Takamichi Murakami
Department of Radiology, Kobe University Graduate School of Medicine, Kobe, Japan
Exploring Multilingual Large Language Models for Enhanced TNM classification of Radiology Report in lung cancer staging
Jun 12, 2024Abstract:Background: Structured radiology reports remains underdeveloped due to labor-intensive structuring and narrative-style reporting. Deep learning, particularly large language models (LLMs) like GPT-3.5, offers promise in automating the structuring of radiology reports in natural languages. However, although it has been reported that LLMs are less effective in languages other than English, their radiological performance has not been extensively studied. Purpose: This study aimed to investigate the accuracy of TNM classification based on radiology reports using GPT3.5-turbo (GPT3.5) and the utility of multilingual LLMs in both Japanese and English. Material and Methods: Utilizing GPT3.5, we developed a system to automatically generate TNM classifications from chest CT reports for lung cancer and evaluate its performance. We statistically analyzed the impact of providing full or partial TNM definitions in both languages using a Generalized Linear Mixed Model. Results: Highest accuracy was attained with full TNM definitions and radiology reports in English (M = 94%, N = 80%, T = 47%, and ALL = 36%). Providing definitions for each of the T, N, and M factors statistically improved their respective accuracies (T: odds ratio (OR) = 2.35, p < 0.001; N: OR = 1.94, p < 0.01; M: OR = 2.50, p < 0.001). Japanese reports exhibited decreased N and M accuracies (N accuracy: OR = 0.74 and M accuracy: OR = 0.21). Conclusion: This study underscores the potential of multilingual LLMs for automatic TNM classification in radiology reports. Even without additional model training, performance improvements were evident with the provided TNM definitions, indicating LLMs' relevance in radiology contexts.
Development of pericardial fat count images using a combination of three different deep-learning models
Jul 25, 2023



Abstract:Rationale and Objectives: Pericardial fat (PF), the thoracic visceral fat surrounding the heart, promotes the development of coronary artery disease by inducing inflammation of the coronary arteries. For evaluating PF, this study aimed to generate pericardial fat count images (PFCIs) from chest radiographs (CXRs) using a dedicated deep-learning model. Materials and Methods: The data of 269 consecutive patients who underwent coronary computed tomography (CT) were reviewed. Patients with metal implants, pleural effusion, history of thoracic surgery, or that of malignancy were excluded. Thus, the data of 191 patients were used. PFCIs were generated from the projection of three-dimensional CT images, where fat accumulation was represented by a high pixel value. Three different deep-learning models, including CycleGAN, were combined in the proposed method to generate PFCIs from CXRs. A single CycleGAN-based model was used to generate PFCIs from CXRs for comparison with the proposed method. To evaluate the image quality of the generated PFCIs, structural similarity index measure (SSIM), mean squared error (MSE), and mean absolute error (MAE) of (i) the PFCI generated using the proposed method and (ii) the PFCI generated using the single model were compared. Results: The mean SSIM, MSE, and MAE were as follows: 0.856, 0.0128, and 0.0357, respectively, for the proposed model; and 0.762, 0.0198, and 0.0504, respectively, for the single CycleGAN-based model. Conclusion: PFCIs generated from CXRs with the proposed model showed better performance than those with the single model. PFCI evaluation without CT may be possible with the proposed method.
Unsupervised-learning-based method for chest MRI-CT transformation using structure constrained unsupervised generative attention networks
Jun 16, 2021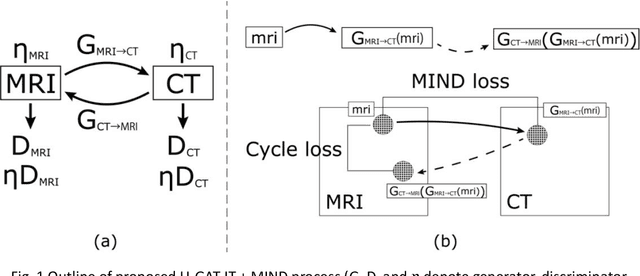
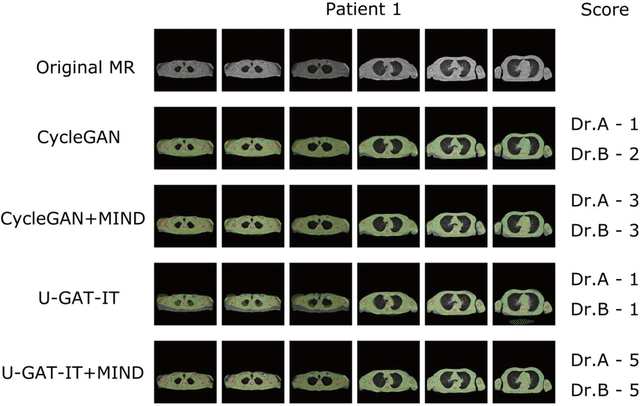
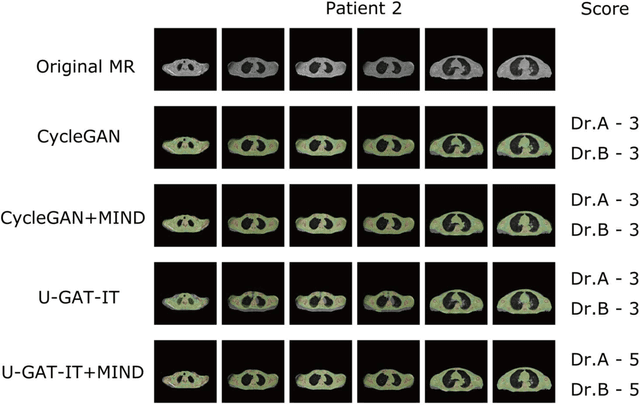
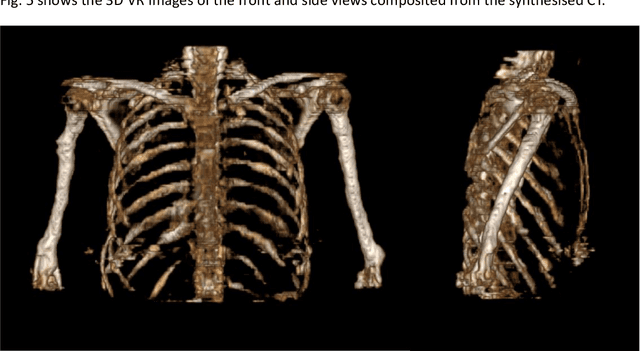
Abstract:The integrated positron emission tomography/magnetic resonance imaging (PET/MRI) scanner facilitates the simultaneous acquisition of metabolic information via PET and morphological information with high soft-tissue contrast using MRI. Although PET/MRI facilitates the capture of high-accuracy fusion images, its major drawback can be attributed to the difficulty encountered when performing attenuation correction, which is necessary for quantitative PET evaluation. The combined PET/MRI scanning requires the generation of attenuation-correction maps from MRI owing to no direct relationship between the gamma-ray attenuation information and MRIs. While MRI-based bone-tissue segmentation can be readily performed for the head and pelvis regions, the realization of accurate bone segmentation via chest CT generation remains a challenging task. This can be attributed to the respiratory and cardiac motions occurring in the chest as well as its anatomically complicated structure and relatively thin bone cortex. This paper presents a means to minimise the anatomical structural changes without human annotation by adding structural constraints using a modality-independent neighbourhood descriptor (MIND) to a generative adversarial network (GAN) that can transform unpaired images. The results obtained in this study revealed the proposed U-GAT-IT + MIND approach to outperform all other competing approaches. The findings of this study hint towards possibility of synthesising clinically acceptable CT images from chest MRI without human annotation, thereby minimising the changes in the anatomical structure.
Automatic classification between COVID-19 pneumonia, non-COVID-19 pneumonia, and the healthy on chest X-ray image: combination of data augmentation methods
Jun 12, 2020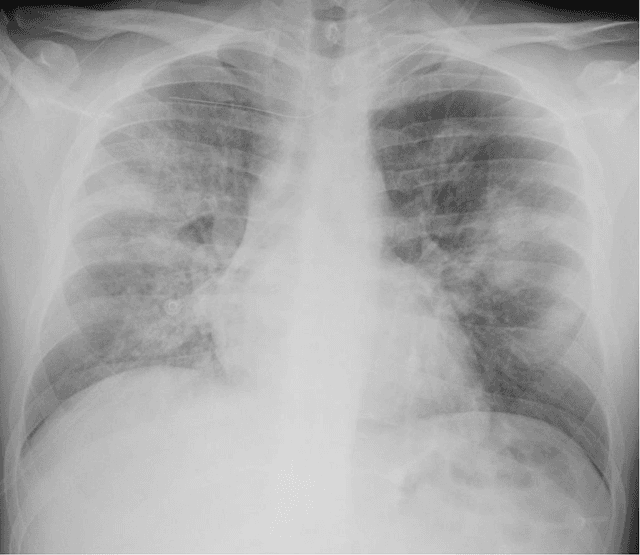
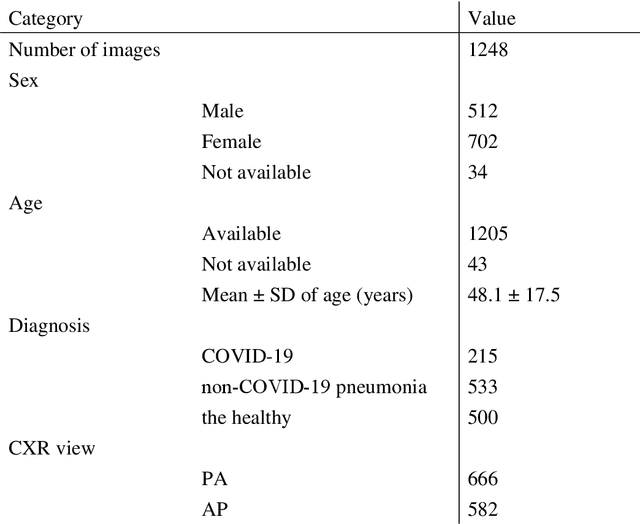
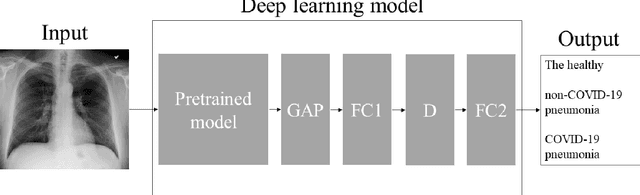
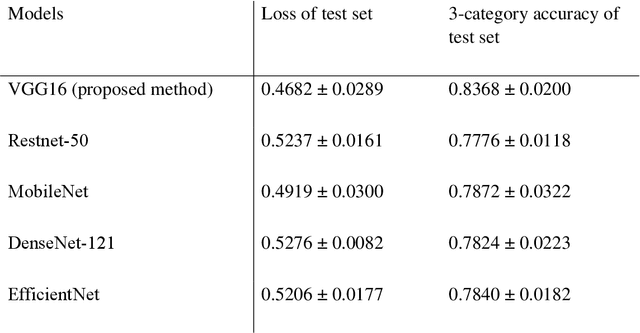
Abstract:Purpose: This study aimed to develop and validate computer-aided diagnosis (CXDx) system for classification between COVID-19 pneumonia, non-COVID-19 pneumonia, and the healthy on chest X-ray (CXR) images. Materials and Methods: From two public datasets, 1248 CXR images were obtained, which included 215, 533, and 500 CXR images of COVID-19 pneumonia patients, non-COVID-19 pneumonia patients, and the healthy samples. The proposed CADx system utilized VGG16 as a pre-trained model and combination of conventional method and mixup as data augmentation methods. Other types of pre-trained models were compared with the VGG16-based model. Single type or no data augmentation methods were also evaluated. Splitting of training/validation/test sets was used when building and evaluating the CADx system. Three-category accuracy was evaluated for test set with 125 CXR images. Results: The three-category accuracy of the CAD system was 83.6% between COVID-19 pneumonia, non-COVID-19 pneumonia, and the healthy. Sensitivity for COVID-19 pneumonia was more than 90%. The combination of conventional method and mixup was more useful than single type or no data augmentation method. Conclusion: This study was able to create an accurate CADx system for the 3-category classification. Source code of our CADx system is available as open source for COVID-19 research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge