Keiland Cooper
ConformalHDC: Uncertainty-Aware Hyperdimensional Computing with Application to Neural Decoding
Feb 24, 2026Abstract:Hyperdimensional Computing (HDC) offers a computationally efficient paradigm for neuromorphic learning. Yet, it lacks rigorous uncertainty quantification, leading to open decision boundaries and, consequently, vulnerability to outliers, adversarial perturbations, and out-of-distribution inputs. To address these limitations, we introduce ConformalHDC, a unified framework that combines the statistical guarantees of conformal prediction with the computational efficiency of HDC. For this framework, we propose two complementary variations. First, the set-valued formulation provides finite-sample, distribution-free coverage guarantees. Using carefully designed conformity scores, it forms enclosed decision boundaries that improve robustness to non-conforming inputs. Second, the point-valued formulation leverages the same conformity scores to produce a single prediction when desired, potentially improving accuracy over traditional HDC by accounting for class interactions. We demonstrate the broad applicability of the proposed framework through evaluations on multiple real-world datasets. In particular, we apply our method to the challenging problem of decoding non-spatial stimulus information from the spiking activity of hippocampal neurons recorded as subjects performed a sequence memory task. Our results show that ConformalHDC not only accurately decodes the stimulus information represented in the neural activity data, but also provides rigorous uncertainty estimates and correctly abstains when presented with data from other behavioral states. Overall, these capabilities position the framework as a reliable, uncertainty-aware foundation for neuromorphic computing.
Optimal Transport for Latent Integration with An Application to Heterogeneous Neuronal Activity Data
Jun 27, 2024



Abstract:Detecting dynamic patterns of task-specific responses shared across heterogeneous datasets is an essential and challenging problem in many scientific applications in medical science and neuroscience. In our motivating example of rodent electrophysiological data, identifying the dynamical patterns in neuronal activity associated with ongoing cognitive demands and behavior is key to uncovering the neural mechanisms of memory. One of the greatest challenges in investigating a cross-subject biological process is that the systematic heterogeneity across individuals could significantly undermine the power of existing machine learning methods to identify the underlying biological dynamics. In addition, many technically challenging neurobiological experiments are conducted on only a handful of subjects where rich longitudinal data are available for each subject. The low sample sizes of such experiments could further reduce the power to detect common dynamic patterns among subjects. In this paper, we propose a novel heterogeneous data integration framework based on optimal transport to extract shared patterns in complex biological processes. The key advantages of the proposed method are that it can increase discriminating power in identifying common patterns by reducing heterogeneity unrelated to the signal by aligning the extracted latent spatiotemporal information across subjects. Our approach is effective even with a small number of subjects, and does not require auxiliary matching information for the alignment. In particular, our method can align longitudinal data across heterogeneous subjects in a common latent space to capture the dynamics of shared patterns while utilizing temporal dependency within subjects.
Avalanche: an End-to-End Library for Continual Learning
Apr 01, 2021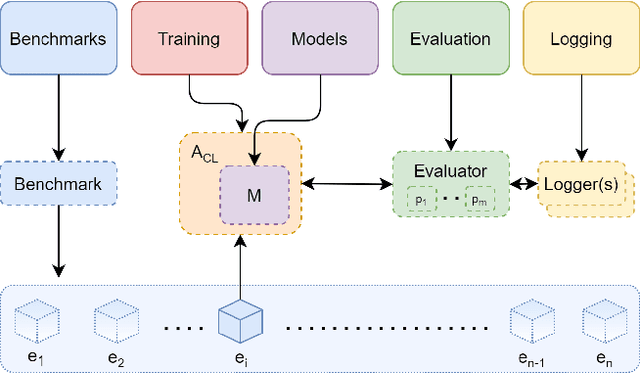
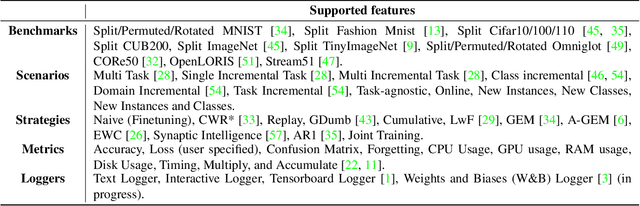
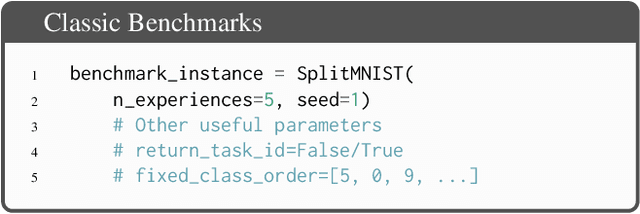

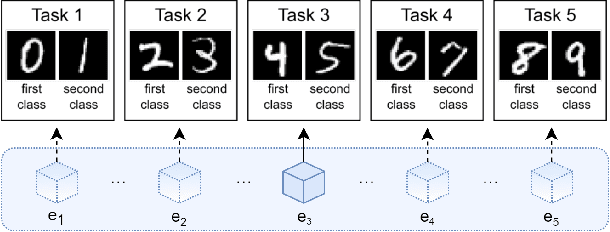

Abstract:Learning continually from non-stationary data streams is a long-standing goal and a challenging problem in machine learning. Recently, we have witnessed a renewed and fast-growing interest in continual learning, especially within the deep learning community. However, algorithmic solutions are often difficult to re-implement, evaluate and port across different settings, where even results on standard benchmarks are hard to reproduce. In this work, we propose Avalanche, an open-source end-to-end library for continual learning research based on PyTorch. Avalanche is designed to provide a shared and collaborative codebase for fast prototyping, training, and reproducible evaluation of continual learning algorithms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge