7 simulated and 8 real datasets and several Bayesian network baselines. TGAN outperforms Bayesian methods on most of the real datasets whereas other deep learning methods could not.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge