Luke Kumar
Using Scaling Laws for Data Source Utility Estimation in Domain-Specific Pre-Training
Jul 29, 2025
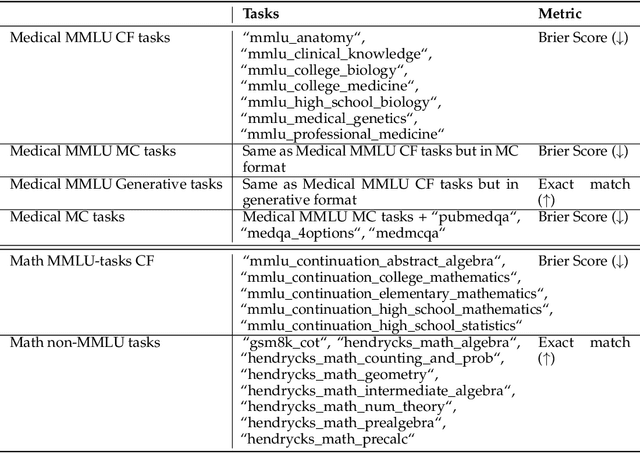
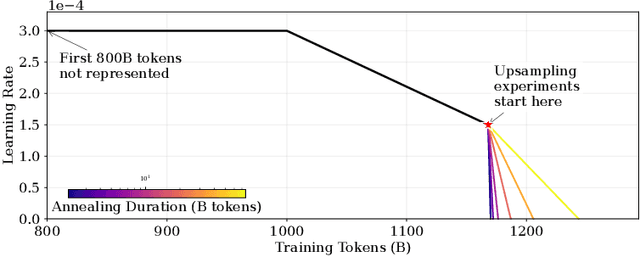
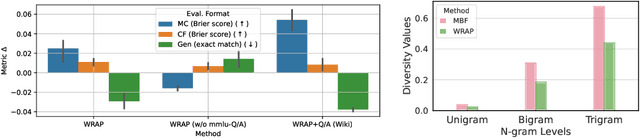
Abstract:We introduce a framework for optimizing domain-specific dataset construction in foundation model training. Specifically, we seek a cost-efficient way to estimate the quality of data sources (e.g. synthetically generated or filtered web data, etc.) in order to make optimal decisions about resource allocation for data sourcing from these sources for the stage two pre-training phase, aka annealing, with the goal of specializing a generalist pre-trained model to specific domains. Our approach extends the usual point estimate approaches, aka micro-annealing, to estimating scaling laws by performing multiple annealing runs of varying compute spent on data curation and training. This addresses a key limitation in prior work, where reliance on point estimates for data scaling decisions can be misleading due to the lack of rank invariance across compute scales -- a phenomenon we confirm in our experiments. By systematically analyzing performance gains relative to acquisition costs, we find that scaling curves can be estimated for different data sources. Such scaling laws can inform cost effective resource allocation across different data acquisition methods (e.g. synthetic data), data sources (e.g. user or web data) and available compute resources. We validate our approach through experiments on a pre-trained model with 7 billion parameters. We adapt it to: a domain well-represented in the pre-training data -- the medical domain, and a domain underrepresented in the pretraining corpora -- the math domain. We show that one can efficiently estimate the scaling behaviors of a data source by running multiple annealing runs, which can lead to different conclusions, had one used point estimates using the usual micro-annealing technique instead. This enables data-driven decision-making for selecting and optimizing data sources.
Predictive Representation Learning for Language Modeling
May 29, 2021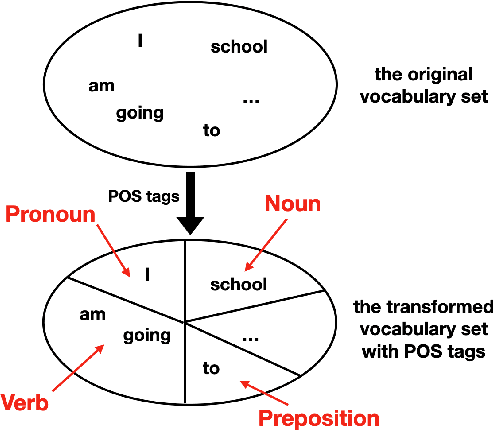
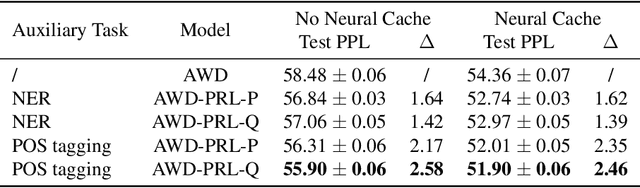
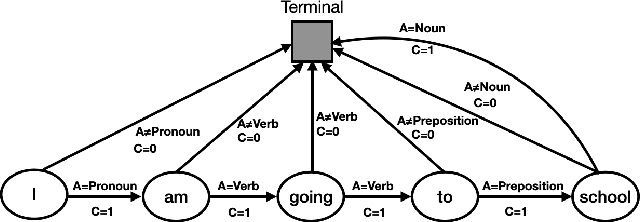
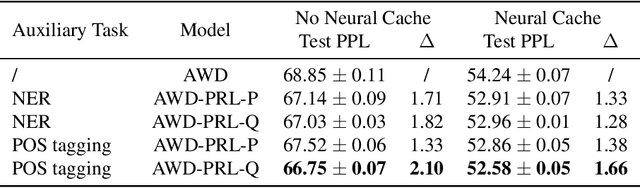
Abstract:To effectively perform the task of next-word prediction, long short-term memory networks (LSTMs) must keep track of many types of information. Some information is directly related to the next word's identity, but some is more secondary (e.g. discourse-level features or features of downstream words). Correlates of secondary information appear in LSTM representations even though they are not part of an \emph{explicitly} supervised prediction task. In contrast, in reinforcement learning (RL), techniques that explicitly supervise representations to predict secondary information have been shown to be beneficial. Inspired by that success, we propose Predictive Representation Learning (PRL), which explicitly constrains LSTMs to encode specific predictions, like those that might need to be learned implicitly. We show that PRL 1) significantly improves two strong language modeling methods, 2) converges more quickly, and 3) performs better when data is limited. Our work shows that explicitly encoding a simple predictive task facilitates the search for a more effective language model.
Gene Expression based Survival Prediction for Cancer Patients: A Topic Modeling Approach
Mar 25, 2019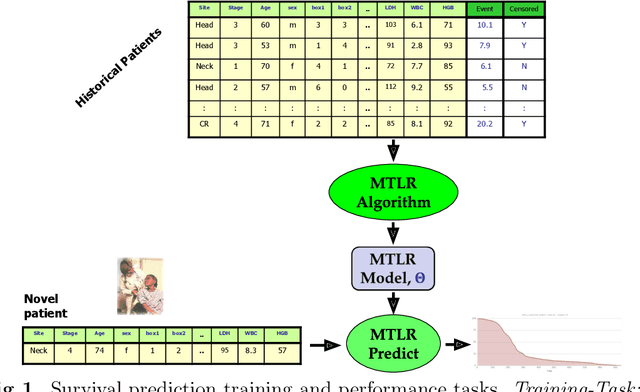
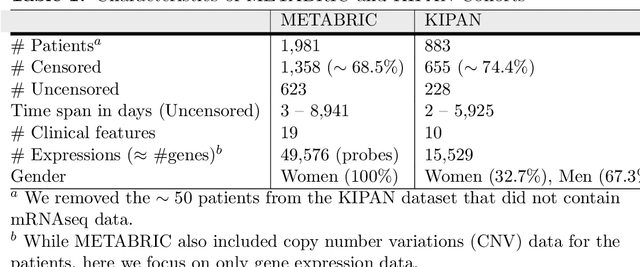
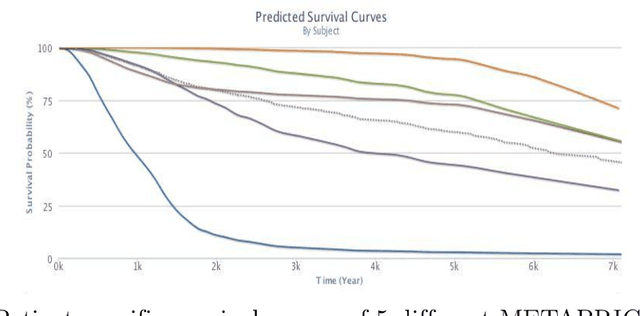
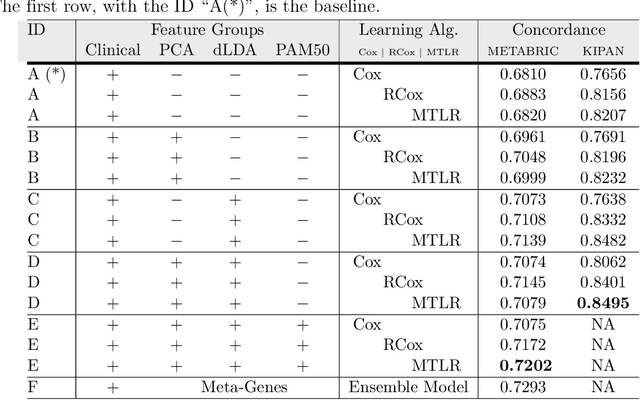
Abstract:Cancer is one of the leading cause of death, worldwide. Many believe that genomic data will enable us to better predict the survival time of these patients, which will lead to better, more personalized treatment options and patient care. As standard survival prediction models have a hard time coping with the high-dimensionality of such gene expression (GE) data, many projects use some dimensionality reduction techniques to overcome this hurdle. We introduce a novel methodology, inspired by topic modeling from the natural language domain, to derive expressive features from the high-dimensional GE data. There, a document is represented as a mixture over a relatively small number of topics, where each topic corresponds to a distribution over the words; here, to accommodate the heterogeneity of a patient's cancer, we represent each patient (~document) as a mixture over cancer-topics, where each cancer-topic is a mixture over GE values (~words). This required some extensions to the standard LDA model eg: to accommodate the "real-valued" expression values - leading to our novel "discretized" Latent Dirichlet Allocation (dLDA) procedure. We initially focus on the METABRIC dataset, which describes breast cancer patients using the r=49,576 GE values, from microarrays. Our results show that our approach provides survival estimates that are more accurate than standard models, in terms of the standard Concordance measure. We then validate this approach by running it on the Pan-kidney (KIPAN) dataset, over r=15,529 GE values - here using the mRNAseq modality - and find that it again achieves excellent results. In both cases, we also show that the resulting model is calibrated, using the recent "D-calibrated" measure. These successes, in two different cancer types and expression modalities, demonstrates the generality, and the effectiveness, of this approach.
On Generality and Knowledge Transferability in Cross-Domain Duplicate Question Detection for Heterogeneous Community Question Answering
Nov 15, 2018
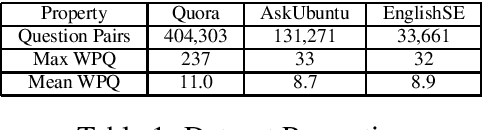
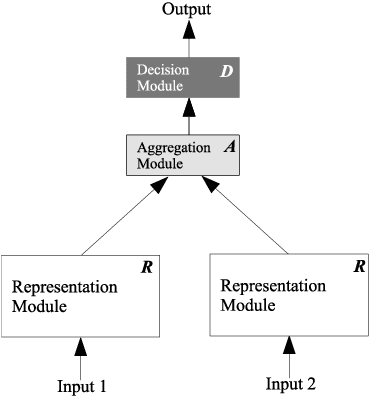
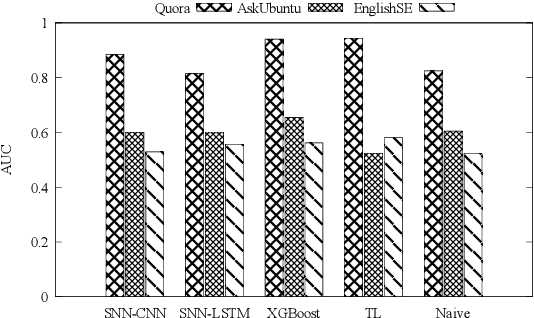
Abstract:Duplicate question detection is an ongoing challenge in community question answering because semantically equivalent questions can have significantly different words and structures. In addition, the identification of duplicate questions can reduce the resources required for retrieval, when the same questions are not repeated. This study compares the performance of deep neural networks and gradient tree boosting, and explores the possibility of domain adaptation with transfer learning to improve the under-performing target domains for the text-pair duplicates classification task, using three heterogeneous datasets: general-purpose Quora, technical Ask Ubuntu, and academic English Stack Exchange. Ultimately, our study exposes the alternative hypothesis that the meaning of a "duplicate" is not inherently general-purpose, but rather is dependent on the domain of learning, hence reducing the chance of transfer learning through adapting to the domain.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge