Julien Cohen-Adad
Impact of individual rater style on deep learning uncertainty in medical imaging segmentation
May 05, 2021
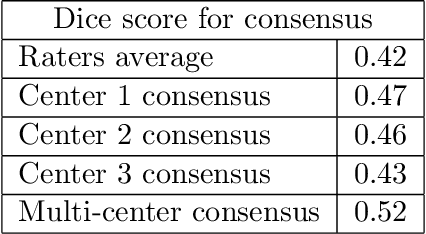
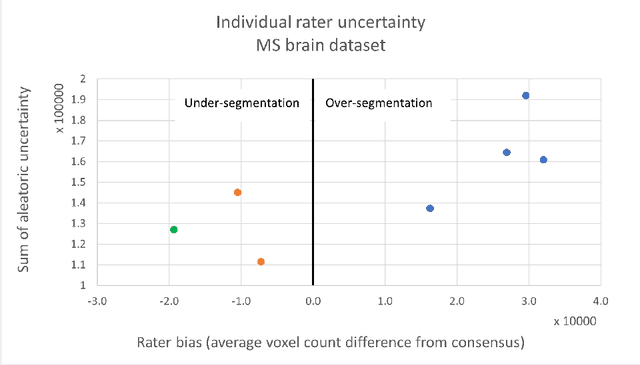
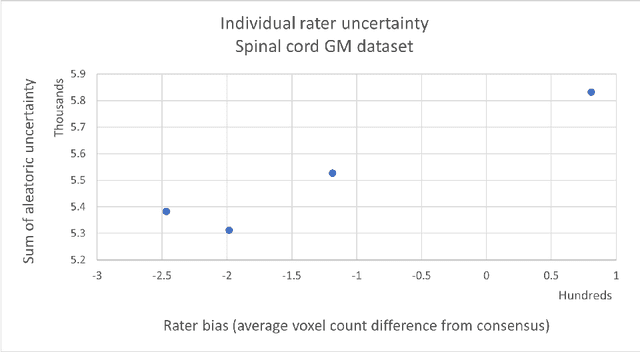
Abstract:While multiple studies have explored the relation between inter-rater variability and deep learning model uncertainty in medical segmentation tasks, little is known about the impact of individual rater style. This study quantifies rater style in the form of bias and consistency and explores their impacts when used to train deep learning models. Two multi-rater public datasets were used, consisting of brain multiple sclerosis lesion and spinal cord grey matter segmentation. On both datasets, results show a correlation ($R^2 = 0.60$ and $0.93$) between rater bias and deep learning uncertainty. The impact of label fusion between raters' annotations on this relationship is also explored, and we show that multi-center consensuses are more effective than single-center consensuses to reduce uncertainty, since rater style is mostly center-specific.
Benefits of Linear Conditioning for Segmentation using Metadata
Feb 18, 2021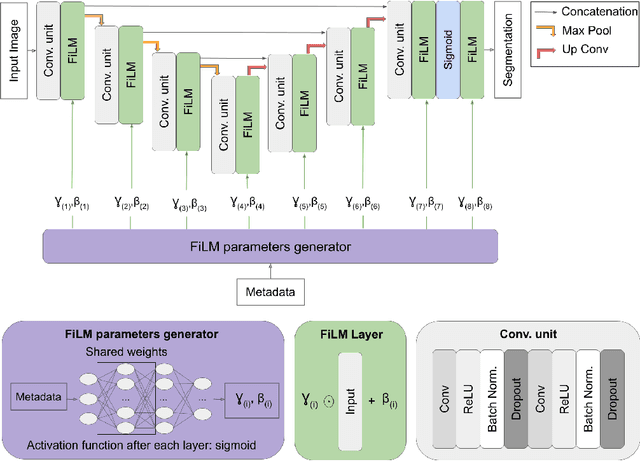
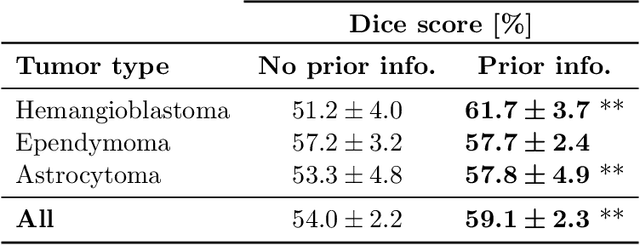
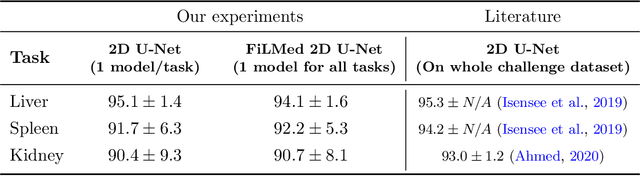
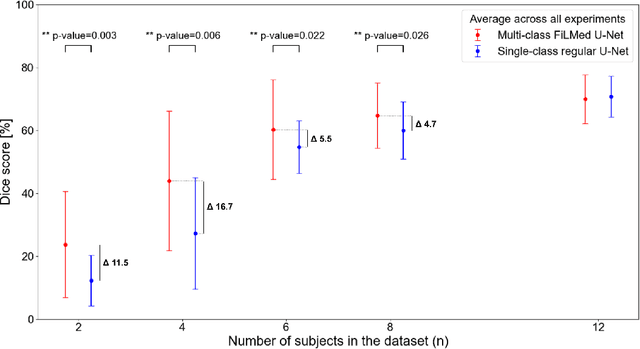
Abstract:Medical images are often accompanied by metadata describing the image (vendor, acquisition parameters) and the patient (disease type or severity, demographics, genomics). This metadata is usually disregarded by image segmentation methods. In this work, we adapt a linear conditioning method called FiLM (Feature-wise Linear Modulation) for image segmentation tasks. This FiLM adaptation enables integrating metadata into segmentation models for better performance. We observed an average Dice score increase of 5.1% on spinal cord tumor segmentation when incorporating the tumor type with FiLM. The metadata modulates the segmentation process through low-cost affine transformations applied on feature maps which can be included in any neural network's architecture. Additionally, we assess the relevance of segmentation FiLM layers for tackling common challenges in medical imaging: training with limited or unbalanced number of annotated data, multi-class training with missing segmentations, and model adaptation to multiple tasks. Our results demonstrated the following benefits of FiLM for segmentation: FiLMed U-Net was robust to missing labels and reached higher Dice scores with few labels (up to 16.7%) compared to single-task U-Net. The code is open-source and available at www.ivadomed.org.
Multiclass Spinal Cord Tumor Segmentation on MRI with Deep Learning
Jan 14, 2021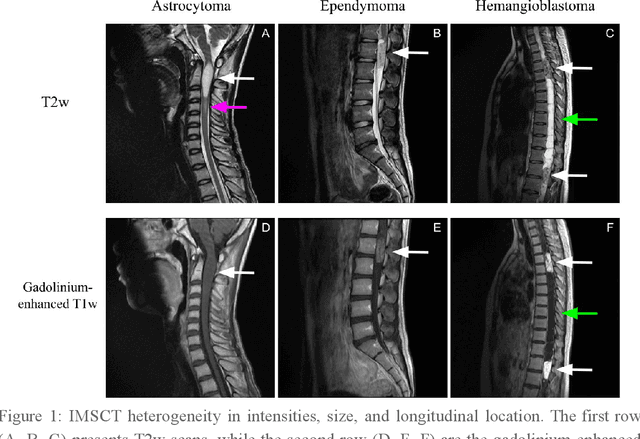
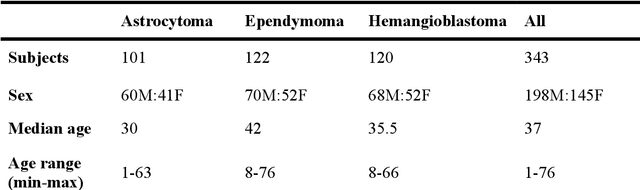

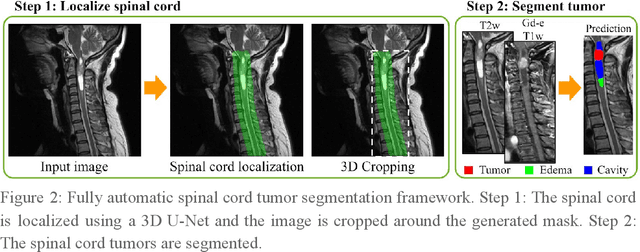
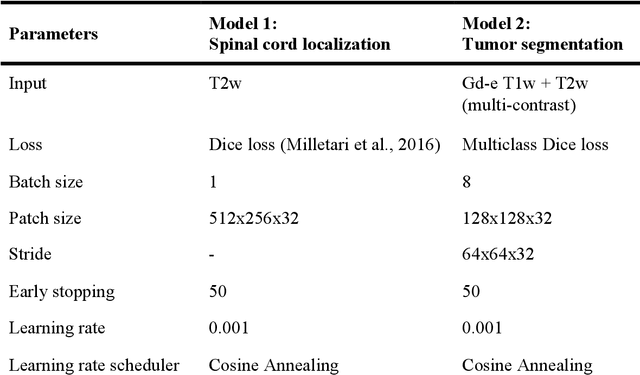
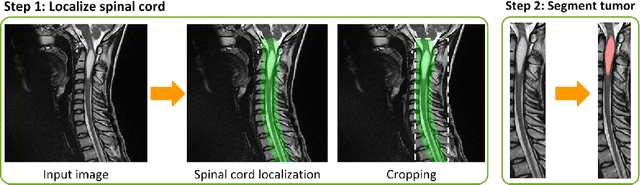
Abstract:Spinal cord tumors lead to neurological morbidity and mortality. Being able to obtain morphometric quantification (size, location, growth rate) of the tumor, edema, and cavity can result in improved monitoring and treatment planning. Such quantification requires the segmentation of these structures into three separate classes. However, manual segmentation of 3-dimensional structures is time-consuming and tedious, motivating the development of automated methods. Here, we tailor a model adapted to the spinal cord tumor segmentation task. Data were obtained from 343 patients using gadolinium-enhanced T1-weighted and T2-weighted MRI scans with cervical, thoracic, and/or lumbar coverage. The dataset includes the three most common intramedullary spinal cord tumor types: astrocytomas, ependymomas, and hemangioblastomas. The proposed approach is a cascaded architecture with U-Net-based models that segments tumors in a two-stage process: locate and label. The model first finds the spinal cord and generates bounding box coordinates. The images are cropped according to this output, leading to a reduced field of view, which mitigates class imbalance. The tumor is then segmented. The segmentation of the tumor, cavity, and edema (as a single class) reached 76.7 $\pm$ 1.5% of Dice score and the segmentation of tumors alone reached 61.8 $\pm$ 4.0% Dice score. The true positive detection rate was above 87% for tumor, edema, and cavity. To the best of our knowledge, this is the first fully automatic deep learning model for spinal cord tumor segmentation. The multiclass segmentation pipeline is available in the Spinal Cord Toolbox (https://spinalcordtoolbox.com/). It can be run with custom data on a regular computer within seconds.
SoftSeg: Advantages of soft versus binary training for image segmentation
Nov 18, 2020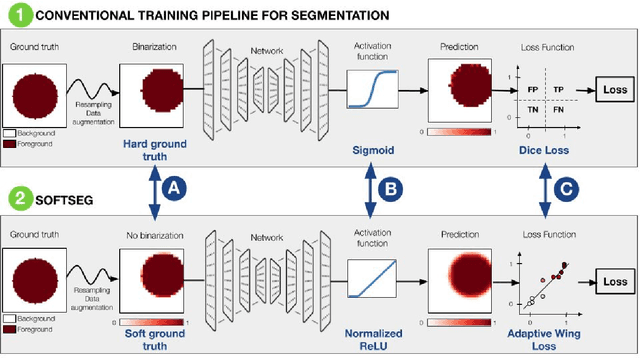
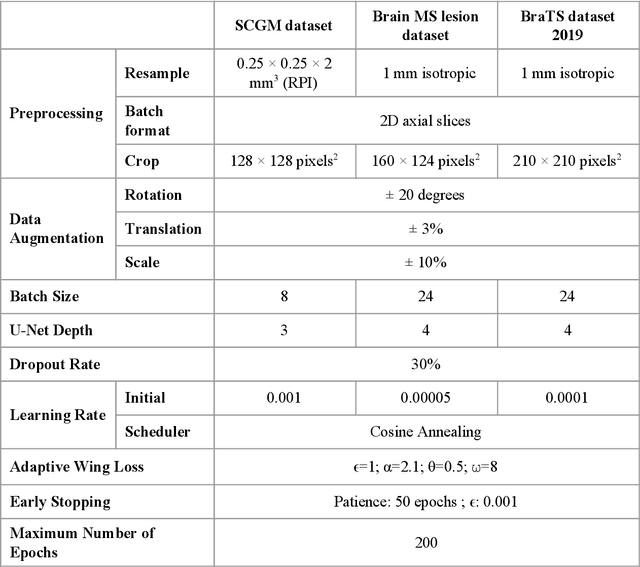
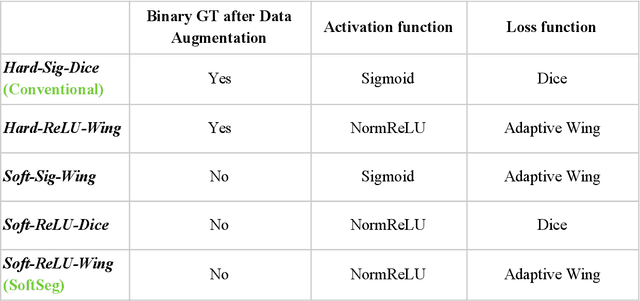
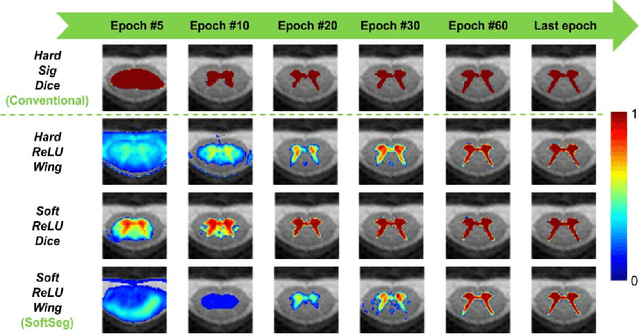
Abstract:Most image segmentation algorithms are trained on binary masks formulated as a classification task per pixel. However, in applications such as medical imaging, this "black-and-white" approach is too constraining because the contrast between two tissues is often ill-defined, i.e., the voxels located on objects' edges contain a mixture of tissues. Consequently, assigning a single "hard" label can result in a detrimental approximation. Instead, a soft prediction containing non-binary values would overcome that limitation. We introduce SoftSeg, a deep learning training approach that takes advantage of soft ground truth labels, and is not bound to binary predictions. SoftSeg aims at solving a regression instead of a classification problem. This is achieved by using (i) no binarization after preprocessing and data augmentation, (ii) a normalized ReLU final activation layer (instead of sigmoid), and (iii) a regression loss function (instead of the traditional Dice loss). We assess the impact of these three features on three open-source MRI segmentation datasets from the spinal cord gray matter, the multiple sclerosis brain lesion, and the multimodal brain tumor segmentation challenges. Across multiple cross-validation iterations, SoftSeg outperformed the conventional approach, leading to an increase in Dice score of 2.0% on the gray matter dataset (p=0.001), 3.3% for the MS lesions, and 6.5% for the brain tumors. SoftSeg produces consistent soft predictions at tissues' interfaces and shows an increased sensitivity for small objects. The richness of soft labels could represent the inter-expert variability, the partial volume effect, and complement the model uncertainty estimation. The developed training pipeline can easily be incorporated into most of the existing deep learning architectures. It is already implemented in the freely-available deep learning toolbox ivadomed (https://ivadomed.org).
ivadomed: A Medical Imaging Deep Learning Toolbox
Oct 20, 2020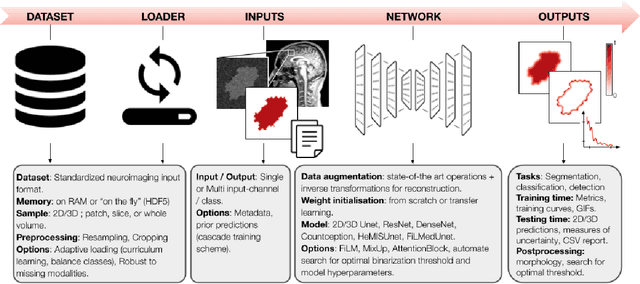
Abstract:ivadomed is an open-source Python package for designing, end-to-end training, and evaluating deep learning models applied to medical imaging data. The package includes APIs, command-line tools, documentation, and tutorials. ivadomed also includes pre-trained models such as spinal tumor segmentation and vertebral labeling. Original features of ivadomed include a data loader that can parse image metadata (e.g., acquisition parameters, image contrast, resolution) and subject metadata (e.g., pathology, age, sex) for custom data splitting or extra information during training and evaluation. Any dataset following the Brain Imaging Data Structure (BIDS) convention will be compatible with ivadomed without the need to manually organize the data, which is typically a tedious task. Beyond the traditional deep learning methods, ivadomed features cutting-edge architectures, such as FiLM and HeMis, as well as various uncertainty estimation methods (aleatoric and epistemic), and losses adapted to imbalanced classes and non-binary predictions. Each step is conveniently configurable via a single file. At the same time, the code is highly modular to allow addition/modification of an architecture or pre/post-processing steps. Example applications of ivadomed include MRI object detection, segmentation, and labeling of anatomical and pathological structures. Overall, ivadomed enables easy and quick exploration of the latest advances in deep learning for medical imaging applications. ivadomed's main project page is available at https://ivadomed.org.
Spine intervertebral disc labeling using a fully convolutional redundant counting model
Mar 11, 2020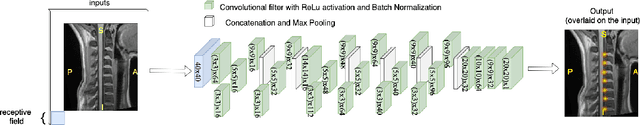
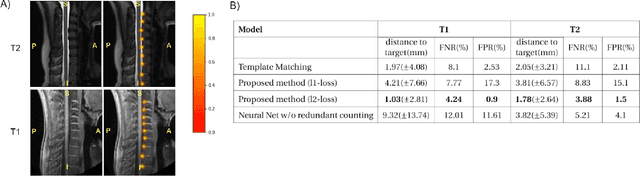
Abstract:Labeling intervertebral discs is relevant as it notably enables clinicians to understand the relationship between a patient's symptoms (pain, paralysis) and the exact level of spinal cord injury. However manually labeling those discs is a tedious and user-biased task which would benefit from automated methods. While some automated methods already exist for MRI and CT-scan, they are either not publicly available, or fail to generalize across various imaging contrasts. In this paper we combine a Fully Convolutional Network (FCN) with inception modules to localize and label intervertebral discs. We demonstrate a proof-of-concept application in a publicly-available multi-center and multi-contrast MRI database (n=235 subjects). The code is publicly available at https://github.com/neuropoly/vertebral-labeling-deep-learning.
Automatic segmentation of spinal multiple sclerosis lesions: How to generalize across MRI contrasts?
Mar 11, 2020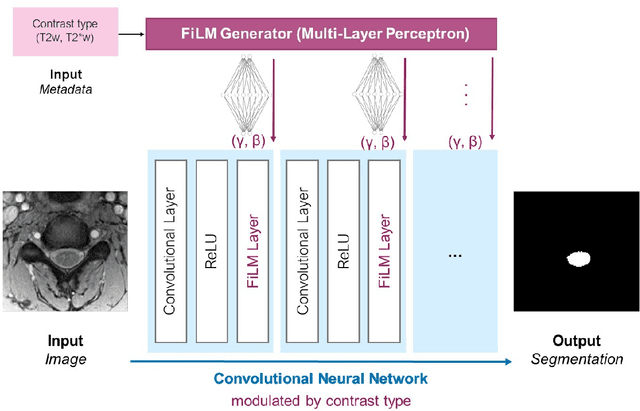
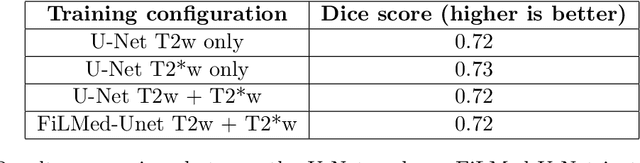
Abstract:Despite recent improvements in medical image segmentation, the ability to generalize across imaging contrasts remains an open issue. To tackle this challenge, we implement Feature-wise Linear Modulation (FiLM) to leverage physics knowledge within the segmentation model and learn the characteristics of each contrast. Interestingly, a well-optimised U-Net reached the same performance as our FiLMed-Unet on a multi-contrast dataset (0.72 of Dice score), which suggests that there is a bottleneck in spinal MS lesion segmentation different from the generalization across varying contrasts. This bottleneck likely stems from inter-rater variability, which is estimated at 0.61 of Dice score in our dataset.
Deep Semantic Segmentation of Natural and Medical Images: A Review
Nov 02, 2019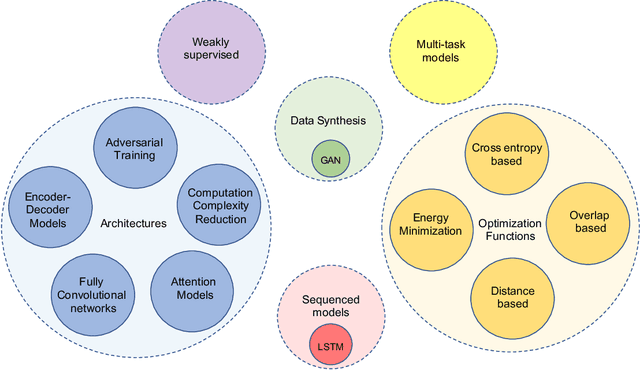
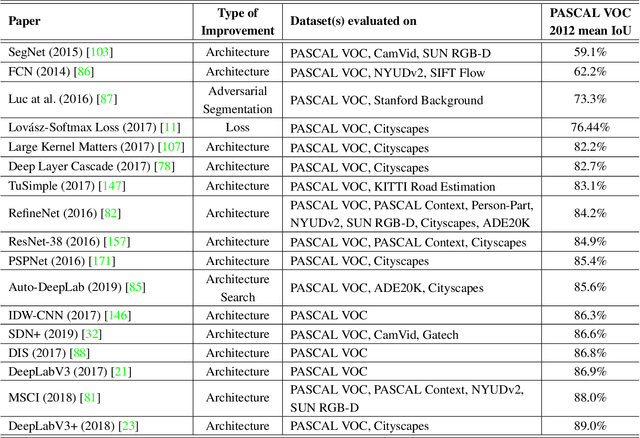
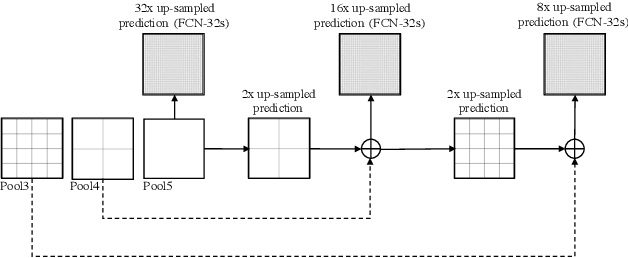
Abstract:The (medical) image semantic segmentation task consists of classifying each pixel of an image (or just several ones) into an instance, where each instance (or category) corresponding to a class. This task is a part of the concept of scene understanding or better explaining the global context of an image. In the medical image analysis domain, image segmentation can be used for image-guided interventions, radiotherapy, or improved radiological diagnostics. In this review, we categorize the main deep learning-based medical and non-medical image segmentation solutions into six main groups of deep architectural improvements, data synthesis-based, loss function-based improvements, sequenced models, weakly supervised, and multi-task methods and further for each group we analyzed each variant of these groups and discuss limitations of the current approaches and future research directions for semantic image segmentation.
U-Net Fixed-Point Quantization for Medical Image Segmentation
Sep 09, 2019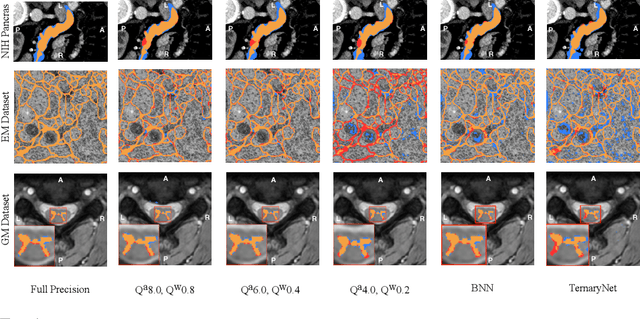
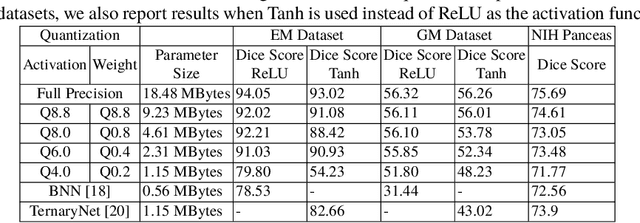
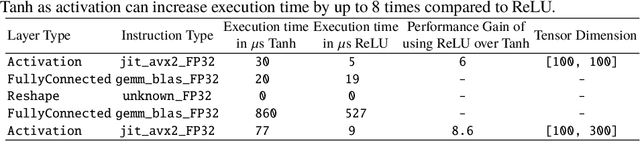
Abstract:Model quantization is leveraged to reduce the memory consumption and the computation time of deep neural networks. This is achieved by representing weights and activations with a lower bit resolution when compared to their high precision floating point counterparts. The suitable level of quantization is directly related to the model performance. Lowering the quantization precision (e.g. 2 bits), reduces the amount of memory required to store model parameters and the amount of logic required to implement computational blocks, which contributes to reducing the power consumption of the entire system. These benefits typically come at the cost of reduced accuracy. The main challenge is to quantize a network as much as possible, while maintaining the performance accuracy. In this work, we present a quantization method for the U-Net architecture, a popular model in medical image segmentation. We then apply our quantization algorithm to three datasets: (1) the Spinal Cord Gray Matter Segmentation (GM), (2) the ISBI challenge for segmentation of neuronal structures in Electron Microscopic (EM), and (3) the public National Institute of Health (NIH) dataset for pancreas segmentation in abdominal CT scans. The reported results demonstrate that with only 4 bits for weights and 6 bits for activations, we obtain 8 fold reduction in memory requirements while loosing only 2.21%, 0.57% and 2.09% dice overlap score for EM, GM and NIH datasets respectively. Our fixed point quantization provides a flexible trade off between accuracy and memory requirement which is not provided by previous quantization methods for U-Net such as TernaryNet.
Deep Active Learning for Axon-Myelin Segmentation on Histology Data
Jul 11, 2019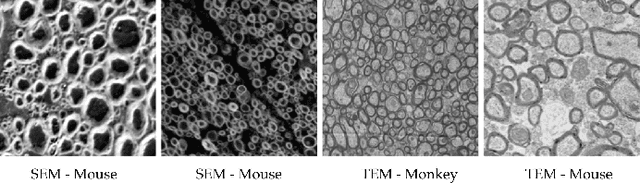
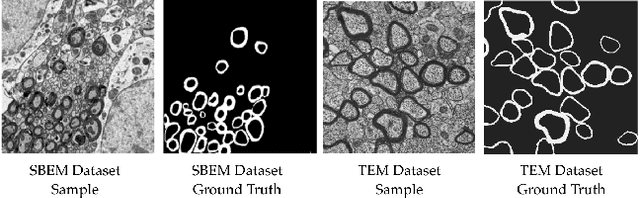
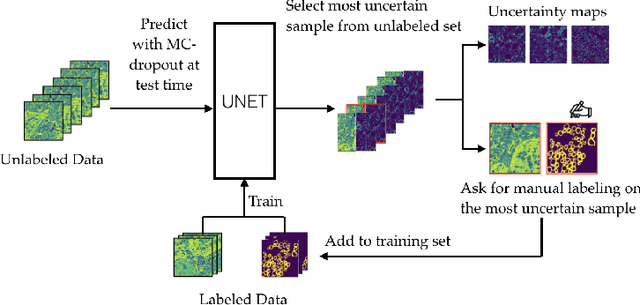
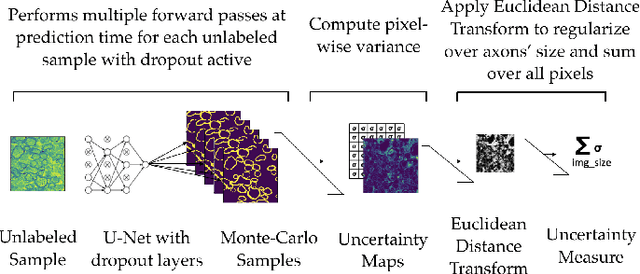

Abstract:Semantic segmentation is a crucial task in biomedical image processing, which recent breakthroughs in deep learning have allowed to improve. However, deep learning methods in general are not yet widely used in practice since they require large amount of data for training complex models. This is particularly challenging for biomedical images, because data and ground truths are a scarce resource. Annotation efforts for biomedical images come with a real cost, since experts have to manually label images at pixel-level on samples usually containing many instances of the target anatomy (e.g. in histology samples: neurons, astrocytes, mitochondria, etc.). In this paper we provide a framework for Deep Active Learning applied to a real-world scenario. Our framework relies on the U-Net architecture and overall uncertainty measure to suggest which sample to annotate. It takes advantage of the uncertainty measure obtained by taking Monte Carlo samples while using Dropout regularization scheme. Experiments were done on spinal cord and brain microscopic histology samples to perform a myelin segmentation task. Two realistic small datasets of 14 and 24 images were used, from different acquisition settings (Serial Block-Face Electron Microscopy and Transmitting Electron Microscopy) and showed that our method reached a maximum Dice value after adding 3 uncertainty-selected samples to the initial training set, versus 15 randomly-selected samples, thereby significantly reducing the annotation effort. We focused on a plausible scenario and showed evidence that this straightforward implementation achieves a high segmentation performance with very few labelled samples. We believe our framework may benefit any biomedical researcher willing to obtain fast and accurate image segmentation on their own dataset. The code is freely available at https://github.com/neuropoly/deep-active-learning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge