Lifeng Han
DreamerNLplus: Interpretable Modeling of Mental Health Dynamics from Social Media Timelines using Hybrid Rule-Based and RAG Methods
May 21, 2026Abstract:We present DreamerNLplus, a hybrid framework for modeling mental health dynamics from social media timelines in the CLPsych 2026 shared task. Our system addresses three tasks: psychological state modeling, temporal change detection, and sequence-level summarization. For Task 1, we combine LLM-based data augmentation, DeBERTa classification, and Random Forest regression for structured state prediction. For Task 2, we use few-shot prompting with a locally deployed Llama 3.1 model to detect Switch and Escalation events using short-term temporal context. For Task 3.1, we explore both a deterministic rule-based summarization pipeline and a few-shot LLM-based approach, ranking \textbf{2nd} officially. Our RAG-based method achieves strong performance in Task 3.2, ranking \textbf{1st} for Improvement and \textbf{3rd} for Deterioration, demonstrating its ability to capture recurrent psychological change patterns across timelines. Our analysis reveals key challenges, including the mismatch between classification and regression performance, the difficulty of modeling temporal transitions, and the disagreement between semantic and similarity-based evaluation metrics. These findings highlight the complexity of modeling mental health dynamics and motivate future work on unified evaluation frameworks. We share our code and prompts at https://github.com/4dpicture/CLPsych2026
Cluster-Aware Attention-Based Deep Reinforcement Learning for Pickup and Delivery Problems
Mar 09, 2026Abstract:The Pickup and Delivery Problem (PDP) is a fundamental and challenging variant of the Vehicle Routing Problem, characterized by tightly coupled pickup--delivery pairs, precedence constraints, and spatial layouts that often exhibit clustering. Existing deep reinforcement learning (DRL) approaches either model all nodes on a flat graph, relying on implicit learning to enforce constraints, or achieve strong performance through inference-time collaborative search at the cost of substantial latency. In this paper, we propose \emph{CAADRL} (Cluster-Aware Attention-based Deep Reinforcement Learning), a DRL framework that explicitly exploits the multi-scale structure of PDP instances via cluster-aware encoding and hierarchical decoding. The encoder builds on a Transformer and combines global self-attention with intra-cluster attention over depot, pickup, and delivery nodes, producing embeddings that are both globally informative and locally role-aware. Based on these embeddings, we introduce a Dynamic Dual-Decoder with a learnable gate that balances intra-cluster routing and inter-cluster transitions at each step. The policy is trained end-to-end with a POMO-style policy gradient scheme using multiple symmetric rollouts per instance. Experiments on synthetic clustered and uniform PDP benchmarks show that CAADRL matches or improves upon strong state-of-the-art baselines on clustered instances and remains highly competitive on uniform instances, particularly as problem size increases. Crucially, our method achieves these results with substantially lower inference time than neural collaborative-search baselines, suggesting that explicitly modeling cluster structure provides an effective and efficient inductive bias for neural PDP solvers.
QuadAI at SemEval-2026 Task 3: Ensemble Learning of Hybrid RoBERTa and LLMs for Dimensional Aspect-Based Sentiment Analysis
Mar 08, 2026Abstract:We present our system for SemEval-2026 Task 3 on dimensional aspect-based sentiment regression. Our approach combines a hybrid RoBERTa encoder, which jointly predicts sentiment using regression and discretized classification heads, with large language models (LLMs) via prediction-level ensemble learning. The hybrid encoder improves prediction stability by combining continuous and discretized sentiment representations. We further explore in-context learning with LLMs and ridge-regression stacking to combine encoder and LLM predictions. Experimental results on the development set show that ensemble learning significantly improves performance over individual models, achieving substantial reductions in RMSE and improvements in correlation scores. Our findings demonstrate the complementary strengths of encoder-based and LLM-based approaches for dimensional sentiment analysis. Our development code and resources will be shared at https://github.com/aaronlifenghan/ABSentiment
FLANS at SemEval-2026 Task 7: RAG with Open-Sourced Smaller LLMs for Everyday Knowledge Across Diverse Languages and Cultures
Mar 02, 2026Abstract:This system paper describes our participation in the SemEval-2025 Task-7 ``Everyday Knowledge Across Diverse Languages and Cultures''. We attended two subtasks, i.e., Track 1: Short Answer Questions (SAQ), and Track 2: Multiple-Choice Questions (MCQ). The methods we used are retrieval augmented generation (RAGs) with open-sourced smaller LLMs (OS-sLLMs). To better adapt to this shared task, we created our own culturally aware knowledge base (CulKBs) by extracting Wikipedia content using keyword lists we prepared. We extracted both culturally-aware wiki-text and country-specific wiki-summary. In addition to the local CulKBs, we also have one system integrating live online search output via DuckDuckGo. Towards better privacy and sustainability, we aimed to deploy smaller LLMs (sLLMs) that are open-sourced on the Ollama platform. We share the prompts we developed using refinement techniques and report the learning curve of such prompts. The tested languages are English, Spanish, and Chinese for both tracks. Our resources and codes are shared via https://github.com/aaronlifenghan/FLANS-2026
Analyzing Cancer Patients' Experiences with Embedding-based Topic Modeling and LLMs
Jan 17, 2026Abstract:This study investigates the use of neural topic modeling and LLMs to uncover meaningful themes from patient storytelling data, to offer insights that could contribute to more patient-oriented healthcare practices. We analyze a collection of transcribed interviews with cancer patients (132,722 words in 13 interviews). We first evaluate BERTopic and Top2Vec for individual interview summarization by using similar preprocessing, chunking, and clustering configurations to ensure a fair comparison on Keyword Extraction. LLMs (GPT4) are then used for the next step topic labeling. Their outputs for a single interview (I0) are rated through a small-scale human evaluation, focusing on {coherence}, {clarity}, and {relevance}. Based on the preliminary results and evaluation, BERTopic shows stronger performance and is selected for further experimentation using three {clinically oriented embedding} models. We then analyzed the full interview collection with the best model setting. Results show that domain-specific embeddings improved topic \textit{precision} and \textit{interpretability}, with BioClinicalBERT producing the most consistent results across transcripts. The global analysis of the full dataset of 13 interviews, using the BioClinicalBERT embedding model, reveals the most dominant topics throughout all 13 interviews, namely ``Coordination and Communication in Cancer Care Management" and ``Patient Decision-Making in Cancer Treatment Journey''. Although the interviews are machine translations from Dutch to English, and clinical professionals are not involved in this evaluation, the findings suggest that neural topic modeling, particularly BERTopic, can help provide useful feedback to clinicians from patient interviews. This pipeline could support more efficient document navigation and strengthen the role of patients' voices in healthcare workflows.
Ara-HOPE: Human-Centric Post-Editing Evaluation for Dialectal Arabic to Modern Standard Arabic Translation
Dec 25, 2025Abstract:Dialectal Arabic to Modern Standard Arabic (DA-MSA) translation is a challenging task in Machine Translation (MT) due to significant lexical, syntactic, and semantic divergences between Arabic dialects and MSA. Existing automatic evaluation metrics and general-purpose human evaluation frameworks struggle to capture dialect-specific MT errors, hindering progress in translation assessment. This paper introduces Ara-HOPE, a human-centric post-editing evaluation framework designed to systematically address these challenges. The framework includes a five-category error taxonomy and a decision-tree annotation protocol. Through comparative evaluation of three MT systems (Arabic-centric Jais, general-purpose GPT-3.5, and baseline NLLB-200), Ara-HOPE effectively highlights systematic performance differences between these systems. The results show that dialect-specific terminology and semantic preservation remain the most persistent challenges in DA-MSA translation. Ara-HOPE establishes a new framework for evaluating Dialectal Arabic MT quality and provides actionable guidance for improving dialect-aware MT systems.
An Empirical Study on Chinese Character Decomposition in Multiword Expression-Aware Neural Machine Translation
Dec 17, 2025
Abstract:Word meaning, representation, and interpretation play fundamental roles in natural language understanding (NLU), natural language processing (NLP), and natural language generation (NLG) tasks. Many of the inherent difficulties in these tasks stem from Multi-word Expressions (MWEs), which complicate the tasks by introducing ambiguity, idiomatic expressions, infrequent usage, and a wide range of variations. Significant effort and substantial progress have been made in addressing the challenging nature of MWEs in Western languages, particularly English. This progress is attributed in part to the well-established research communities and the abundant availability of computational resources. However, the same level of progress is not true for language families such as Chinese and closely related Asian languages, which continue to lag behind in this regard. While sub-word modelling has been successfully applied to many Western languages to address rare words improving phrase comprehension, and enhancing machine translation (MT) through techniques like byte-pair encoding (BPE), it cannot be applied directly to ideograph language scripts like Chinese. In this work, we conduct a systematic study of the Chinese character decomposition technology in the context of MWE-aware neural machine translation (NMT). Furthermore, we report experiments to examine how Chinese character decomposition technology contributes to the representation of the original meanings of Chinese words and characters, and how it can effectively address the challenges of translating MWEs.
HealthcareNLP: where are we and what is next?
Dec 09, 2025

Abstract:This proposed tutorial focuses on Healthcare Domain Applications of NLP, what we have achieved around HealthcareNLP, and the challenges that lie ahead for the future. Existing reviews in this domain either overlook some important tasks, such as synthetic data generation for addressing privacy concerns, or explainable clinical NLP for improved integration and implementation, or fail to mention important methodologies, including retrieval augmented generation and the neural symbolic integration of LLMs and KGs. In light of this, the goal of this tutorial is to provide an introductory overview of the most important sub-areas of a patient- and resource-oriented HealthcareNLP, with three layers of hierarchy: data/resource layer: annotation guidelines, ethical approvals, governance, synthetic data; NLP-Eval layer: NLP tasks such as NER, RE, sentiment analysis, and linking/coding with categorised methods, leading to explainable HealthAI; patients layer: Patient Public Involvement and Engagement (PPIE), health literacy, translation, simplification, and summarisation (also NLP tasks), and shared decision-making support. A hands-on session will be included in the tutorial for the audience to use HealthcareNLP applications. The target audience includes NLP practitioners in the healthcare application domain, NLP researchers who are interested in domain applications, healthcare researchers, and students from NLP fields. The type of tutorial is "Introductory to CL/NLP topics (HealthcareNLP)" and the audience does not need prior knowledge to attend this. Tutorial materials: https://github.com/4dpicture/HealthNLP
Non-Linear Scoring Model for Translation Quality Evaluation
Nov 17, 2025Abstract:Analytic Translation Quality Evaluation (TQE), based on Multidimensional Quality Metrics (MQM), traditionally uses a linear error-to-penalty scale calibrated to a reference sample of 1000-2000 words. However, linear extrapolation biases judgment on samples of different sizes, over-penalizing short samples and under-penalizing long ones, producing misalignment with expert intuition. Building on the Multi-Range framework, this paper presents a calibrated, non-linear scoring model that better reflects how human content consumers perceive translation quality across samples of varying length. Empirical data from three large-scale enterprise environments shows that acceptable error counts grow logarithmically, not linearly, with sample size. Psychophysical and cognitive evidence, including the Weber-Fechner law and Cognitive Load Theory, supports this premise by explaining why the perceptual impact of additional errors diminishes while the cognitive burden grows with scale. We propose a two-parameter model E(x) = a * ln(1 + b * x), a, b > 0, anchored to a reference tolerance and calibrated from two tolerance points using a one-dimensional root-finding step. The model yields an explicit interval within which the linear approximation stays within +/-20 percent relative error and integrates into existing evaluation workflows with only a dynamic tolerance function added. The approach improves interpretability, fairness, and inter-rater reliability across both human and AI-generated translations. By operationalizing a perceptually valid scoring paradigm, it advances translation quality evaluation toward more accurate and scalable assessment. The model also provides a stronger basis for AI-based document-level evaluation aligned with human judgment. Implementation considerations for CAT/LQA systems and implications for human and AI-generated text evaluation are discussed.
Dutch Metaphor Extraction from Cancer Patients' Interviews and Forum Data using LLMs and Human in the Loop
Nov 09, 2025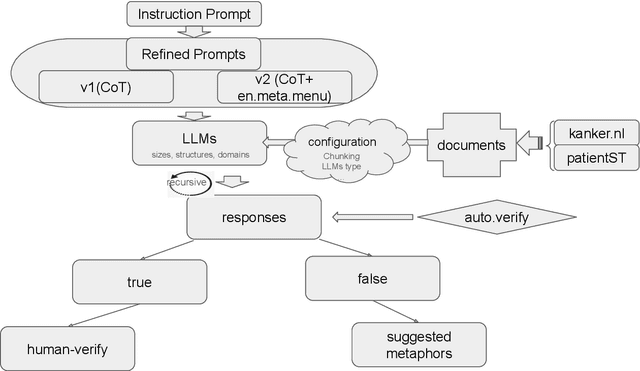

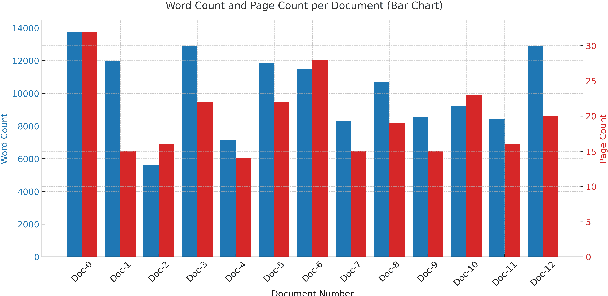
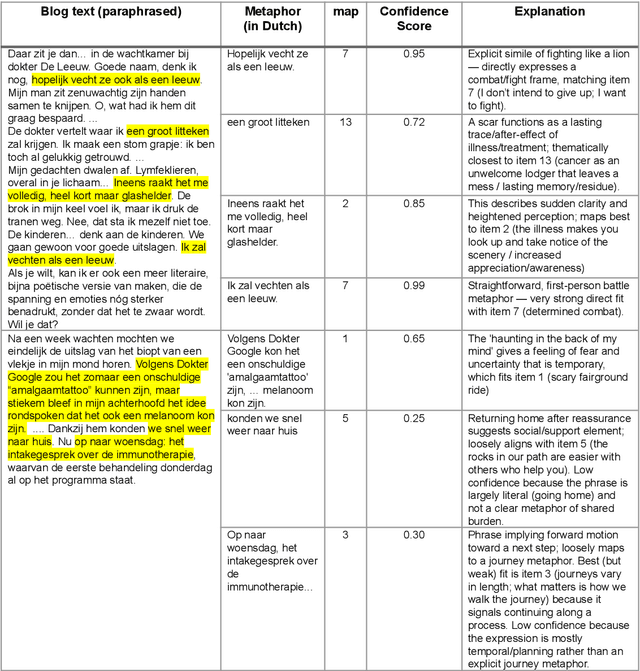
Abstract:Metaphors and metaphorical language (MLs) play an important role in healthcare communication between clinicians, patients, and patients' family members. In this work, we focus on Dutch language data from cancer patients. We extract metaphors used by patients using two data sources: (1) cancer patient storytelling interview data and (2) online forum data, including patients' posts, comments, and questions to professionals. We investigate how current state-of-the-art large language models (LLMs) perform on this task by exploring different prompting strategies such as chain of thought reasoning, few-shot learning, and self-prompting. With a human-in-the-loop setup, we verify the extracted metaphors and compile the outputs into a corpus named HealthQuote.NL. We believe the extracted metaphors can support better patient care, for example shared decision making, improved communication between patients and clinicians, and enhanced patient health literacy. They can also inform the design of personalized care pathways. We share prompts and related resources at https://github.com/aaronlifenghan/HealthQuote.NL
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge