Joshua T. Vogelstein
Teresa
A New Age of Computing and the Brain
Apr 27, 2020
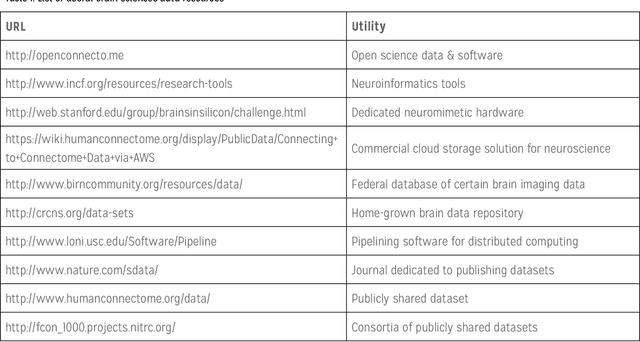
Abstract:The history of computer science and brain sciences are intertwined. In his unfinished manuscript "The Computer and the Brain," von Neumann debates whether or not the brain can be thought of as a computing machine and identifies some of the similarities and differences between natural and artificial computation. Turing, in his 1950 article in Mind, argues that computing devices could ultimately emulate intelligence, leading to his proposed Turing test. Herbert Simon predicted in 1957 that most psychological theories would take the form of a computer program. In 1976, David Marr proposed that the function of the visual system could be abstracted and studied at computational and algorithmic levels that did not depend on the underlying physical substrate. In December 2014, a two-day workshop supported by the Computing Community Consortium (CCC) and the National Science Foundation's Computer and Information Science and Engineering Directorate (NSF CISE) was convened in Washington, DC, with the goal of bringing together computer scientists and brain researchers to explore these new opportunities and connections, and develop a new, modern dialogue between the two research communities. Specifically, our objectives were: 1. To articulate a conceptual framework for research at the interface of brain sciences and computing and to identify key problems in this interface, presented in a way that will attract both CISE and brain researchers into this space. 2. To inform and excite researchers within the CISE research community about brain research opportunities and to identify and explain strategic roles they can play in advancing this initiative. 3. To develop new connections, conversations and collaborations between brain sciences and CISE researchers that will lead to highly relevant and competitive proposals, high-impact research, and influential publications.
The Chi-Square Test of Distance Correlation
Jan 22, 2020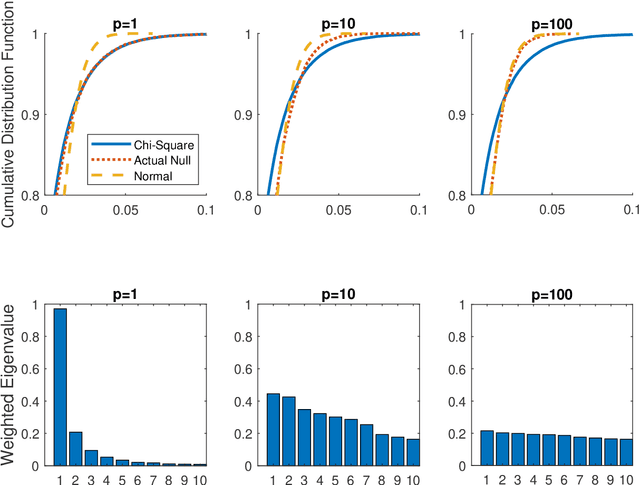
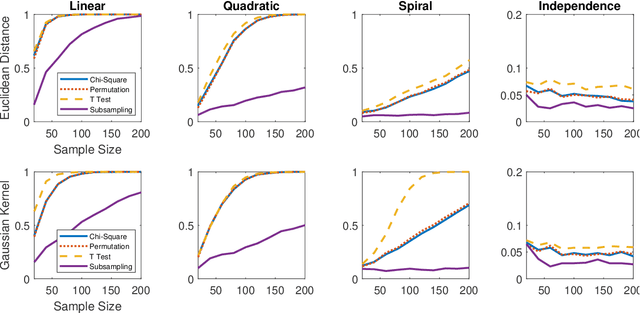
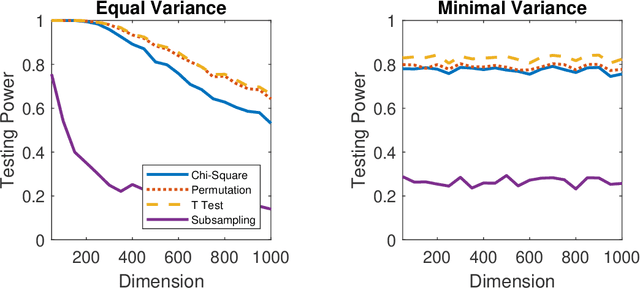
Abstract:Distance correlation has gained much recent attention in the data science community: the sample statistic is straightforward to compute and asymptotically equals zero if and only if independence, making it an ideal choice to test any type of dependency structure given sufficient sample size. One major bottleneck is the testing process: because the null distribution of distance correlation depends on the underlying random variables and metric choice, it typically requires a permutation test to estimate the null and compute the p-value, which is very costly for large amount of data. To overcome the difficulty, we propose a centered chi-square distribution, demonstrate it well-approximates the null distribution of unbiased distance correlation, and prove upper tail dominance and distribution bound between them. The resulting distance correlation chi-square test is a nonparametric test for independence, is valid and universally consistent using any strong negative type metric or characteristic kernel, enjoys a similar finite-sample testing power as the standard permutation test, and is provably the most powerful test of distance correlation among all valid tests with known distribution.
The Exact Equivalence of Independence Testing and Two-Sample Testing
Oct 20, 2019
Abstract:Testing independence and testing equality of distributions are two tightly related statistical hypotheses. Several distance and kernel-based statistics are recently proposed to achieve universally consistent testing for either hypothesis. On the distance side, the distance correlation is proposed for independence testing, and the energy statistic is proposed for two-sample testing. On the kernel side, the Hilbert-Schmidt independence criterion is proposed for independence testing and the maximum mean discrepancy is proposed for two-sample testing. In this paper, we show that two-sample testing are special cases of independence testing via an auxiliary label vector, and prove that distance correlation is exactly equivalent to the energy statistic in terms of the population statistic, the sample statistic, and the testing p-value via permutation test. The equivalence can be further generalized to K-sample testing and extended to the kernel regime. As a consequence, it suffices to always use an independence statistic to test equality of distributions, which enables better interpretability of the test statistic and more efficient testing.
AutoGMM: Automatic Gaussian Mixture Modeling in Python
Oct 04, 2019
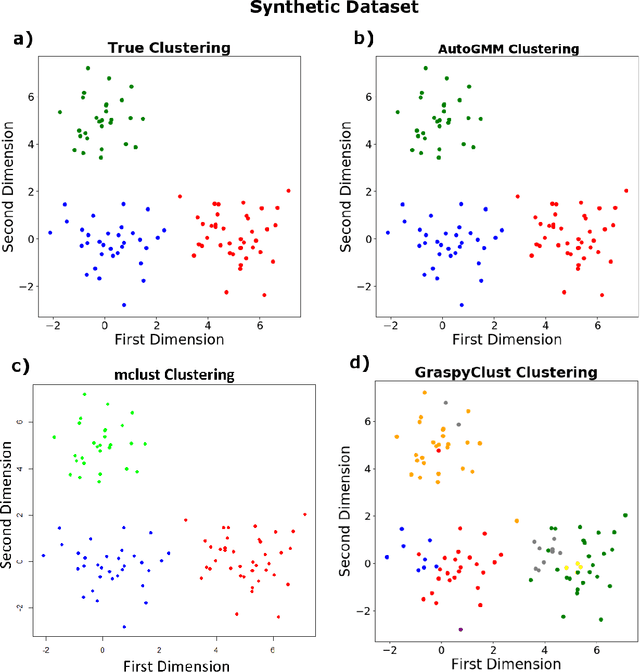
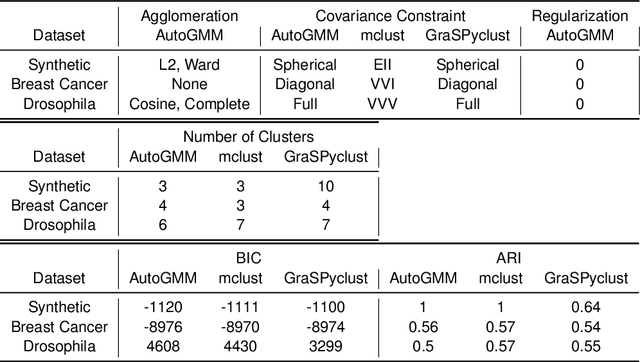
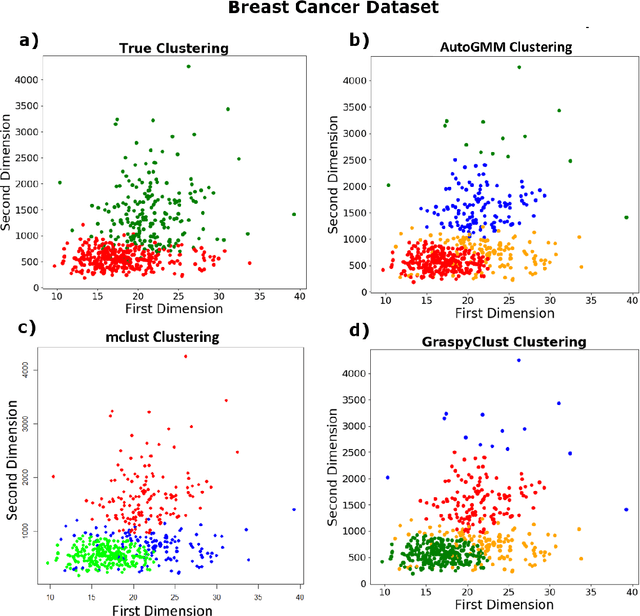
Abstract:Gaussian mixture modeling is a fundamental tool in clustering, as well as discriminant analysis and semiparametric density estimation. However, estimating the optimal model for any given number of components is an NP-hard problem, and estimating the number of components is in some respects an even harder problem. In R, a popular package called mclust addresses both of these problems. However, Python has lacked such a package. We therefore introduce AutoGMM, a Python algorithm for automatic Gaussian mixture modeling. AutoGMM builds upon scikit-learn's AgglomerativeClustering and GaussianMixture classes, with certain modifications to make the results more stable. Empirically, on several different applications, AutoGMM performs approximately as well as mclust. This algorithm is freely available and therefore further shrinks the gap between functionality of R and Python for data science.
Manifold Forests: Closing the Gap on Neural Networks
Sep 25, 2019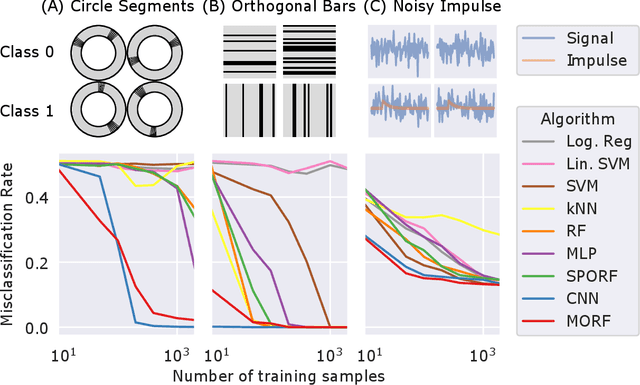
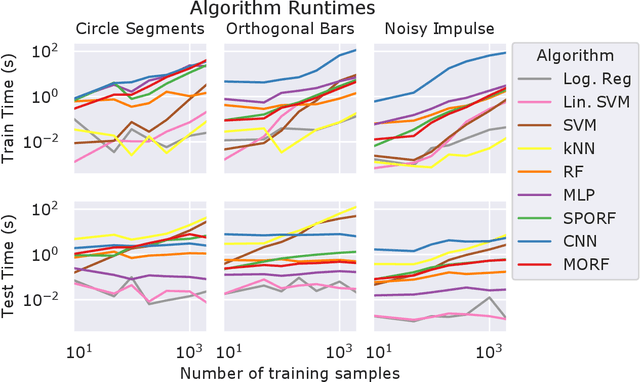
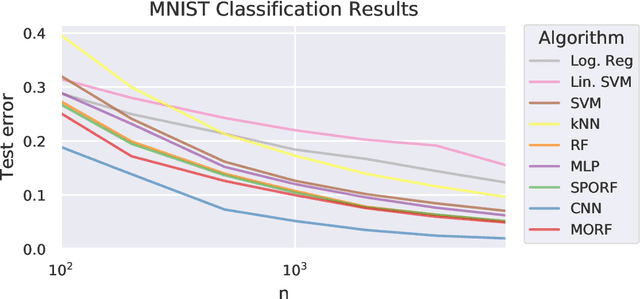
Abstract:Decision forests (DF), in particular random forests and gradient boosting trees, have demonstrated state-of-the-art accuracy compared to other methods in many supervised learning scenarios. In particular, DFs dominate other methods in tabular data, that is, when the feature space is unstructured, so that the signal is invariant to permuting feature indices. However, in structured data lying on a manifold---such as images, text, and speech---neural nets (NN) tend to outperform DFs. We conjecture that at least part of the reason for this is that the input to NN is not simply the feature magnitudes, but also their indices (for example, the convolution operation uses "feature locality"). In contrast, na\"ive DF implementations fail to explicitly consider feature indices. A recently proposed DF approach demonstrates that DFs, for each node, implicitly sample a random matrix from some specific distribution. Here, we build on that to show that one can choose distributions in a \emph{manifold aware fashion}. For example, for image classification, rather than randomly selecting pixels, one can randomly select contiguous patches. We demonstrate the empirical performance of data living on three different manifolds: images, time-series, and a torus. In all three cases, our Manifold Forest (\Mf) algorithm empirically dominates other state-of-the-art approaches that ignore feature space structure, achieving a lower classification error on all sample sizes. This dominance extends to the MNIST data set as well. Moreover, both training and test time is significantly faster for manifold forests as compared to deep nets. This approach, therefore, has promise to enable DFs and other machine learning methods to close the gap with deep nets on manifold-valued data.
mgcpy: A Comprehensive High Dimensional Independence Testing Python Package
Jul 18, 2019
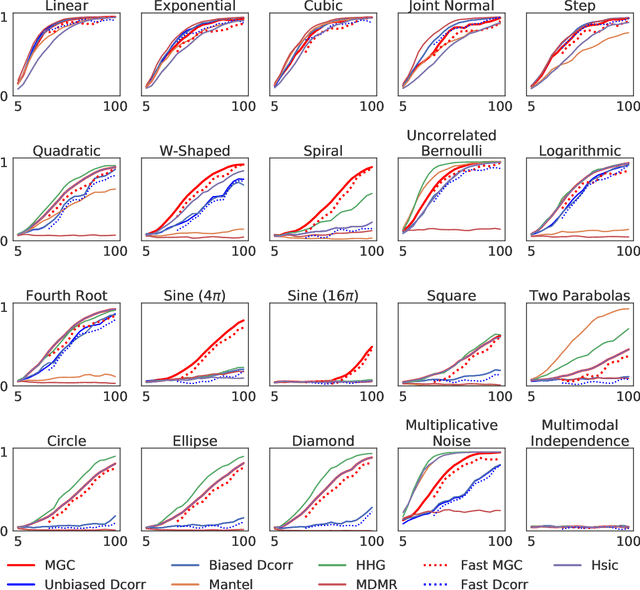
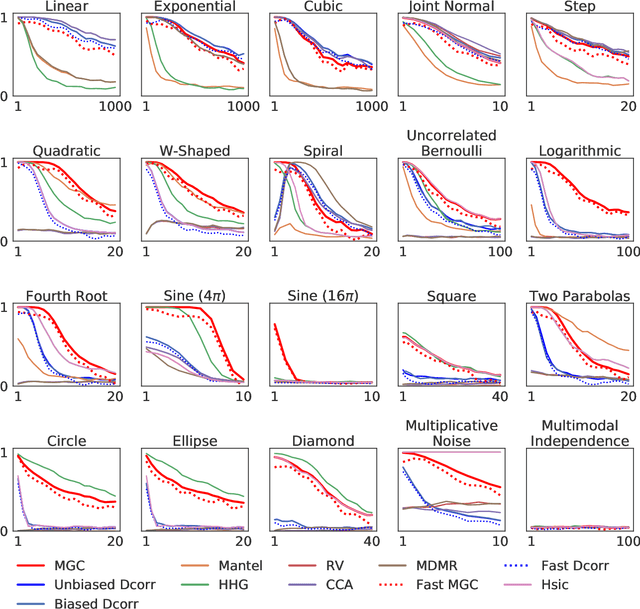
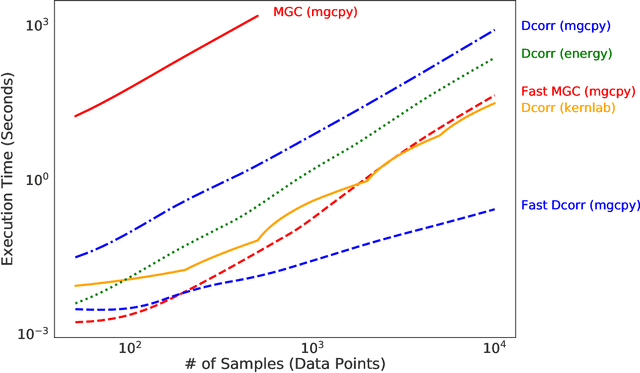
Abstract:With the increase in the amount of data in many fields, a method to consistently and efficiently decipher relationships within high dimensional data sets is important. Because many modern datasets are high-dimensional, univariate independence tests are not applicable. While many multivariate independence tests have R packages available, the interfaces are inconsistent, most are not available in Python. mgcpy is an extensive Python library that includes many state of the art high-dimensional independence testing procedures using a common interface. The package is easy-to-use and is flexible enough to enable future extensions. This manuscript provides details for each of the tests as well as extensive power and run-time benchmarks on a suite of high-dimensional simulations previously used in different publications. The appendix includes demonstrations of how the user can interact with the package, as well as links and documentation.
Geodesic Learning via Unsupervised Decision Forests
Jul 05, 2019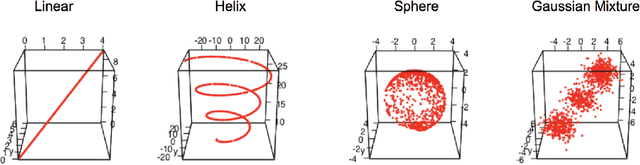
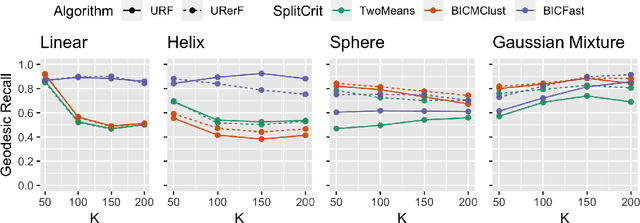
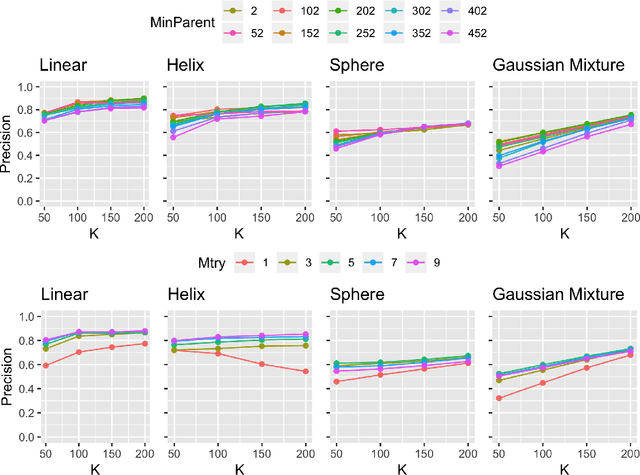
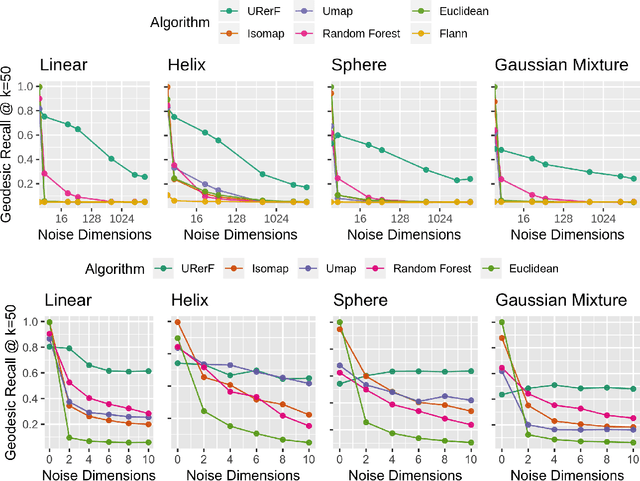
Abstract:Geodesic distance is the shortest path between two points in a Riemannian manifold. Manifold learning algorithms, such as Isomap, seek to learn a manifold that preserves geodesic distances. However, such methods operate on the ambient dimensionality, and are therefore fragile to noise dimensions. We developed an unsupervised random forest method (URerF) to approximately learn geodesic distances in linear and nonlinear manifolds with noise. URerF operates on low-dimensional sparse linear combinations of features, rather than the full observed dimensionality. To choose the optimal split in a computationally efficient fashion, we developed a fast Bayesian Information Criterion statistic for Gaussian mixture models. We introduce geodesic precision-recall curves which quantify performance relative to the true latent manifold. Empirical results on simulated and real data demonstrate that URerF is robust to high-dimensional noise, where as other methods, such as Isomap, UMAP, and FLANN, quickly deteriorate in such settings. In particular, URerF is able to estimate geodesic distances on a real connectome dataset better than other approaches.
Estimating Information-Theoretic Quantities with Random Forests
Jul 03, 2019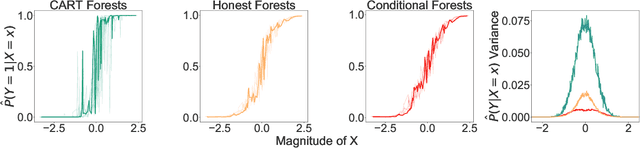
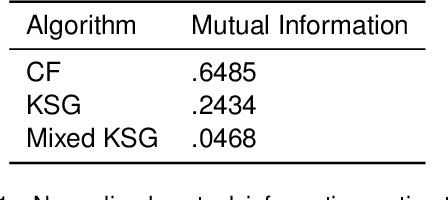
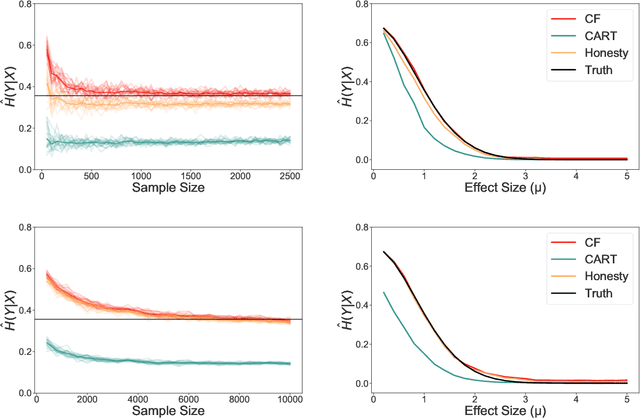

Abstract:Information-theoretic quantities, such as mutual information and conditional entropy, are useful statistics for measuring the dependence between two random variables. However, estimating these quantities in a non-parametric fashion is difficult, especially when the variables are high-dimensional, a mixture of continuous and discrete values, or both. In this paper, we propose a decision forest method, Conditional Forests (CF), to estimate these quantities. By combining quantile regression forests with honest sampling, and introducing a finite sample correction, CF improves finite sample bias in a range of settings. We demonstrate through simulations that CF achieves smaller bias and variance in both low- and high-dimensional settings for estimating posteriors, conditional entropy, and mutual information. We then use CF to estimate the amount of information between neuron class and other ceulluar feautres.
GraSPy: Graph Statistics in Python
Mar 29, 2019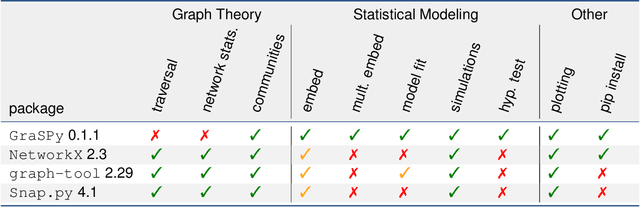
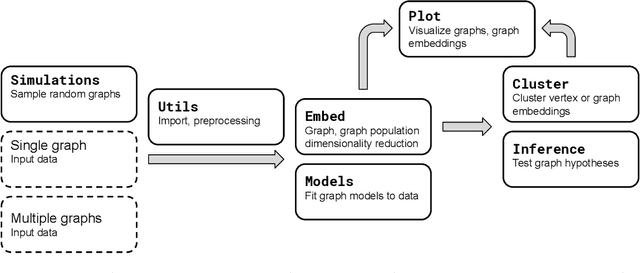
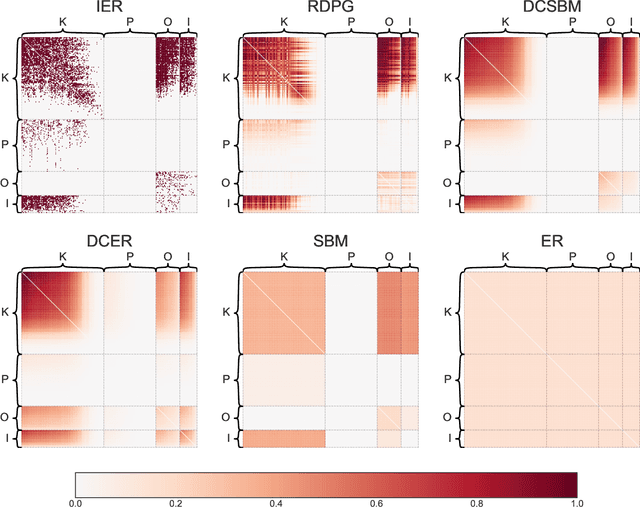
Abstract:We introduce GraSPy, a Python library devoted to statistical inference, machine learning, and visualization of random graphs and graph populations. This package provides flexible and easy-to-use algorithms for analyzing and understanding graphs with a scikit-learn compliant API. GraSPy can be downloaded from Python Package Index (PyPi), and is released under the Apache 2.0 open-source license. The documentation and all releases are available at https://neurodata.io/graspy.
Decision Forests Induce Characteristic Kernels
Nov 30, 2018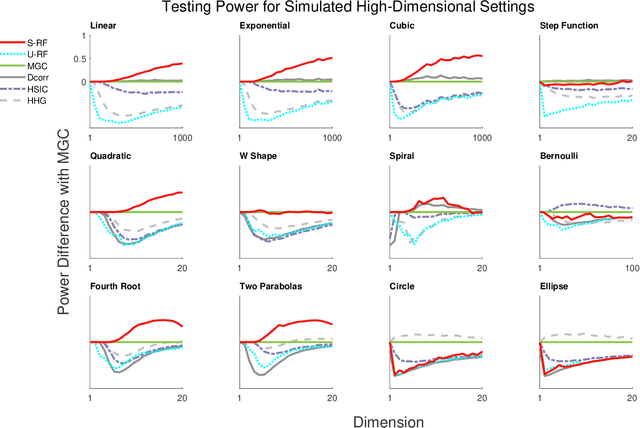
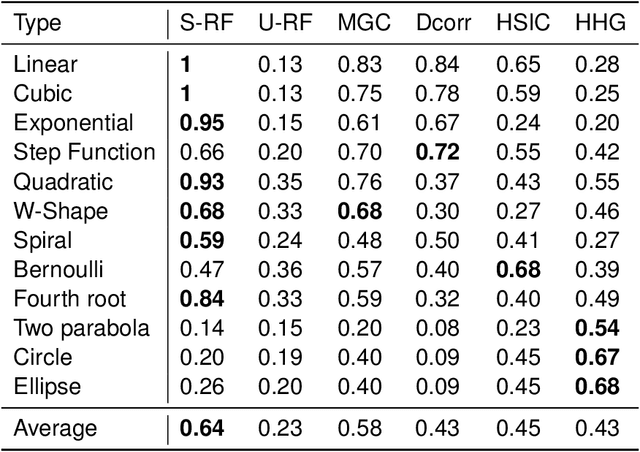
Abstract:Decision forests are popular tools for classification and regression. These forests naturally produce proximity matrices measuring how often each pair of observations lies in the same leaf node. Recently it has been demonstrated that these proximity matrices can be thought of as kernels, connecting the decision forest literature to the extensive kernel machine literature. While other kernels are known to have strong theoretical properties, such as being characteristic kernels, no similar result is available for any decision forest based kernel. We show that a decision forest induced proximity can be made into a characteristic kernel, which can be used within an independence test to obtain a universally consistent test. We therefore empirically evaluate this kernel on a suite of 12 high-dimensional independence test settings: the decision forest induced kernel is shown to typically achieve substantially higher power than other methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge