Koen Van Leemput
Department of Applied Mathematics and Computer Science, Technical University of Denmark, Denmark, Athinoula A. Martinos Center for Biomedical Imaging, Massachusetts General Hospital, Harvard Medical School, USA
The ASNR-MICCAI Brain Tumor Segmentation (BraTS) Challenge 2023: Intracranial Meningioma
May 12, 2023



Abstract:Meningiomas are the most common primary intracranial tumor in adults and can be associated with significant morbidity and mortality. Radiologists, neurosurgeons, neuro-oncologists, and radiation oncologists rely on multiparametric MRI (mpMRI) for diagnosis, treatment planning, and longitudinal treatment monitoring; yet automated, objective, and quantitative tools for non-invasive assessment of meningiomas on mpMRI are lacking. The BraTS meningioma 2023 challenge will provide a community standard and benchmark for state-of-the-art automated intracranial meningioma segmentation models based on the largest expert annotated multilabel meningioma mpMRI dataset to date. Challenge competitors will develop automated segmentation models to predict three distinct meningioma sub-regions on MRI including enhancing tumor, non-enhancing tumor core, and surrounding nonenhancing T2/FLAIR hyperintensity. Models will be evaluated on separate validation and held-out test datasets using standardized metrics utilized across the BraTS 2023 series of challenges including the Dice similarity coefficient and Hausdorff distance. The models developed during the course of this challenge will aid in incorporation of automated meningioma MRI segmentation into clinical practice, which will ultimately improve care of patients with meningioma.
An Open-Source Tool for Longitudinal Whole-Brain and White Matter Lesion Segmentation
Jul 10, 2022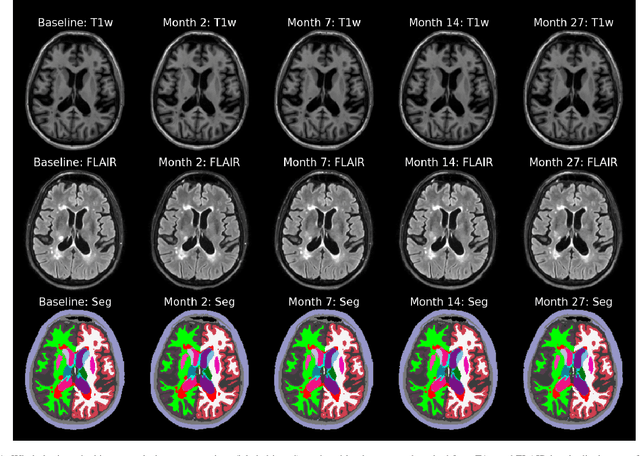
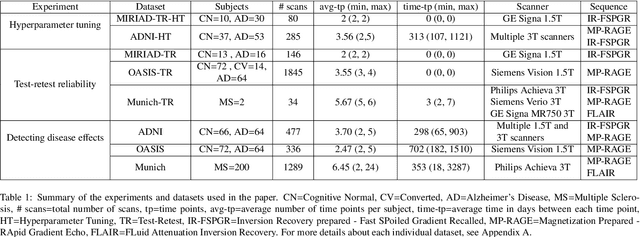
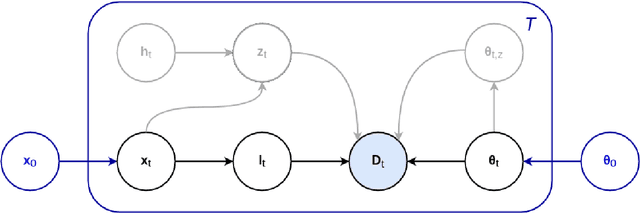

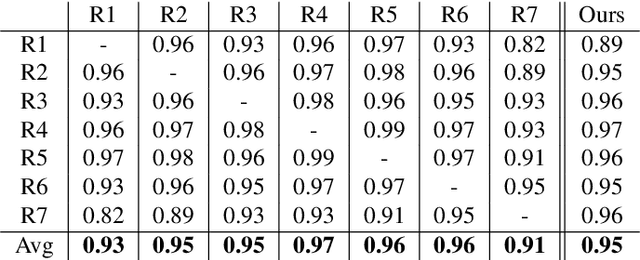
Abstract:In this paper we describe and validate a longitudinal method for whole-brain segmentation of longitudinal MRI scans. It builds upon an existing whole-brain segmentation method that can handle multi-contrast data and robustly analyze images with white matter lesions. This method is here extended with subject-specific latent variables that encourage temporal consistency between its segmentation results, enabling it to better track subtle morphological changes in dozens of neuroanatomical structures and white matter lesions. We validate the proposed method on multiple datasets of control subjects and patients suffering from Alzheimer's disease and multiple sclerosis, and compare its results against those obtained with its original cross-sectional formulation and two benchmark longitudinal methods. The results indicate that the method attains a higher test-retest reliability, while being more sensitive to longitudinal disease effect differences between patient groups. An implementation is publicly available as part of the open-source neuroimaging package FreeSurfer.
Prediction of MGMT Methylation Status of Glioblastoma using Radiomics and Latent Space Shape Features
Sep 25, 2021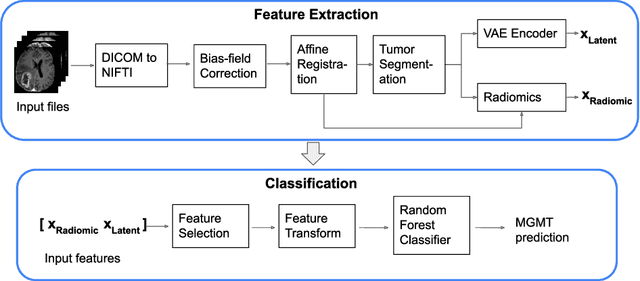
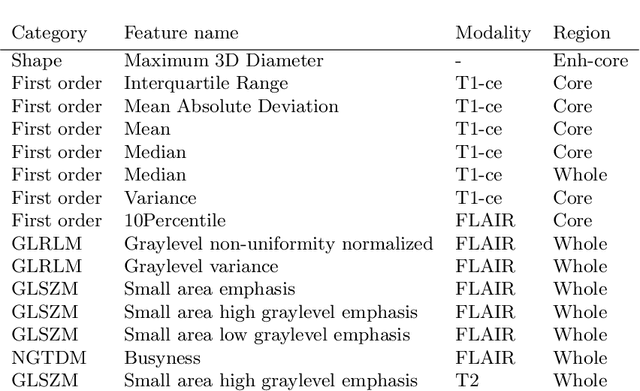
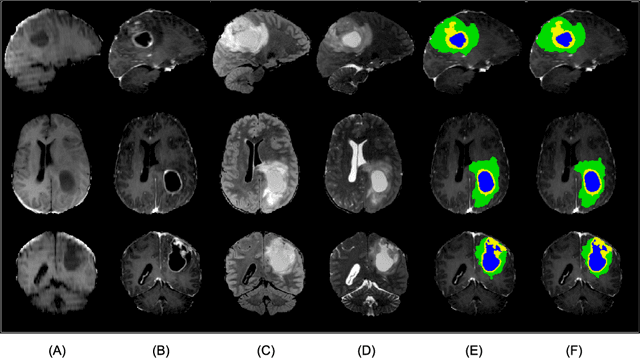
Abstract:In this paper we propose a method for predicting the status of MGMT promoter methylation in high-grade gliomas. From the available MR images, we segment the tumor using deep convolutional neural networks and extract both radiomic features and shape features learned by a variational autoencoder. We implemented a standard machine learning workflow to obtain predictions, consisting of feature selection followed by training of a random forest classification model. We trained and evaluated our method on the RSNA-ASNR-MICCAI BraTS 2021 challenge dataset and submitted our predictions to the challenge.
Predicting survival of glioblastoma from automatic whole-brain and tumor segmentation of MR images
Sep 25, 2021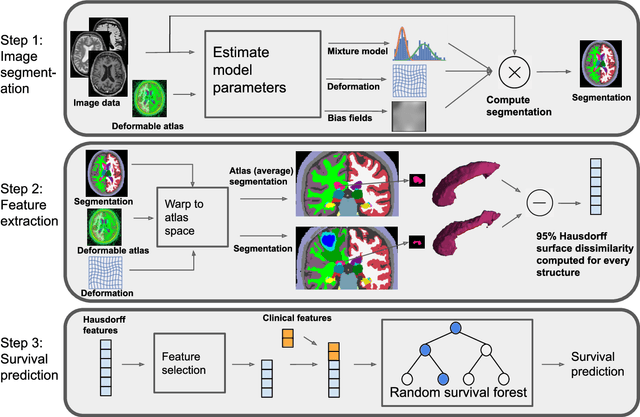
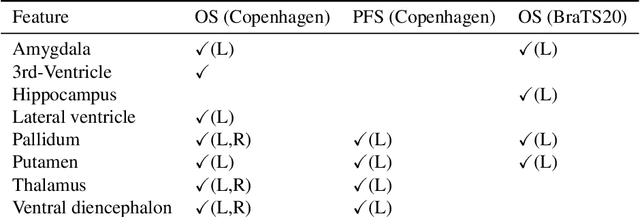

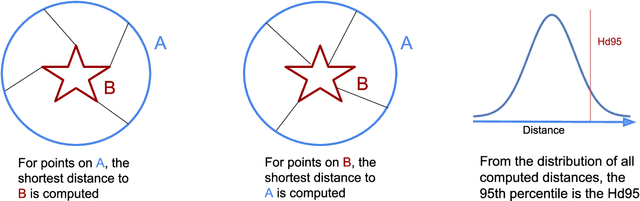
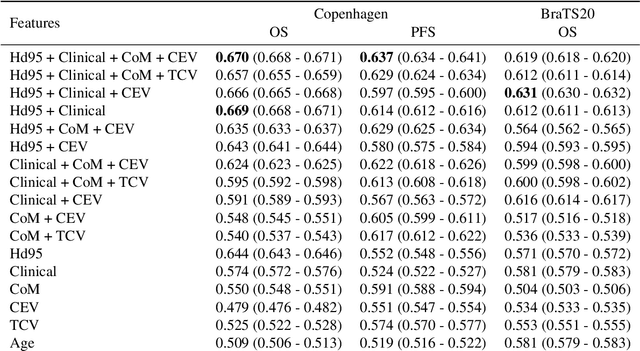
Abstract:Survival prediction models can potentially be used to guide treatment of glioblastoma patients. However, currently available MR imaging biomarkers holding prognostic information are often challenging to interpret, have difficulties generalizing across data acquisitions, or are only applicable to pre-operative MR data. In this paper we aim to address these issues by introducing novel imaging features that can be automatically computed from MR images and fed into machine learning models to predict patient survival. The features we propose have a direct biological interpretation: They measure the deformation caused by the tumor on the surrounding brain structures, comparing the shape of various structures in the patient's brain to their expected shape in healthy individuals. To obtain the required segmentations, we use an automatic method that is contrast-adaptive and robust to missing modalities, making the features generalizable across scanners and imaging protocols. Since the features we propose do not depend on characteristics of the tumor region itself, they are also applicable to post-operative images, which have been much less studied in the context of survival prediction. Using experiments involving both pre- and post-operative data, we show that the proposed features carry prognostic value in terms of overall- and progression-free survival, over and above that of conventional non-imaging features.
SynthSeg: Domain Randomisation for Segmentation of Brain MRI Scans of any Contrast and Resolution
Jul 20, 2021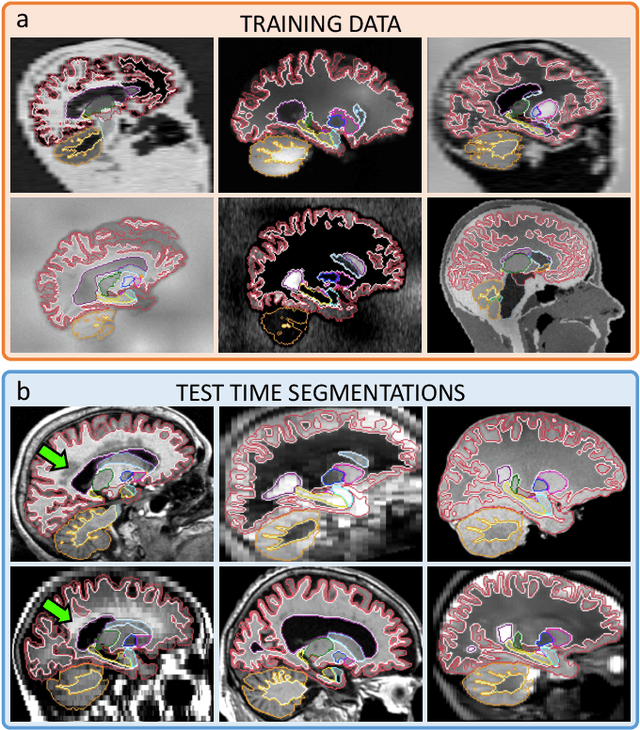
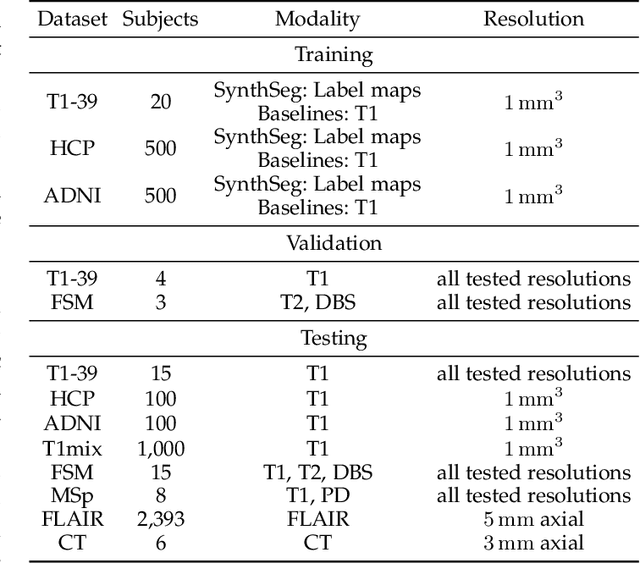
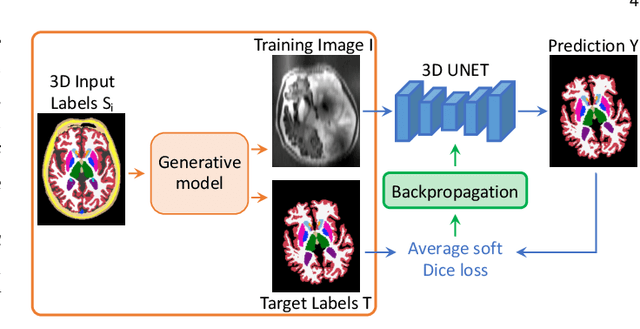
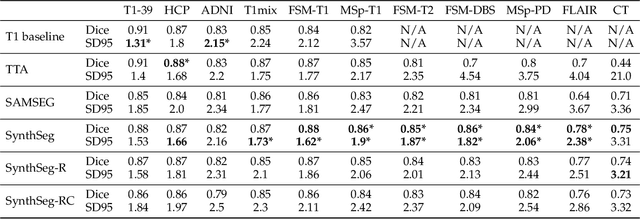
Abstract:Despite advances in data augmentation and transfer learning, convolutional neural networks (CNNs) have difficulties generalising to unseen target domains. When applied to segmentation of brain MRI scans, CNNs are highly sensitive to changes in resolution and contrast: even within the same MR modality, decreases in performance can be observed across datasets. We introduce SynthSeg, the first segmentation CNN agnostic to brain MRI scans of any contrast and resolution. SynthSeg is trained with synthetic data sampled from a generative model inspired by Bayesian segmentation. Crucially, we adopt a \textit{domain randomisation} strategy where we fully randomise the generation parameters to maximise the variability of the training data. Consequently, SynthSeg can segment preprocessed and unpreprocessed real scans of any target domain, without retraining or fine-tuning. Because SynthSeg only requires segmentations to be trained (no images), it can learn from label maps obtained automatically from existing datasets of different populations (e.g., with atrophy and lesions), thus achieving robustness to a wide range of morphological variability. We demonstrate SynthSeg on 5,500 scans of 6 modalities and 10 resolutions, where it exhibits unparalleled generalisation compared to supervised CNNs, test time adaptation, and Bayesian segmentation. The code and trained model are available at https://github.com/BBillot/SynthSeg.
An augmentation strategy to mimic multi-scanner variability in MRI
Mar 23, 2021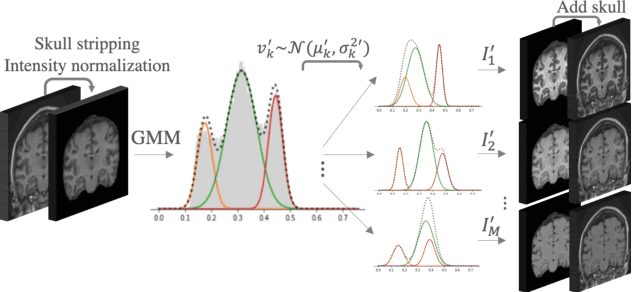
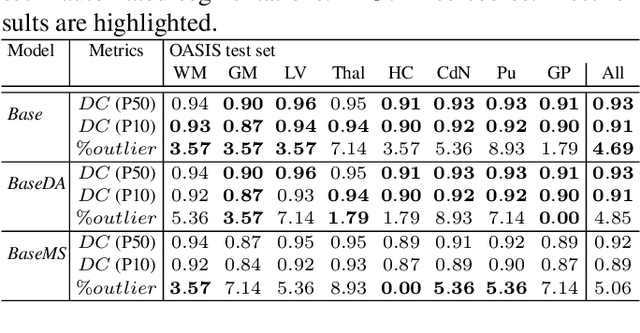
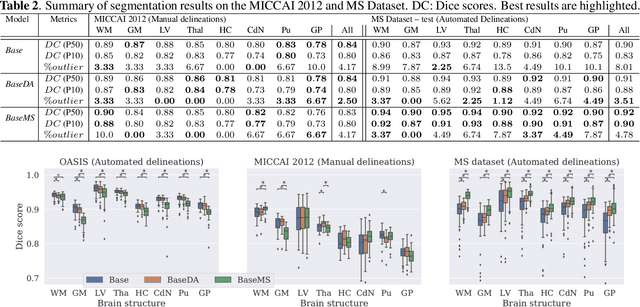
Abstract:Most publicly available brain MRI datasets are very homogeneous in terms of scanner and protocols, and it is difficult for models that learn from such data to generalize to multi-center and multi-scanner data. We propose a novel data augmentation approach with the aim of approximating the variability in terms of intensities and contrasts present in real world clinical data. We use a Gaussian Mixture Model based approach to change tissue intensities individually, producing new contrasts while preserving anatomical information. We train a deep learning model on a single scanner dataset and evaluate it on a multi-center and multi-scanner dataset. The proposed approach improves the generalization capability of the model to other scanners not present in the training data.
A Longitudinal Method for Simultaneous Whole-Brain and Lesion Segmentation in Multiple Sclerosis
Sep 15, 2020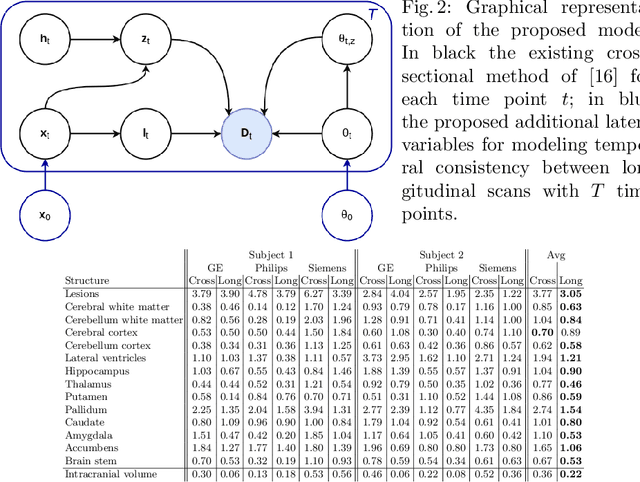
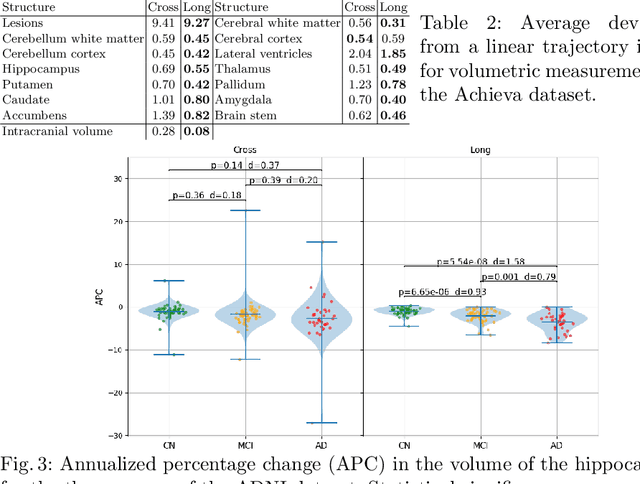
Abstract:In this paper we propose a novel method for the segmentation of longitudinal brain MRI scans of patients suffering from Multiple Sclerosis. The method builds upon an existing cross-sectional method for simultaneous whole-brain and lesion segmentation, introducing subject-specific latent variables to encourage temporal consistency between longitudinal scans. It is very generally applicable, as it does not make any prior assumptions on the scanner, the MRI protocol, or the number and timing of longitudinal follow-up scans. Preliminary experiments on three longitudinal datasets indicate that the proposed method produces more reliable segmentations and detects disease effects better than the cross-sectional method it is based upon.
3D Reconstruction and Segmentation of Dissection Photographs for MRI-free Neuropathology
Sep 11, 2020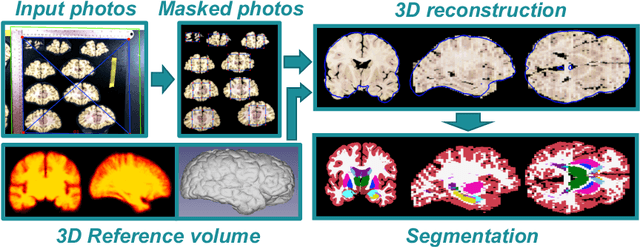
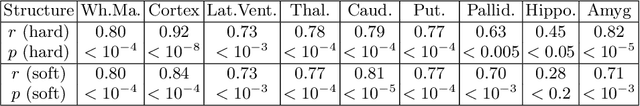
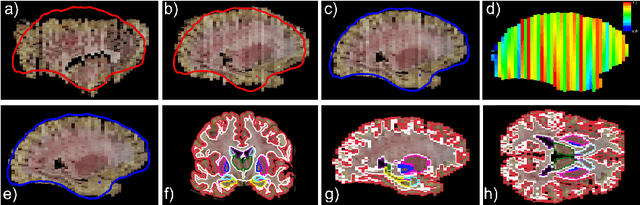
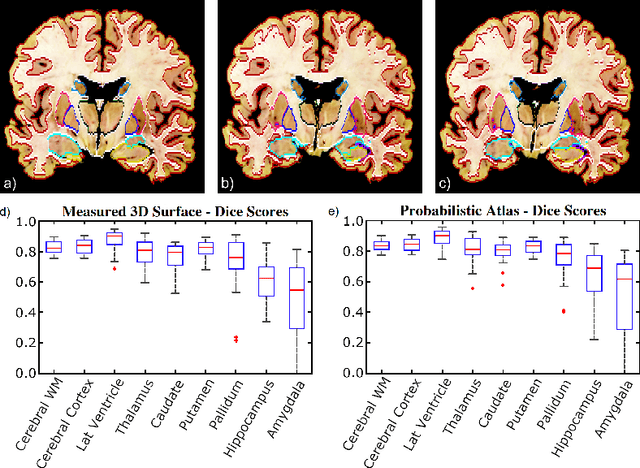
Abstract:Neuroimaging to neuropathology correlation (NTNC) promises to enable the transfer of microscopic signatures of pathology to in vivo imaging with MRI, ultimately enhancing clinical care. NTNC traditionally requires a volumetric MRI scan, acquired either ex vivo or a short time prior to death. Unfortunately, ex vivo MRI is difficult and costly, and recent premortem scans of sufficient quality are seldom available. To bridge this gap, we present methodology to 3D reconstruct and segment full brain image volumes from brain dissection photographs, which are routinely acquired at many brain banks and neuropathology departments. The 3D reconstruction is achieved via a joint registration framework, which uses a reference volume other than MRI. This volume may represent either the sample at hand (e.g., a surface 3D scan) or the general population (a probabilistic atlas). In addition, we present a Bayesian method to segment the 3D reconstructed photographic volumes into 36 neuroanatomical structures, which is robust to nonuniform brightness within and across photographs. We evaluate our methods on a dataset with 24 brains, using Dice scores and volume correlations. The results show that dissection photography is a valid replacement for ex vivo MRI in many volumetric analyses, opening an avenue for MRI-free NTNC, including retrospective data. The code is available at https://github.com/htregidgo/DissectionPhotoVolumes.
A Contrast-Adaptive Method for Simultaneous Whole-Brain and Lesion Segmentation in Multiple Sclerosis
May 11, 2020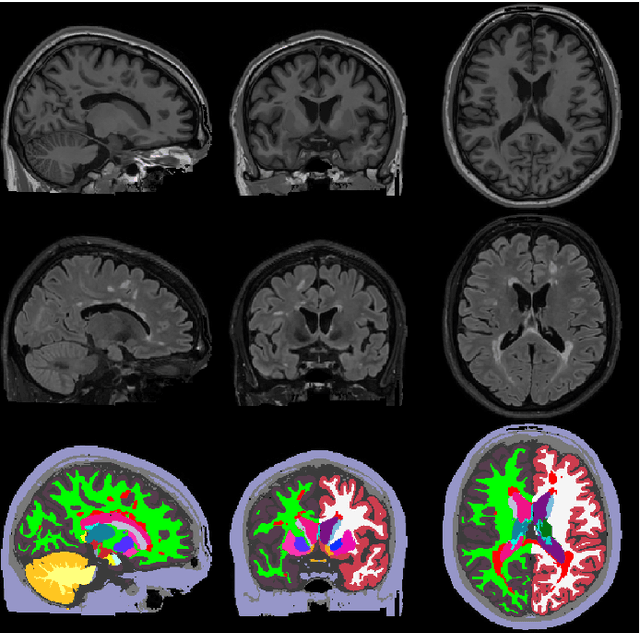
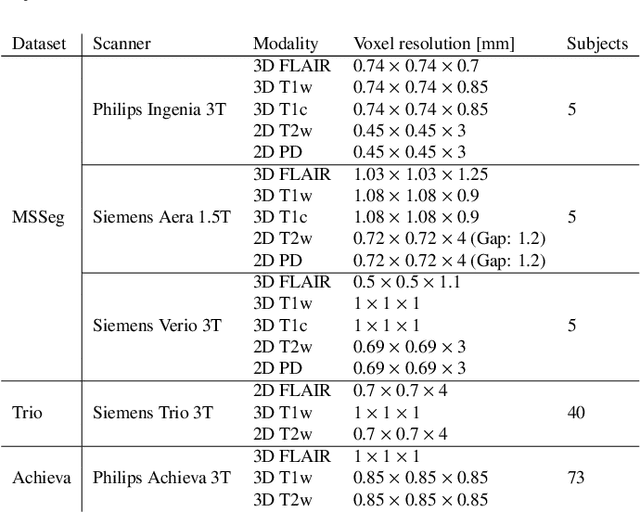
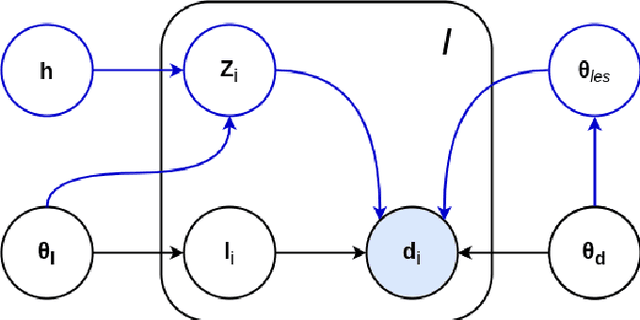
Abstract:Here we present a method for the simultaneous segmentation of white matter lesions and normal-appearing neuroanatomical structures from multi-contrast brain MRI scans of multiple sclerosis patients. The method integrates a novel model for white matter lesions into a previously validated generative model for whole-brain segmentation. By using separate models for the shape of anatomical structures and their appearance in MRI, the algorithm can adapt to data acquired with different scanners and imaging protocols without retraining. We validate the method using three disparate datasets, showing state-of-the-art performance in white matter lesion segmentation while simultaneously segmenting dozens of other brain structures. We further demonstrate that the contrast-adaptive method can also be applied robustly to MRI scans of healthy controls, and replicate previously documented atrophy patterns in deep gray matter structures in MS. The algorithm is publicly available as part of the open-source neuroimaging package FreeSurfer.
A Learning Strategy for Contrast-agnostic MRI Segmentation
Mar 04, 2020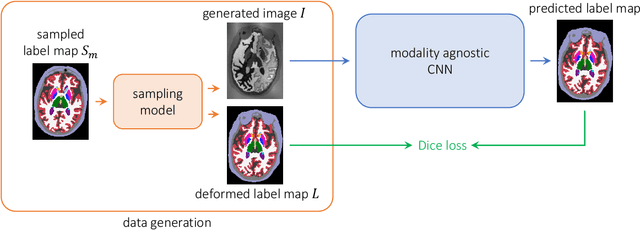

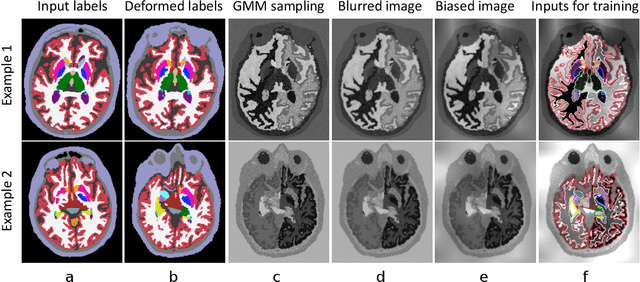

Abstract:We present a deep learning strategy that enables, for the first time, contrast-agnostic semantic segmentation of completely unpreprocessed brain MRI scans, without requiring additional training or fine-tuning for new modalities. Classical Bayesian methods address this segmentation problem with unsupervised intensity models, but require significant computational resources. In contrast, learning-based methods can be fast at test time, but are sensitive to the data available at training. Our proposed learning method, SynthSeg, leverages a set of training segmentations (no intensity images required) to generate synthetic sample images of widely varying contrasts on the fly during training. These samples are produced using the generative model of the classical Bayesian segmentation framework, with randomly sampled parameters for appearance, deformation, noise, and bias field. Because each mini-batch has a different synthetic contrast, the final network is not biased towards any MRI contrast. We comprehensively evaluate our approach on four datasets comprising over 1,000 subjects and four types of MR contrast. The results show that our approach successfully segments every contrast in the data, performing slightly better than classical Bayesian segmentation, and three orders of magnitude faster. Moreover, even within the same type of MRI contrast, our strategy generalizes significantly better across datasets, compared to training using real images. Finally, we find that synthesizing a broad range of contrasts, even if unrealistic, increases the generalization of the neural network. Our code and model are open source at https://github.com/BBillot/SynthSeg.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge