Marc Niethammer
PseudoClick: Interactive Image Segmentation with Click Imitation
Jul 12, 2022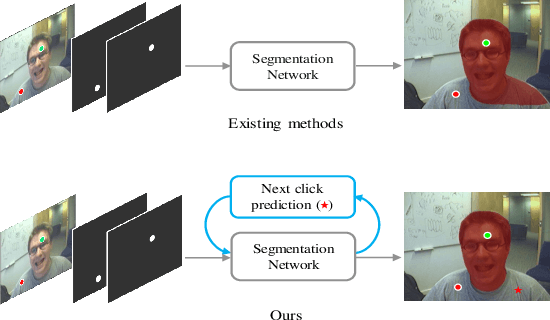
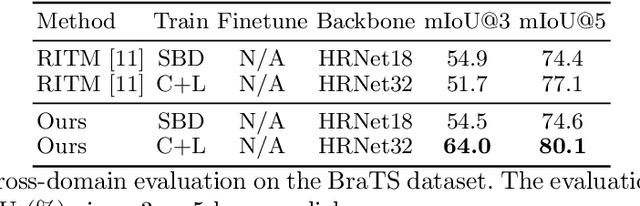
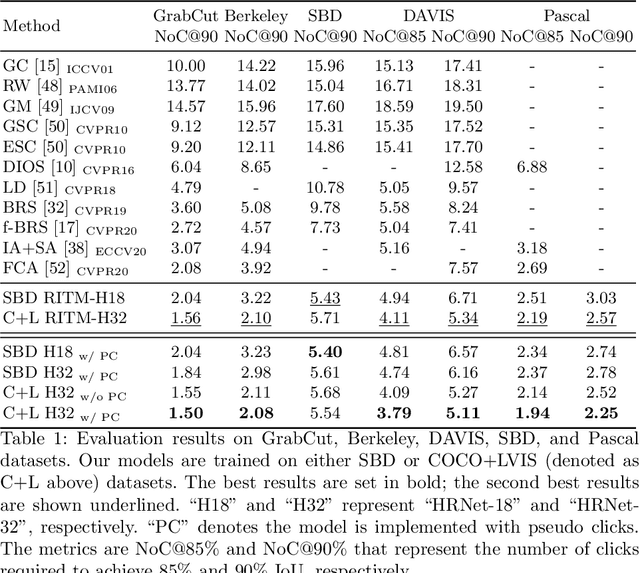
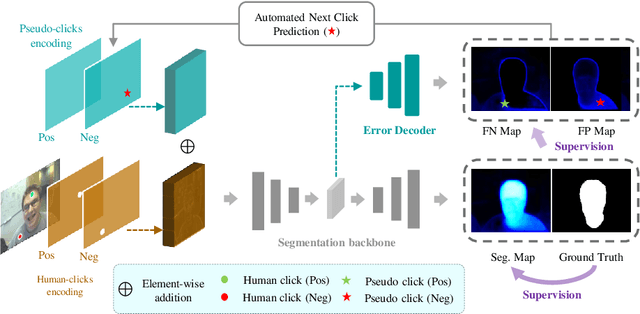

Abstract:The goal of click-based interactive image segmentation is to obtain precise object segmentation masks with limited user interaction, i.e., by a minimal number of user clicks. Existing methods require users to provide all the clicks: by first inspecting the segmentation mask and then providing points on mislabeled regions, iteratively. We ask the question: can our model directly predict where to click, so as to further reduce the user interaction cost? To this end, we propose {\PseudoClick}, a generic framework that enables existing segmentation networks to propose candidate next clicks. These automatically generated clicks, termed pseudo clicks in this work, serve as an imitation of human clicks to refine the segmentation mask.
GradICON: Approximate Diffeomorphisms via Gradient Inverse Consistency
Jun 13, 2022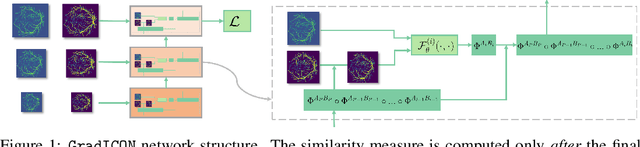
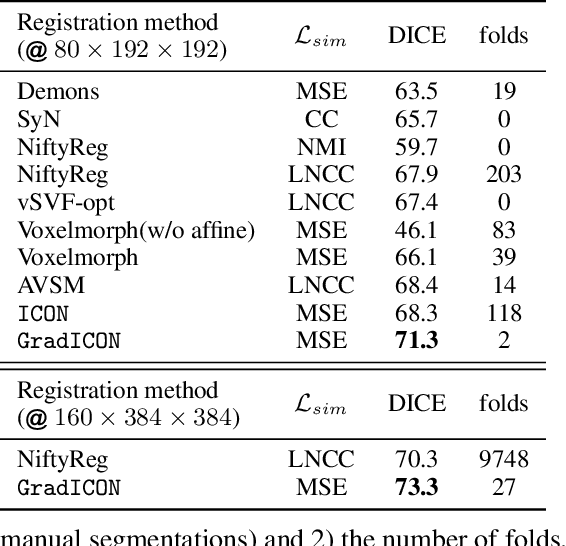
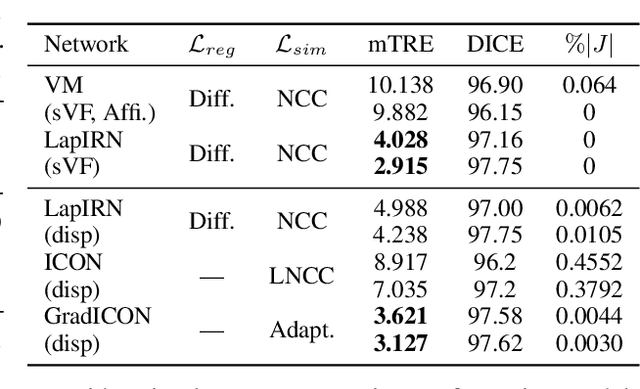
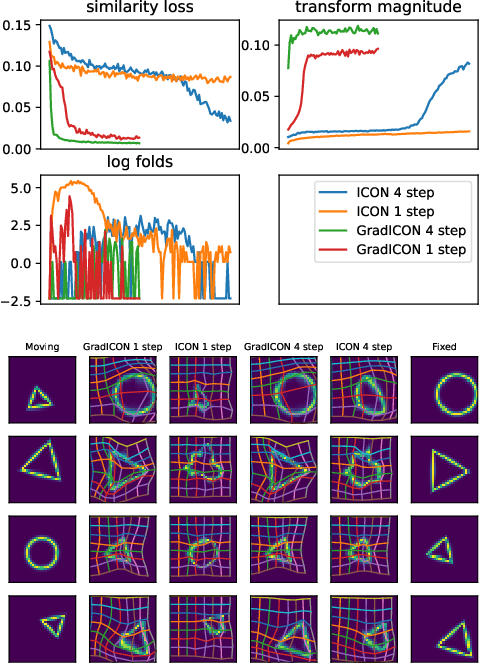
Abstract:Many registration approaches exist with early work focusing on optimization-based approaches for image pairs. Recent work focuses on deep registration networks to predict spatial transformations. In both cases, commonly used non-parametric registration models, which estimate transformation functions instead of low-dimensional transformation parameters, require choosing a suitable regularizer (to encourage smooth transformations) and its parameters. This makes models difficult to tune and restricts deformations to the deformation space permissible by the chosen regularizer. While deep-learning models for optical flow exist that do not regularize transformations and instead entirely rely on the data these might not yield diffeomorphic transformations which are desirable for medical image registration. In this work, we therefore develop GradICON building upon the unsupervised ICON deep-learning registration approach, which only uses inverse-consistency for regularization. However, in contrast to ICON, we prove and empirically verify that using a gradient inverse-consistency loss not only significantly improves convergence, but also results in a similar implicit regularization of the resulting transformation map. Synthetic experiments and experiments on magnetic resonance (MR) knee images and computed tomography (CT) lung images show the excellent performance of GradICON. We achieve state-of-the-art (SOTA) accuracy while retaining a simple registration formulation, which is practically important.
LiftReg: Limited Angle 2D/3D Deformable Registration
Mar 10, 2022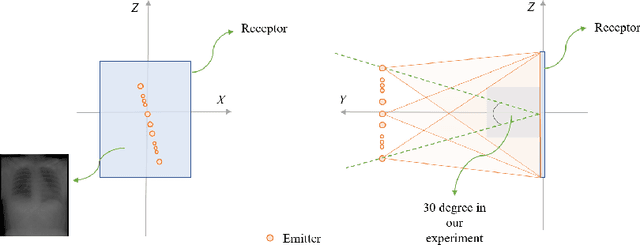
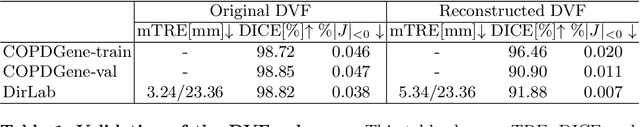
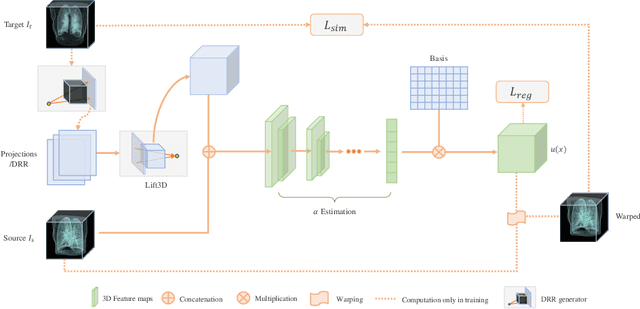
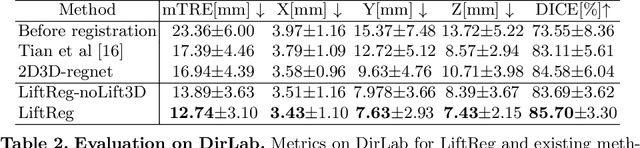
Abstract:We propose LiftReg, a 2D/3D deformable registration approach. LiftReg is a deep registration framework which is trained using sets of digitally reconstructed radiographs (DRR) and computed tomography (CT) image pairs. By using simulated training data, LiftReg can use a high-quality CT-CT image similarity measure, which helps the network to learn a high-quality deformation space. To further improve registration quality and to address the inherent depth ambiguities of very limited angle acquisitions, we propose to use features extracted from the backprojected 2D images and a statistical deformation model. We test our approach on the DirLab lung registration dataset and show that it outperforms an existing learning-based pairwise registration approach.
Fluid registration between lung CT and stationary chest tomosynthesis images
Mar 06, 2022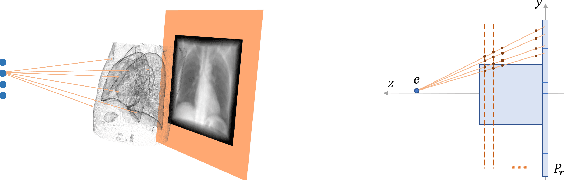
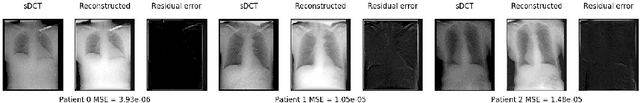
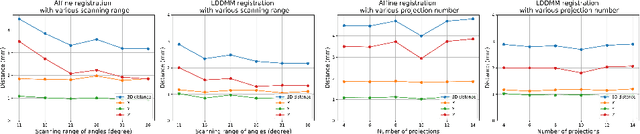
Abstract:Registration is widely used in image-guided therapy and image-guided surgery to estimate spatial correspondences between organs of interest between planning and treatment images. However, while high-quality computed tomography (CT) images are often available at planning time, limited angle acquisitions are frequently used during treatment because of radiation concerns or imaging time constraints. This requires algorithms to register CT images based on limited angle acquisitions. We, therefore, formulate a 3D/2D registration approach which infers a 3D deformation based on measured projections and digitally reconstructed radiographs of the CT. Most 3D/2D registration approaches use simple transformation models or require complex mathematical derivations to formulate the underlying optimization problem. Instead, our approach entirely relies on differentiable operations which can be combined with modern computational toolboxes supporting automatic differentiation. This then allows for rapid prototyping, integration with deep neural networks, and to support a variety of transformation models including fluid flow models. We demonstrate our approach for the registration between CT and stationary chest tomosynthesis (sDCT) images and show how it naturally leads to an iterative image reconstruction approach.
On Measuring Excess Capacity in Neural Networks
Feb 16, 2022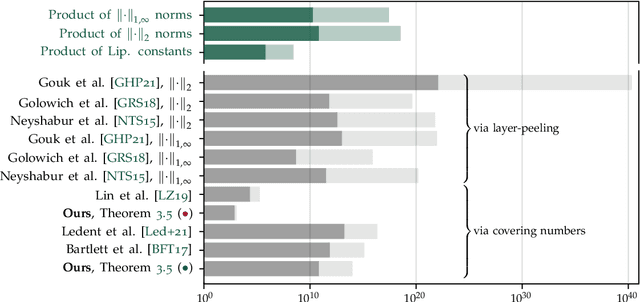
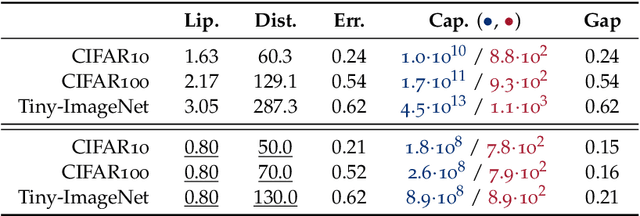
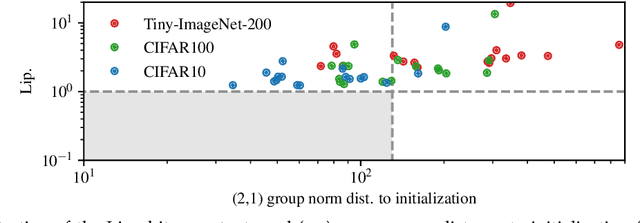
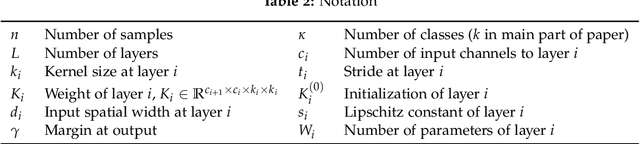
Abstract:We study the excess capacity of deep networks in the context of supervised classification. That is, given a capacity measure of the underlying hypothesis class -- in our case, Rademacher complexity -- how much can we (a-priori) constrain this class while maintaining an empirical error comparable to the unconstrained setting. To assess excess capacity in modern architectures, we first extend an existing generalization bound to accommodate function composition and addition, as well as the specific structure of convolutions. This then facilitates studying residual networks through the lens of the accompanying capacity measure. The key quantities driving this measure are the Lipschitz constants of the layers and the (2,1) group norm distance to the initializations of the convolution weights. We show that these quantities (1) can be kept surprisingly small and, (2) since excess capacity unexpectedly increases with task difficulty, this points towards an unnecessarily large capacity of unconstrained models.
Aladdin: Joint Atlas Building and Diffeomorphic Registration Learning with Pairwise Alignment
Feb 07, 2022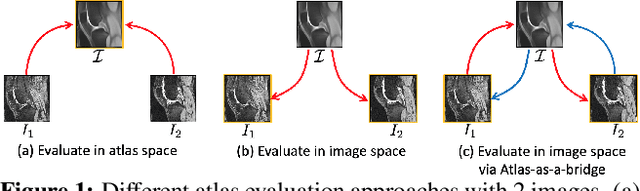
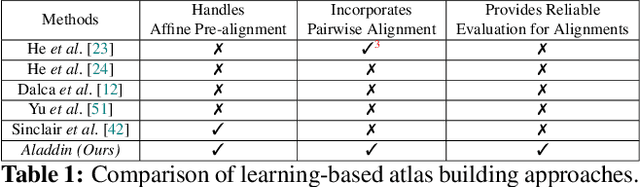
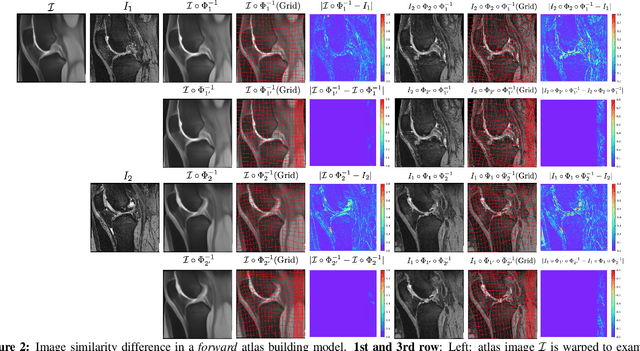
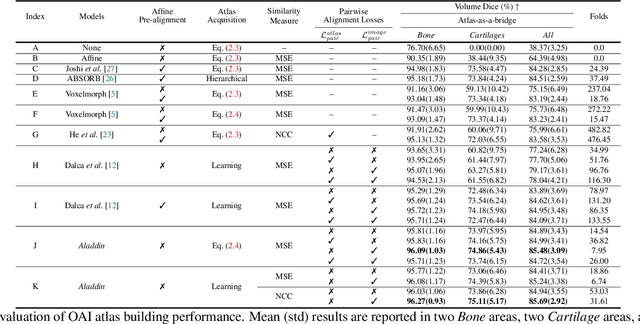
Abstract:Atlas building and image registration are important tasks for medical image analysis. Once one or multiple atlases from an image population have been constructed, commonly (1) images are warped into an atlas space to study intra-subject or inter-subject variations or (2) a possibly probabilistic atlas is warped into image space to assign anatomical labels. Atlas estimation and nonparametric transformations are computationally expensive as they usually require numerical optimization. Additionally, previous approaches for atlas building often define similarity measures between a fuzzy atlas and each individual image, which may cause alignment difficulties because a fuzzy atlas does not exhibit clear anatomical structures in contrast to the individual images. This work explores using a convolutional neural network (CNN) to jointly predict the atlas and a stationary velocity field (SVF) parameterization for diffeomorphic image registration with respect to the atlas. Our approach does not require affine pre-registrations and utilizes pairwise image alignment losses to increase registration accuracy. We evaluate our model on 3D knee magnetic resonance images (MRI) from the OAI-ZIB dataset. Our results show that the proposed framework achieves better performance than other state-of-the-art image registration algorithms, allows for end-to-end training, and for fast inference at test time.
Deep Decomposition for Stochastic Normal-Abnormal Transport
Nov 29, 2021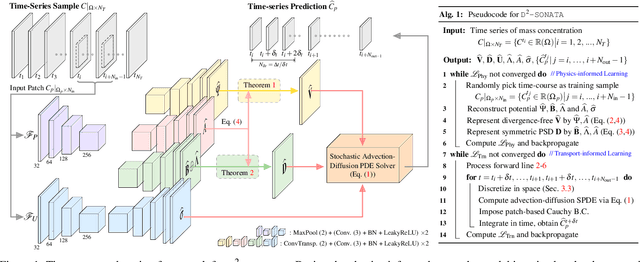
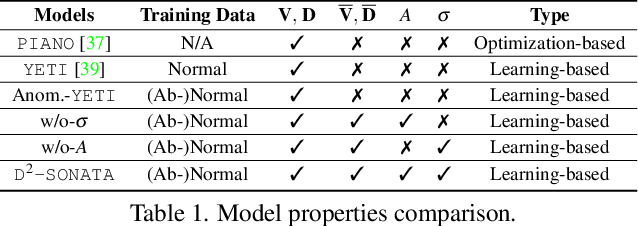
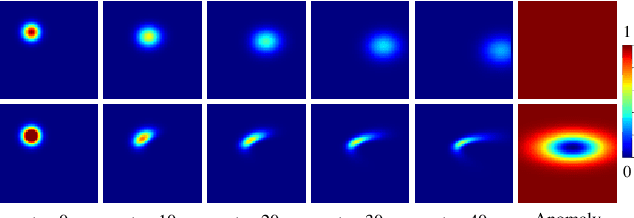
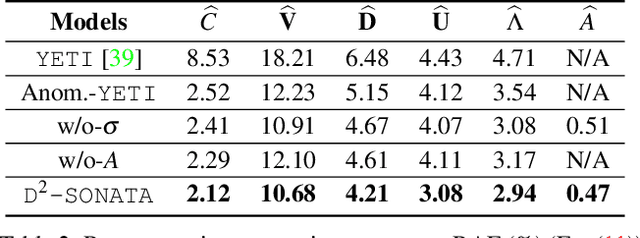
Abstract:Advection-diffusion equations describe a large family of natural transport processes, e.g., fluid flow, heat transfer, and wind transport. They are also used for optical flow and perfusion imaging computations. We develop a machine learning model, D^2-SONATA, built upon a stochastic advection-diffusion equation, which predicts the velocity and diffusion fields that drive 2D/3D image time-series of transport. In particular, our proposed model incorporates a model of transport atypicality, which isolates abnormal differences between expected normal transport behavior and the observed transport. In a medical context such a normal-abnormal decomposition can be used, for example, to quantify pathologies. Specifically, our model identifies the advection and diffusion contributions from the transport time-series and simultaneously predicts an anomaly value field to provide a decomposition into normal and abnormal advection and diffusion behavior. To achieve improved estimation performance for the velocity and diffusion-tensor fields underlying the advection-diffusion process and for the estimation of the anomaly fields, we create a 2D/3D anomaly-encoded advection-diffusion simulator, which allows for supervised learning. We further apply our model on a brain perfusion dataset from ischemic stroke patients via transfer learning. Extensive comparisons demonstrate that our model successfully distinguishes stroke lesions (abnormal) from normal brain regions, while reconstructing the underlying velocity and diffusion tensor fields.
Accurate Point Cloud Registration with Robust Optimal Transport
Nov 01, 2021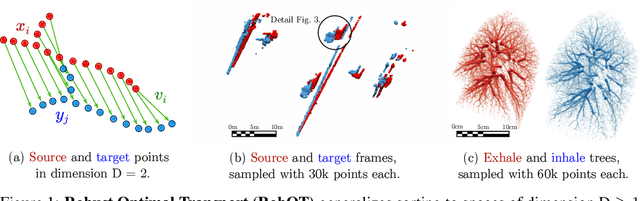
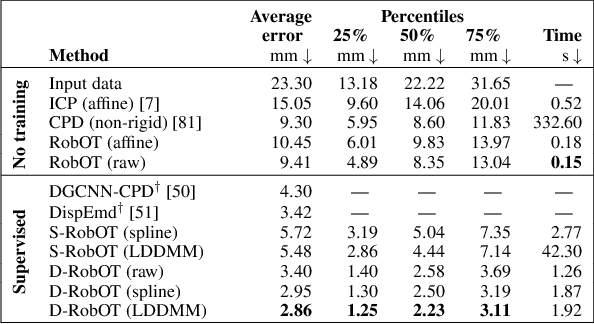
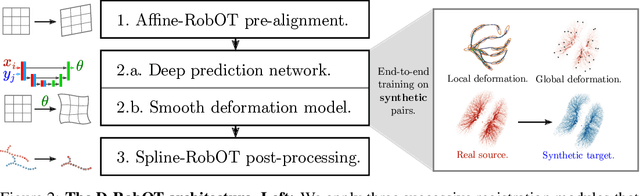
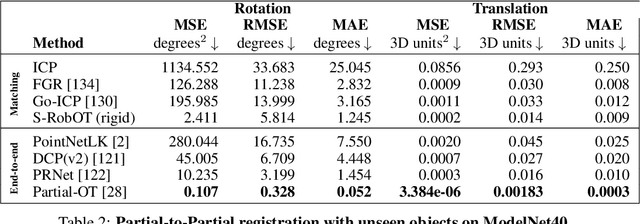
Abstract:This work investigates the use of robust optimal transport (OT) for shape matching. Specifically, we show that recent OT solvers improve both optimization-based and deep learning methods for point cloud registration, boosting accuracy at an affordable computational cost. This manuscript starts with a practical overview of modern OT theory. We then provide solutions to the main difficulties in using this framework for shape matching. Finally, we showcase the performance of transport-enhanced registration models on a wide range of challenging tasks: rigid registration for partial shapes; scene flow estimation on the Kitti dataset; and nonparametric registration of lung vascular trees between inspiration and expiration. Our OT-based methods achieve state-of-the-art results on Kitti and for the challenging lung registration task, both in terms of accuracy and scalability. We also release PVT1010, a new public dataset of 1,010 pairs of lung vascular trees with densely sampled points. This dataset provides a challenging use case for point cloud registration algorithms with highly complex shapes and deformations. Our work demonstrates that robust OT enables fast pre-alignment and fine-tuning for a wide range of registration models, thereby providing a new key method for the computer vision toolbox. Our code and dataset are available online at: https://github.com/uncbiag/robot.
ICON: Learning Regular Maps Through Inverse Consistency
May 21, 2021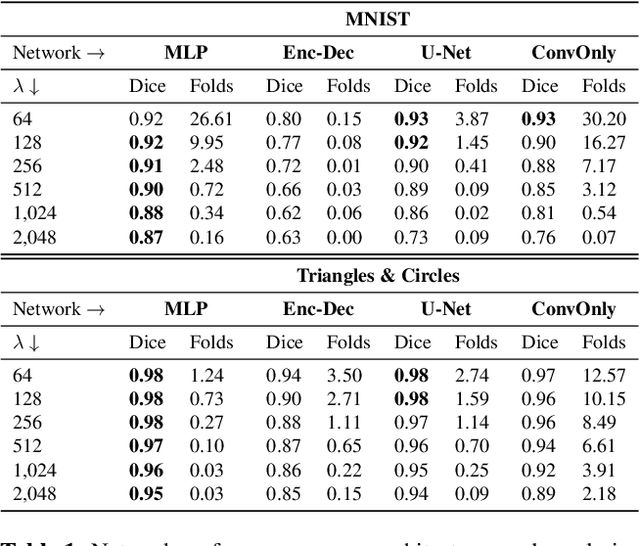
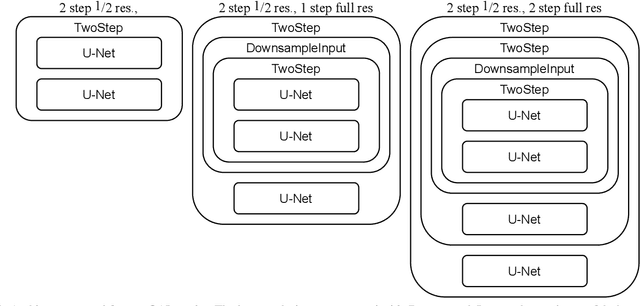
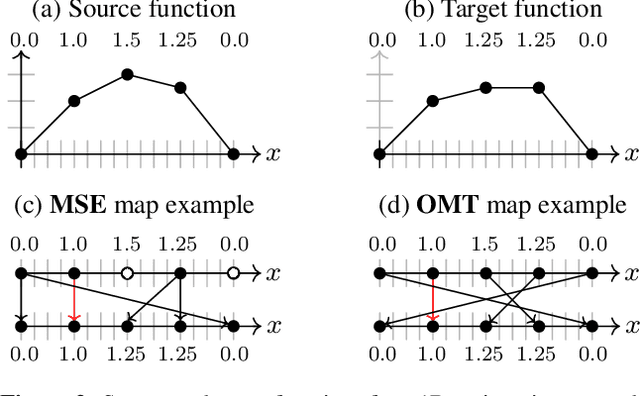
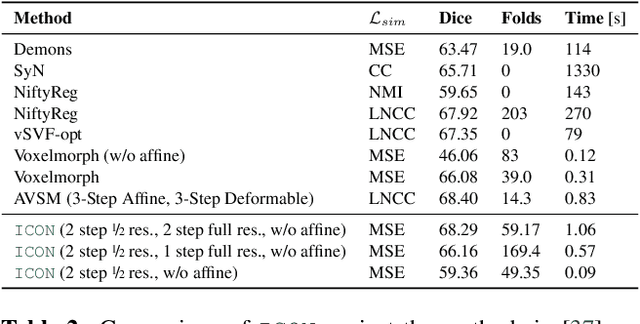
Abstract:Learning maps between data samples is fundamental. Applications range from representation learning, image translation and generative modeling, to the estimation of spatial deformations. Such maps relate feature vectors, or map between feature spaces. Well-behaved maps should be regular, which can be imposed explicitly or may emanate from the data itself. We explore what induces regularity for spatial transformations, e.g., when computing image registrations. Classical optimization-based models compute maps between pairs of samples and rely on an appropriate regularizer for well-posedness. Recent deep learning approaches have attempted to avoid using such regularizers altogether by relying on the sample population instead. We explore if it is possible to obtain spatial regularity using an inverse consistency loss only and elucidate what explains map regularity in such a context. We find that deep networks combined with an inverse consistency loss and randomized off-grid interpolation yield well behaved, approximately diffeomorphic, spatial transformations. Despite the simplicity of this approach, our experiments present compelling evidence, on both synthetic and real data, that regular maps can be obtained without carefully tuned explicit regularizers, while achieving competitive registration performance.
Dissecting Supervised Constrastive Learning
Feb 17, 2021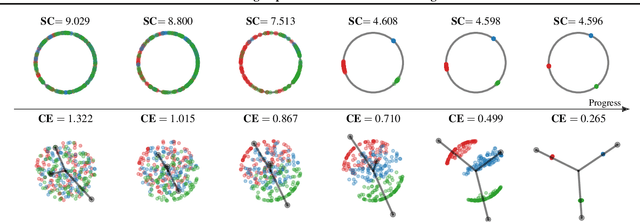
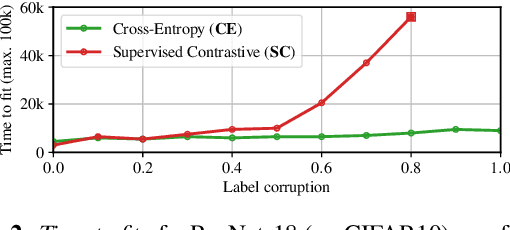
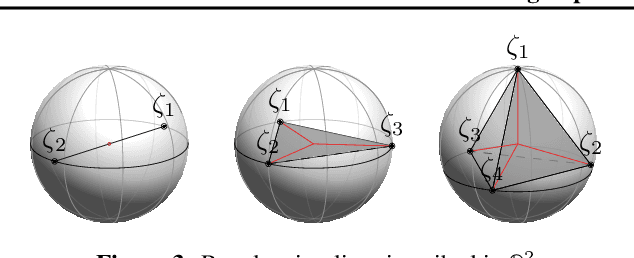
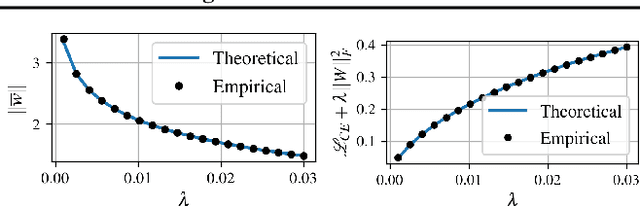
Abstract:Minimizing cross-entropy over the softmax scores of a linear map composed with a high-capacity encoder is arguably the most popular choice for training neural networks on supervised learning tasks. However, recent works show that one can directly optimize the encoder instead, to obtain equally (or even more) discriminative representations via a supervised variant of a contrastive objective. In this work, we address the question whether there are fundamental differences in the sought-for representation geometry in the output space of the encoder at minimal loss. Specifically, we prove, under mild assumptions, that both losses attain their minimum once the representations of each class collapse to the vertices of a regular simplex, inscribed in a hypersphere. We provide empirical evidence that this configuration is attained in practice and that reaching a close-to-optimal state typically indicates good generalization performance. Yet, the two losses show remarkably different optimization behavior. The number of iterations required to perfectly fit to data scales superlinearly with the amount of randomly flipped labels for the supervised contrastive loss. This is in contrast to the approximately linear scaling previously reported for networks trained with cross-entropy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge