Koji Fujimoto
RadVLM: A Multitask Conversational Vision-Language Model for Radiology
Feb 05, 2025



Abstract:The widespread use of chest X-rays (CXRs), coupled with a shortage of radiologists, has driven growing interest in automated CXR analysis and AI-assisted reporting. While existing vision-language models (VLMs) show promise in specific tasks such as report generation or abnormality detection, they often lack support for interactive diagnostic capabilities. In this work we present RadVLM, a compact, multitask conversational foundation model designed for CXR interpretation. To this end, we curate a large-scale instruction dataset comprising over 1 million image-instruction pairs containing both single-turn tasks -- such as report generation, abnormality classification, and visual grounding -- and multi-turn, multi-task conversational interactions. After fine-tuning RadVLM on this instruction dataset, we evaluate it across different tasks along with re-implemented baseline VLMs. Our results show that RadVLM achieves state-of-the-art performance in conversational capabilities and visual grounding while remaining competitive in other radiology tasks. Ablation studies further highlight the benefit of joint training across multiple tasks, particularly for scenarios with limited annotated data. Together, these findings highlight the potential of RadVLM as a clinically relevant AI assistant, providing structured CXR interpretation and conversational capabilities to support more effective and accessible diagnostic workflows.
Exploring Multilingual Large Language Models for Enhanced TNM classification of Radiology Report in lung cancer staging
Jun 12, 2024Abstract:Background: Structured radiology reports remains underdeveloped due to labor-intensive structuring and narrative-style reporting. Deep learning, particularly large language models (LLMs) like GPT-3.5, offers promise in automating the structuring of radiology reports in natural languages. However, although it has been reported that LLMs are less effective in languages other than English, their radiological performance has not been extensively studied. Purpose: This study aimed to investigate the accuracy of TNM classification based on radiology reports using GPT3.5-turbo (GPT3.5) and the utility of multilingual LLMs in both Japanese and English. Material and Methods: Utilizing GPT3.5, we developed a system to automatically generate TNM classifications from chest CT reports for lung cancer and evaluate its performance. We statistically analyzed the impact of providing full or partial TNM definitions in both languages using a Generalized Linear Mixed Model. Results: Highest accuracy was attained with full TNM definitions and radiology reports in English (M = 94%, N = 80%, T = 47%, and ALL = 36%). Providing definitions for each of the T, N, and M factors statistically improved their respective accuracies (T: odds ratio (OR) = 2.35, p < 0.001; N: OR = 1.94, p < 0.01; M: OR = 2.50, p < 0.001). Japanese reports exhibited decreased N and M accuracies (N accuracy: OR = 0.74 and M accuracy: OR = 0.21). Conclusion: This study underscores the potential of multilingual LLMs for automatic TNM classification in radiology reports. Even without additional model training, performance improvements were evident with the provided TNM definitions, indicating LLMs' relevance in radiology contexts.
Radiology-Aware Model-Based Evaluation Metric for Report Generation
Nov 28, 2023Abstract:We propose a new automated evaluation metric for machine-generated radiology reports using the successful COMET architecture adapted for the radiology domain. We train and publish four medically-oriented model checkpoints, including one trained on RadGraph, a radiology knowledge graph. Our results show that our metric correlates moderately to high with established metrics such as BERTscore, BLEU, and CheXbert scores. Furthermore, we demonstrate that one of our checkpoints exhibits a high correlation with human judgment, as assessed using the publicly available annotations of six board-certified radiologists, using a set of 200 reports. We also performed our own analysis gathering annotations with two radiologists on a collection of 100 reports. The results indicate the potential effectiveness of our method as a radiology-specific evaluation metric. The code, data, and model checkpoints to reproduce our findings will be publicly available.
Boosting Radiology Report Generation by Infusing Comparison Prior
May 08, 2023



Abstract:Current transformer-based models achieved great success in generating radiology reports from chest X-ray images. Nonetheless, one of the major issues is the model's lack of prior knowledge, which frequently leads to false references to non-existent prior exams in synthetic reports. This is mainly due to the knowledge gap between radiologists and the generation models: radiologists are aware of the prior information of patients to write a medical report, while models only receive X-ray images at a specific time. To address this issue, we propose a novel approach that employs a labeler to extract comparison prior information from radiology reports in the IU X-ray and MIMIC-CXR datasets. This comparison prior is then incorporated into state-of-the-art transformer-based models, allowing them to generate more realistic and comprehensive reports. We test our method on the IU X-ray and MIMIC-CXR datasets and find that it outperforms previous state-of-the-art models in terms of both automatic and human evaluation metrics. In addition, unlike previous models, our model generates reports that do not contain false references to non-existent prior exams. Our approach provides a promising direction for bridging the gap between radiologists and generation models in medical report generation.
Progressive Transformer-Based Generation of Radiology Reports
Feb 19, 2021
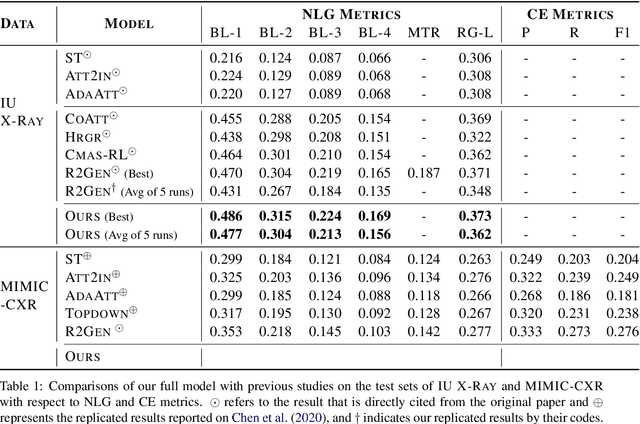
Abstract:Inspired by Curriculum Learning, we propose a consecutive (i.e. image-to-text-to-text) generation framework where we divide the problem of radiology report generation into two steps. Contrary to generating the full radiology report from the image at once, the model generates global concepts from the image in the first step and then reforms them into finer and coherent texts using transformer-based architecture. We follow the transformer-based sequence-to-sequence paradigm at each step. We improve upon the state-of-the-art on two benchmark datasets.
Lung segmentation on chest x-ray images in patients with severe abnormal findings using deep learning
Aug 21, 2019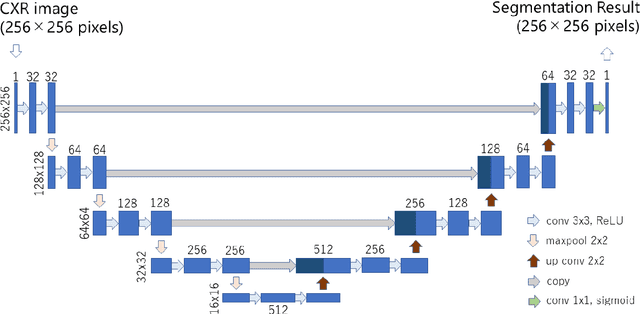
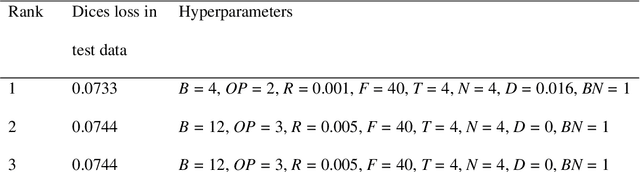

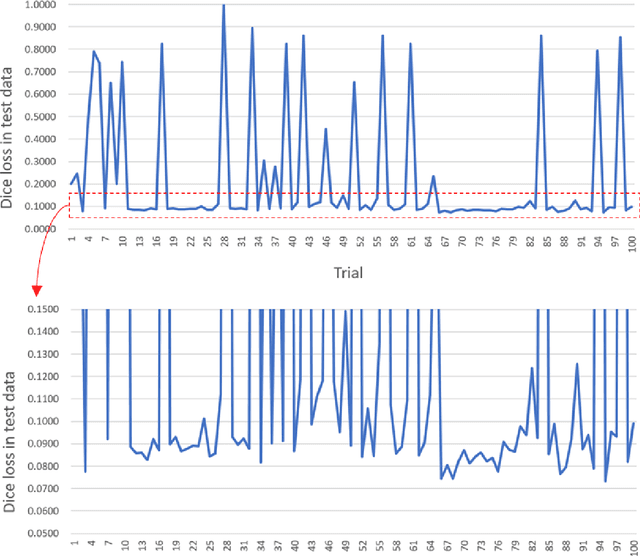
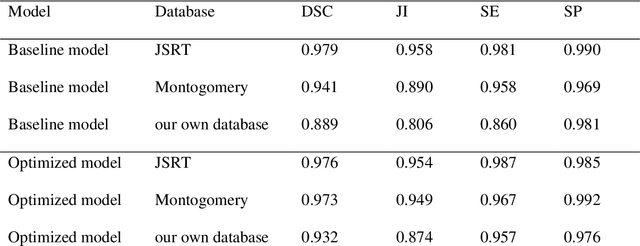
Abstract:Rationale and objectives: Several studies have evaluated the usefulness of deep learning for lung segmentation using chest x-ray (CXR) images with small- or medium-sized abnormal findings. Here, we built a database including both CXR images with severe abnormalities and experts' lung segmentation results, and aimed to evaluate our network's efficacy in lung segmentation from these images. Materials and Methods: For lung segmentation, CXR images from the Japanese Society of Radiological Technology (JSRT, N = 247) and Montgomery databases (N = 138), were included, and 65 additional images depicting severe abnormalities from a public database were evaluated and annotated by a radiologist, thereby adding lung segmentation results to these images. Baseline U-net was used to segment the lungs in images from the three databases. Subsequently, the U-net network architecture was automatically optimized for lung segmentation from CXR images using Bayesian optimization. Dice similarity coefficient (DSC) was calculated to confirm segmentation. Results: Our results demonstrated that using baseline U-net yielded poorer lung segmentation results in our database than those in the JSRT and Montgomery databases, implying that robust segmentation of lungs may be difficult because of severe abnormalities. The DSC values with baseline U-net for the JSRT, Montgomery and our databases were 0.979, 0.941, and 0.889, respectively, and with optimized U-net, 0.976, 0.973, and 0.932, respectively. Conclusion: For robust lung segmentation, the U-net architecture was optimized via Bayesian optimization, and our results demonstrate that the optimized U-net was more robust than baseline U-net in lung segmentation from CXR images with large-sized abnormalities.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge