Lorenzo Torresani
Calibrating Histopathology Image Classifiers using Label Smoothing
Jan 28, 2022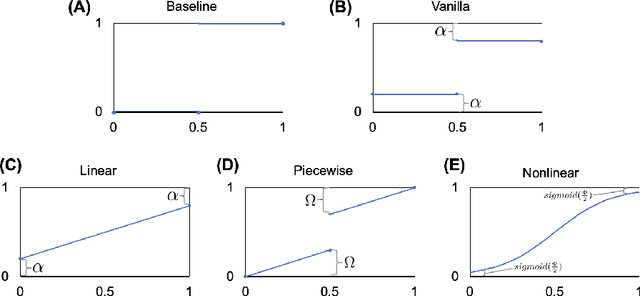
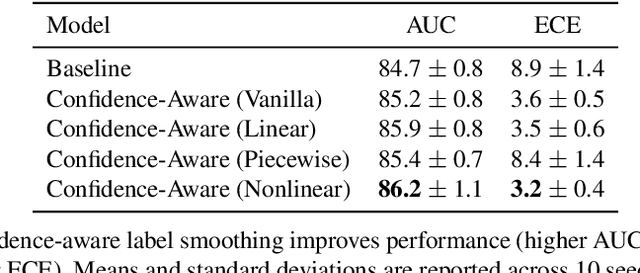
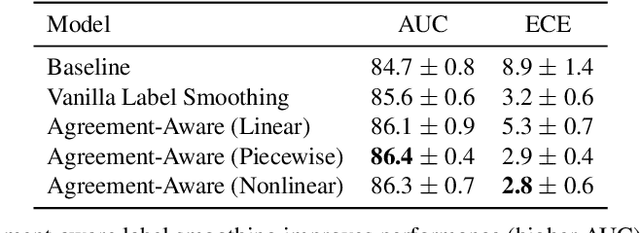
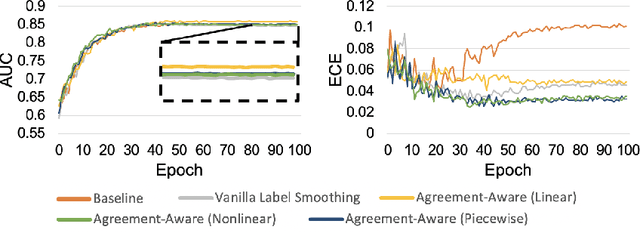

Abstract:The classification of histopathology images fundamentally differs from traditional image classification tasks because histopathology images naturally exhibit a range of diagnostic features, resulting in a diverse range of annotator agreement levels. However, examples with high annotator disagreement are often either assigned the majority label or discarded entirely when training histopathology image classifiers. This widespread practice often yields classifiers that do not account for example difficulty and exhibit poor model calibration. In this paper, we ask: can we improve model calibration by endowing histopathology image classifiers with inductive biases about example difficulty? We propose several label smoothing methods that utilize per-image annotator agreement. Though our methods are simple, we find that they substantially improve model calibration, while maintaining (or even improving) accuracy. For colorectal polyp classification, a common yet challenging task in gastrointestinal pathology, we find that our proposed agreement-aware label smoothing methods reduce calibration error by almost 70%. Moreover, we find that using model confidence as a proxy for annotator agreement also improves calibration and accuracy, suggesting that datasets without multiple annotators can still benefit from our proposed label smoothing methods via our proposed confidence-aware label smoothing methods. Given the importance of calibration (especially in histopathology image analysis), the improvements from our proposed techniques merit further exploration and potential implementation in other histopathology image classification tasks.
Learning To Recognize Procedural Activities with Distant Supervision
Jan 26, 2022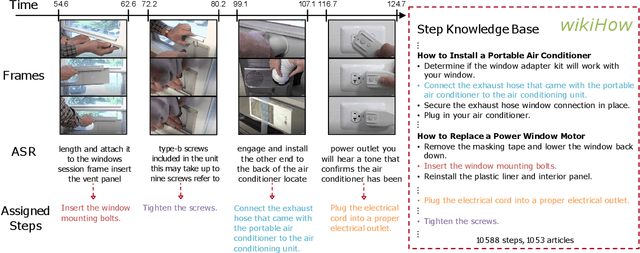
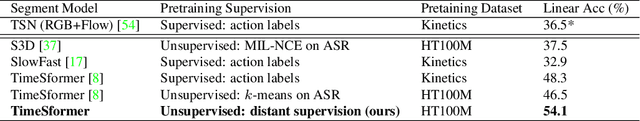
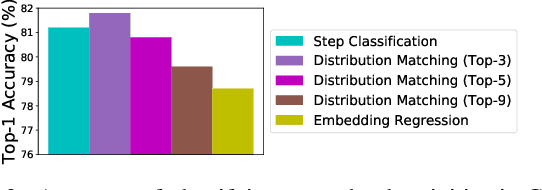

Abstract:In this paper we consider the problem of classifying fine-grained, multi-step activities (e.g., cooking different recipes, making disparate home improvements, creating various forms of arts and crafts) from long videos spanning up to several minutes. Accurately categorizing these activities requires not only recognizing the individual steps that compose the task but also capturing their temporal dependencies. This problem is dramatically different from traditional action classification, where models are typically optimized on videos that span only a few seconds and that are manually trimmed to contain simple atomic actions. While step annotations could enable the training of models to recognize the individual steps of procedural activities, existing large-scale datasets in this area do not include such segment labels due to the prohibitive cost of manually annotating temporal boundaries in long videos. To address this issue, we propose to automatically identify steps in instructional videos by leveraging the distant supervision of a textual knowledge base (wikiHow) that includes detailed descriptions of the steps needed for the execution of a wide variety of complex activities. Our method uses a language model to match noisy, automatically-transcribed speech from the video to step descriptions in the knowledge base. We demonstrate that video models trained to recognize these automatically-labeled steps (without manual supervision) yield a representation that achieves superior generalization performance on four downstream tasks: recognition of procedural activities, step classification, step forecasting and egocentric video classification.
Label Hallucination for Few-Shot Classification
Dec 06, 2021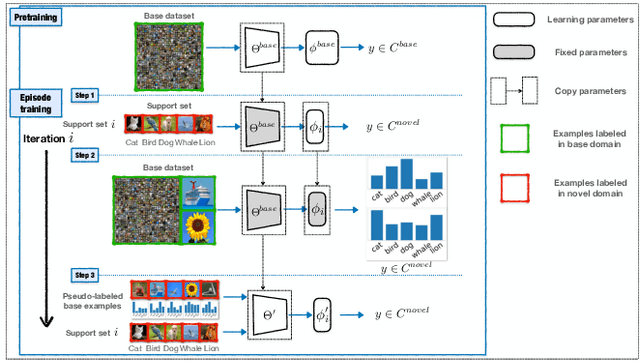
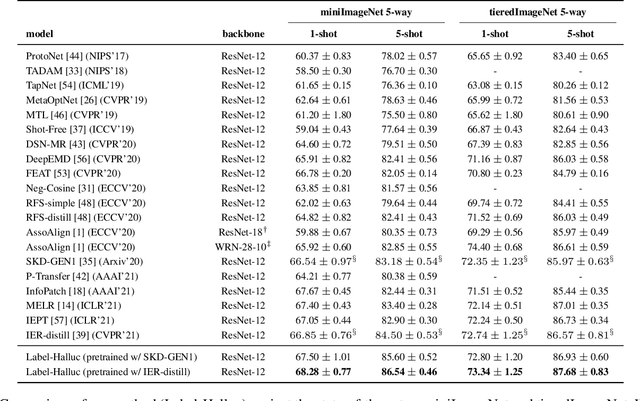
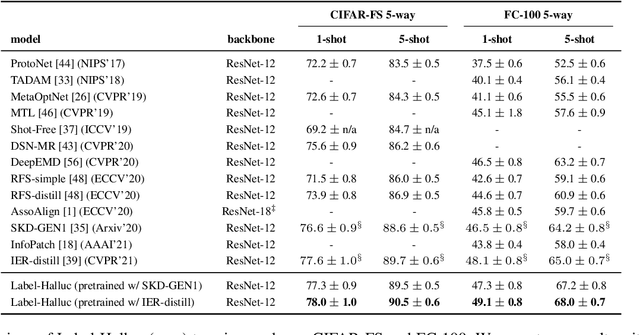
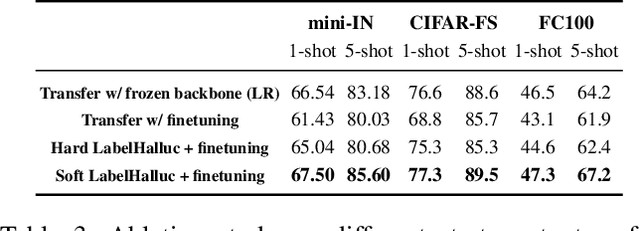
Abstract:Few-shot classification requires adapting knowledge learned from a large annotated base dataset to recognize novel unseen classes, each represented by few labeled examples. In such a scenario, pretraining a network with high capacity on the large dataset and then finetuning it on the few examples causes severe overfitting. At the same time, training a simple linear classifier on top of "frozen" features learned from the large labeled dataset fails to adapt the model to the properties of the novel classes, effectively inducing underfitting. In this paper we propose an alternative approach to both of these two popular strategies. First, our method pseudo-labels the entire large dataset using the linear classifier trained on the novel classes. This effectively "hallucinates" the novel classes in the large dataset, despite the novel categories not being present in the base database (novel and base classes are disjoint). Then, it finetunes the entire model with a distillation loss on the pseudo-labeled base examples, in addition to the standard cross-entropy loss on the novel dataset. This step effectively trains the network to recognize contextual and appearance cues that are useful for the novel-category recognition but using the entire large-scale base dataset and thus overcoming the inherent data-scarcity problem of few-shot learning. Despite the simplicity of the approach, we show that that our method outperforms the state-of-the-art on four well-established few-shot classification benchmarks.
Ego4D: Around the World in 3,000 Hours of Egocentric Video
Oct 13, 2021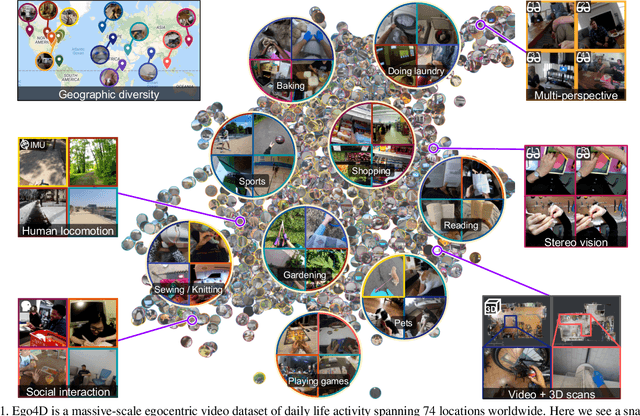
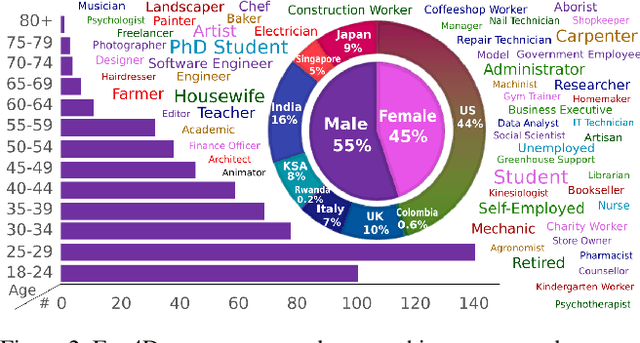

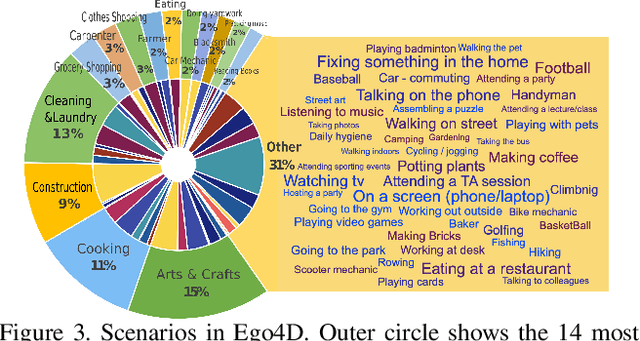
Abstract:We introduce Ego4D, a massive-scale egocentric video dataset and benchmark suite. It offers 3,025 hours of daily-life activity video spanning hundreds of scenarios (household, outdoor, workplace, leisure, etc.) captured by 855 unique camera wearers from 74 worldwide locations and 9 different countries. The approach to collection is designed to uphold rigorous privacy and ethics standards with consenting participants and robust de-identification procedures where relevant. Ego4D dramatically expands the volume of diverse egocentric video footage publicly available to the research community. Portions of the video are accompanied by audio, 3D meshes of the environment, eye gaze, stereo, and/or synchronized videos from multiple egocentric cameras at the same event. Furthermore, we present a host of new benchmark challenges centered around understanding the first-person visual experience in the past (querying an episodic memory), present (analyzing hand-object manipulation, audio-visual conversation, and social interactions), and future (forecasting activities). By publicly sharing this massive annotated dataset and benchmark suite, we aim to push the frontier of first-person perception. Project page: https://ego4d-data.org/
Long-Short Temporal Contrastive Learning of Video Transformers
Jul 08, 2021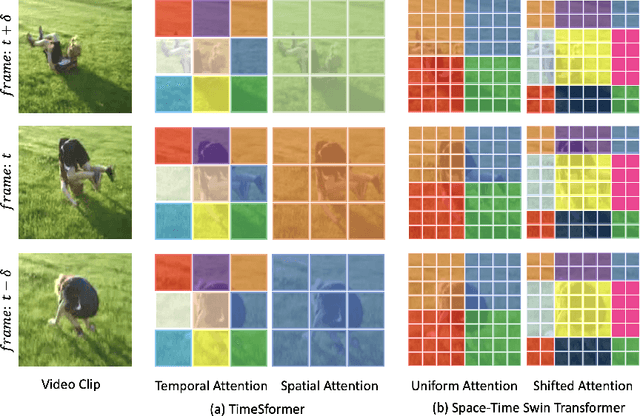
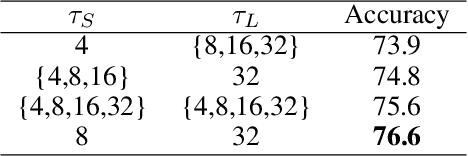

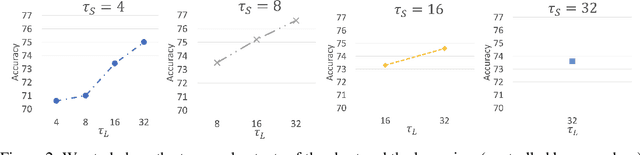
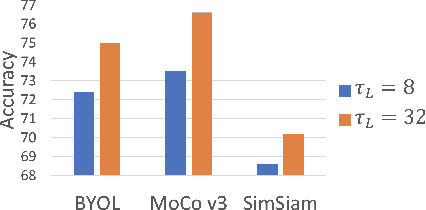
Abstract:Video transformers have recently emerged as a competitive alternative to 3D CNNs for video understanding. However, due to their large number of parameters and reduced inductive biases, these models require supervised pretraining on large-scale image datasets to achieve top performance. In this paper, we empirically demonstrate that self-supervised pretraining of video transformers on video-only datasets can lead to action recognition results that are on par or better than those obtained with supervised pretraining on large-scale image datasets, even massive ones such as ImageNet-21K. Since transformer-based models are effective at capturing dependencies over extended temporal spans, we propose a simple learning procedure that forces the model to match a long-term view to a short-term view of the same video. Our approach, named Long-Short Temporal Contrastive Learning (LSTCL), enables video transformers to learn an effective clip-level representation by predicting temporal context captured from a longer temporal extent. To demonstrate the generality of our findings, we implement and validate our approach under three different self-supervised contrastive learning frameworks (MoCo v3, BYOL, SimSiam) using two distinct video-transformer architectures, including an improved variant of the Swin Transformer augmented with space-time attention. We conduct a thorough ablation study and show that LSTCL achieves competitive performance on multiple video benchmarks and represents a convincing alternative to supervised image-based pretraining.
Beyond Short Clips: End-to-End Video-Level Learning with Collaborative Memories
Apr 02, 2021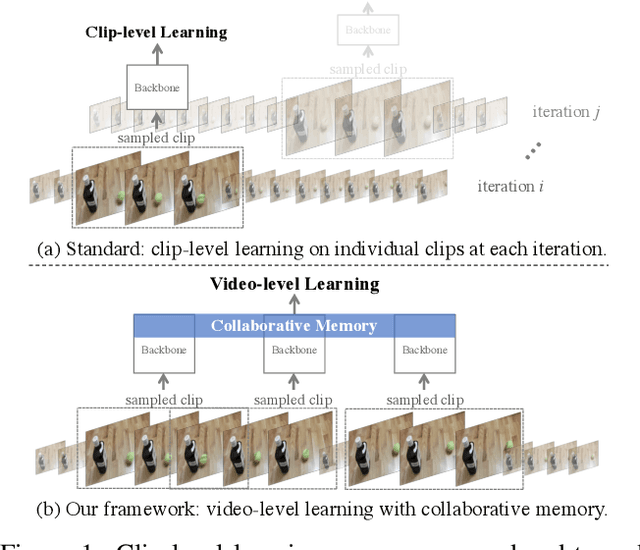
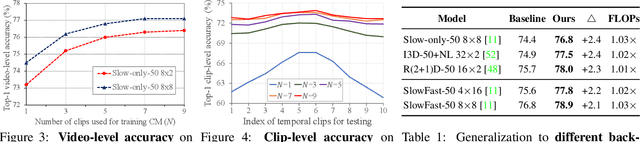
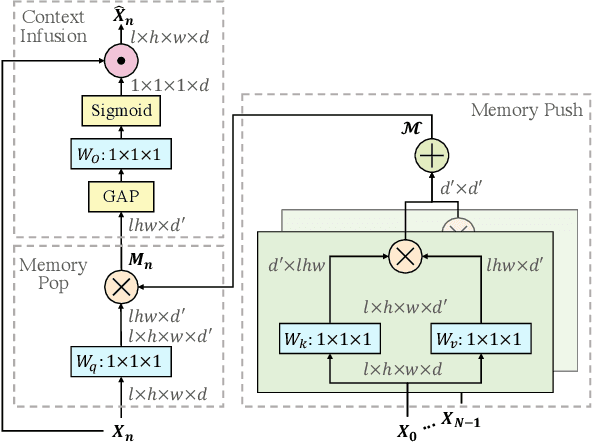
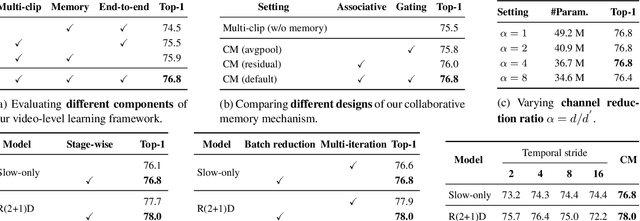
Abstract:The standard way of training video models entails sampling at each iteration a single clip from a video and optimizing the clip prediction with respect to the video-level label. We argue that a single clip may not have enough temporal coverage to exhibit the label to recognize, since video datasets are often weakly labeled with categorical information but without dense temporal annotations. Furthermore, optimizing the model over brief clips impedes its ability to learn long-term temporal dependencies. To overcome these limitations, we introduce a collaborative memory mechanism that encodes information across multiple sampled clips of a video at each training iteration. This enables the learning of long-range dependencies beyond a single clip. We explore different design choices for the collaborative memory to ease the optimization difficulties. Our proposed framework is end-to-end trainable and significantly improves the accuracy of video classification at a negligible computational overhead. Through extensive experiments, we demonstrate that our framework generalizes to different video architectures and tasks, outperforming the state of the art on both action recognition (e.g., Kinetics-400 & 700, Charades, Something-Something-V1) and action detection (e.g., AVA v2.1 & v2.2).
Is Space-Time Attention All You Need for Video Understanding?
Feb 24, 2021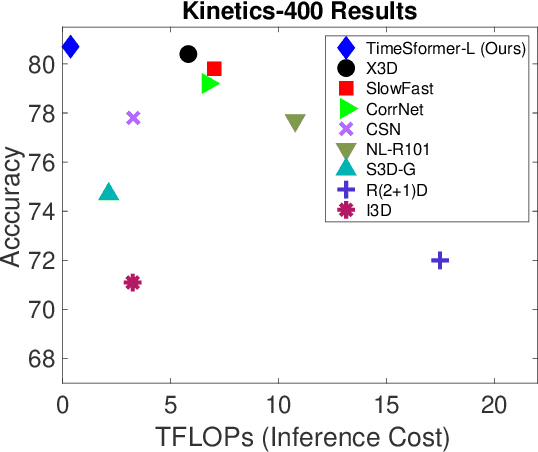
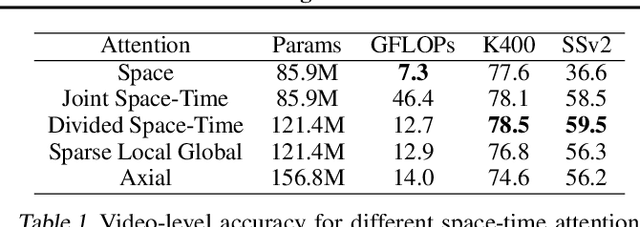
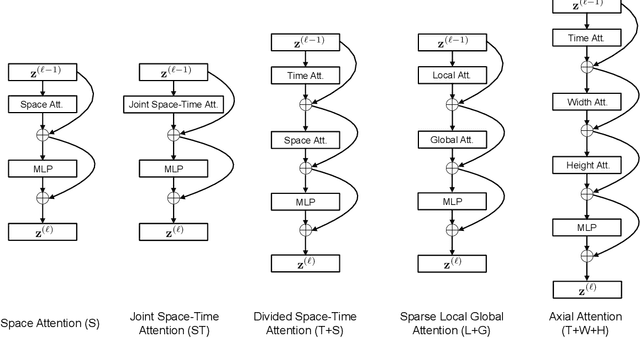
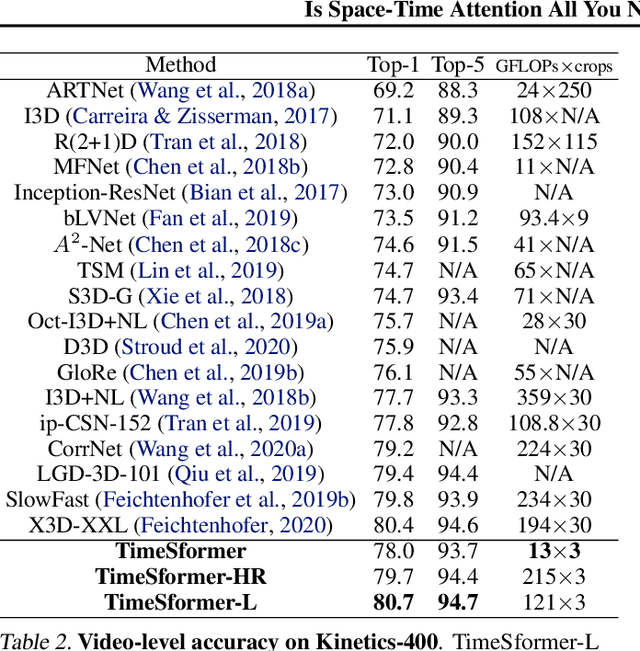
Abstract:We present a convolution-free approach to video classification built exclusively on self-attention over space and time. Our method, named "TimeSformer," adapts the standard Transformer architecture to video by enabling spatiotemporal feature learning directly from a sequence of frame-level patches. Our experimental study compares different self-attention schemes and suggests that "divided attention," where temporal attention and spatial attention are separately applied within each block, leads to the best video classification accuracy among the design choices considered. Despite the radically different design compared to the prominent paradigm of 3D convolutional architectures for video, TimeSformer achieves state-of-the-art results on several major action recognition benchmarks, including the best reported accuracy on Kinetics-400 and Kinetics-600. Furthermore, our model is faster to train and has higher test-time efficiency compared to competing architectures. Code and pretrained models will be made publicly available.
A Multi-View Approach To Audio-Visual Speaker Verification
Feb 11, 2021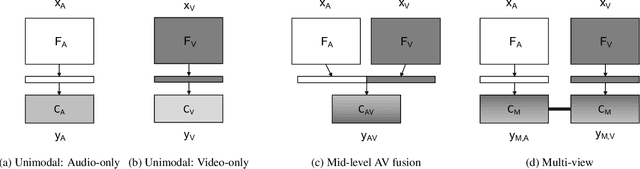
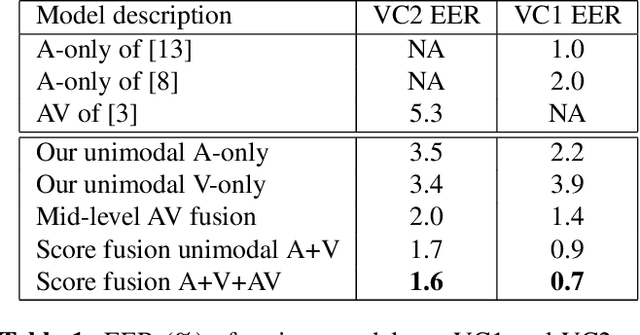

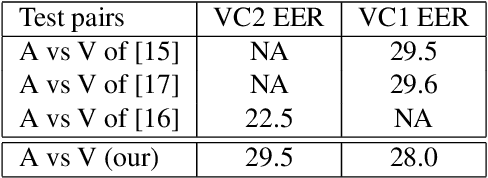
Abstract:Although speaker verification has conventionally been an audio-only task, some practical applications provide both audio and visual streams of input. In these cases, the visual stream provides complementary information and can often be leveraged in conjunction with the acoustics of speech to improve verification performance. In this study, we explore audio-visual approaches to speaker verification, starting with standard fusion techniques to learn joint audio-visual (AV) embeddings, and then propose a novel approach to handle cross-modal verification at test time. Specifically, we investigate unimodal and concatenation based AV fusion and report the lowest AV equal error rate (EER) of 0.7% on the VoxCeleb1 dataset using our best system. As these methods lack the ability to do cross-modal verification, we introduce a multi-view model which uses a shared classifier to map audio and video into the same space. This new approach achieves 28% EER on VoxCeleb1 in the challenging testing condition of cross-modal verification.
VX2TEXT: End-to-End Learning of Video-Based Text Generation From Multimodal Inputs
Jan 29, 2021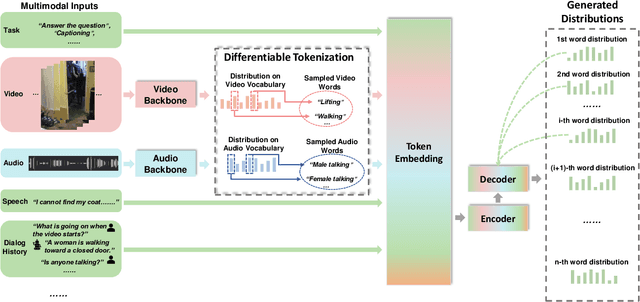
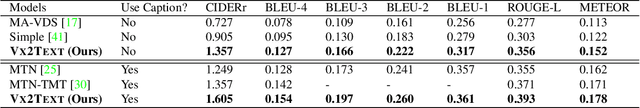
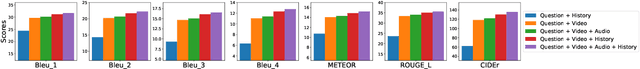

Abstract:We present \textsc{Vx2Text}, a framework for text generation from multimodal inputs consisting of video plus text, speech, or audio. In order to leverage transformer networks, which have been shown to be effective at modeling language, each modality is first converted into a set of language embeddings by a learnable tokenizer. This allows our approach to perform multimodal fusion in the language space, thus eliminating the need for ad-hoc cross-modal fusion modules. To address the non-differentiability of tokenization on continuous inputs (e.g., video or audio), we utilize a relaxation scheme that enables end-to-end training. Furthermore, unlike prior encoder-only models, our network includes an autoregressive decoder to generate open-ended text from the multimodal embeddings fused by the language encoder. This renders our approach fully generative and makes it directly applicable to different "video+$x$ to text" problems without the need to design specialized network heads for each task. The proposed framework is not only conceptually simple but also remarkably effective: experiments demonstrate that our approach based on a single architecture outperforms the state-of-the-art on three video-based text-generation tasks -- captioning, question answering and audio-visual scene-aware dialog.
A Petri Dish for Histopathology Image Analysis
Jan 29, 2021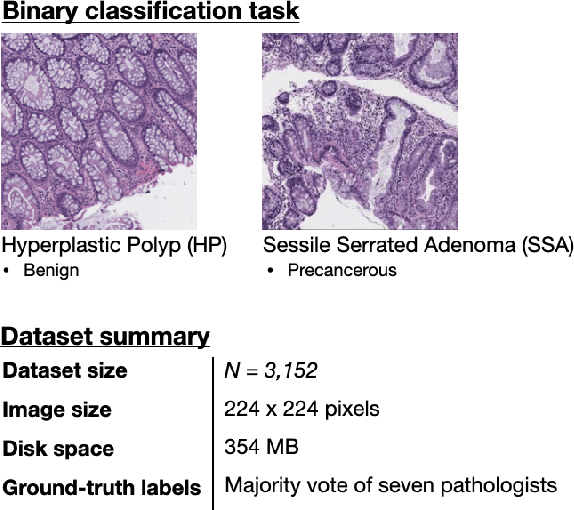
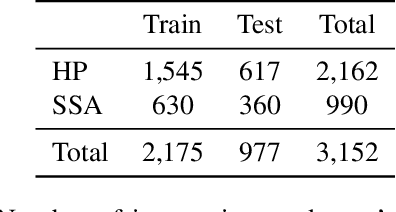
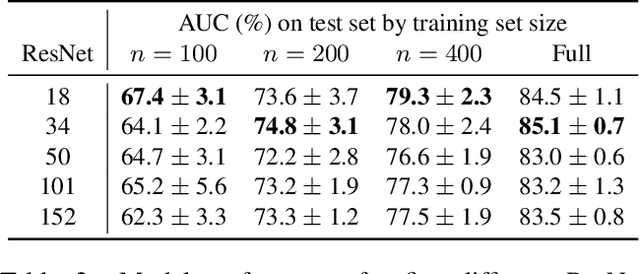
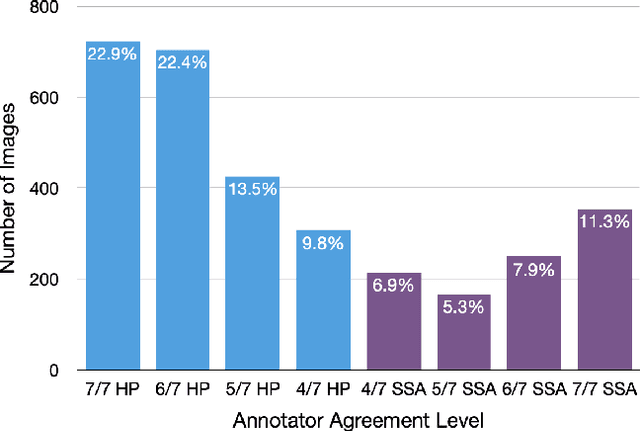
Abstract:With the rise of deep learning, there has been increased interest in using neural networks for histopathology image analysis, a field that investigates the properties of biopsy or resected specimens that are traditionally manually examined under a microscope by pathologists. In histopathology image analysis, however, challenges such as limited data, costly annotation, and processing high-resolution and variable-size images create a high barrier of entry and make it difficult to quickly iterate over model designs. Throughout scientific history, many significant research directions have leveraged small-scale experimental setups as petri dishes to efficiently evaluate exploratory ideas, which are then validated in large-scale applications. For instance, the Drosophila fruit fly in genetics and MNIST in computer vision are well-known petri dishes. In this paper, we introduce a minimalist histopathology image analysis dataset (MHIST), an analogous petri dish for histopathology image analysis. MHIST is a binary classification dataset of 3,152 fixed-size images of colorectal polyps, each with a gold-standard label determined by the majority vote of seven board-certified gastrointestinal pathologists and annotator agreement level. MHIST occupies less than 400 MB of disk space, and a ResNet-18 baseline can be trained to convergence on MHIST in just 6 minutes using 3.5 GB of memory on a NVIDIA RTX 3090. As example use cases, we use MHIST to study natural questions such as how dataset size, network depth, transfer learning, and high-disagreement examples affect model performance. By introducing MHIST, we hope to not only help facilitate the work of current histopathology imaging researchers, but also make histopathology image analysis more accessible to the general computer vision community. Our dataset is available at https://bmirds.github.io/MHIST.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge