Jose Dolz
École de Technologie Supérieure, Montreal, Canada
Looking at the whole picture: constrained unsupervised anomaly segmentation
Sep 01, 2021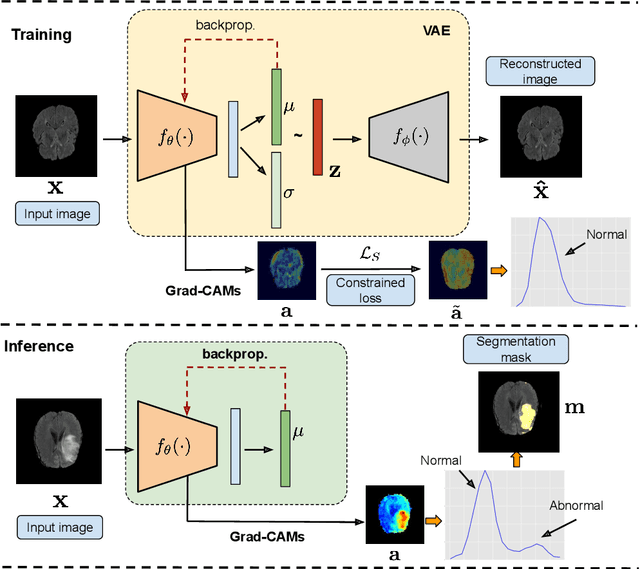
Abstract:Current unsupervised anomaly localization approaches rely on generative models to learn the distribution of normal images, which is later used to identify potential anomalous regions derived from errors on the reconstructed images. However, a main limitation of nearly all prior literature is the need of employing anomalous images to set a class-specific threshold to locate the anomalies. This limits their usability in realistic scenarios, where only normal data is typically accessible. Despite this major drawback, only a handful of works have addressed this limitation, by integrating supervision on attention maps during training. In this work, we propose a novel formulation that does not require accessing images with abnormalities to define the threshold. Furthermore, and in contrast to very recent work, the proposed constraint is formulated in a more principled manner, leveraging well-known knowledge in constrained optimization. In particular, the equality constraint on the attention maps in prior work is replaced by an inequality constraint, which allows more flexibility. In addition, to address the limitations of penalty-based functions we employ an extension of the popular log-barrier methods to handle the constraint. Comprehensive experiments on the popular BRATS'19 dataset demonstrate that the proposed approach substantially outperforms relevant literature, establishing new state-of-the-art results for unsupervised lesion segmentation.
Source-Free Domain Adaptation for Image Segmentation
Aug 06, 2021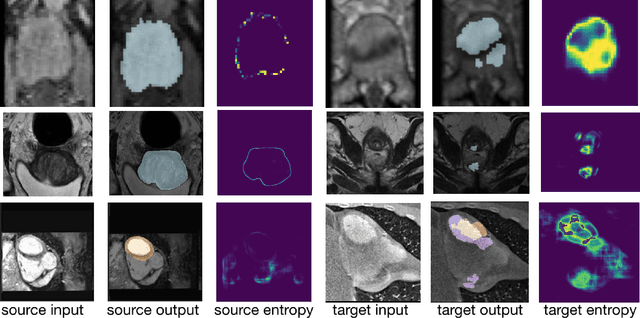
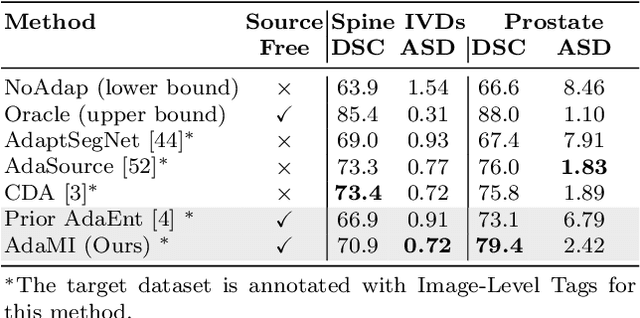
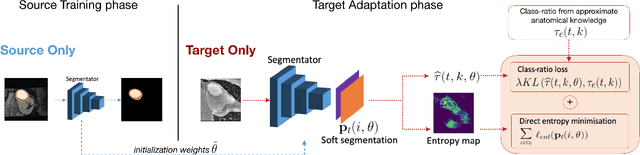
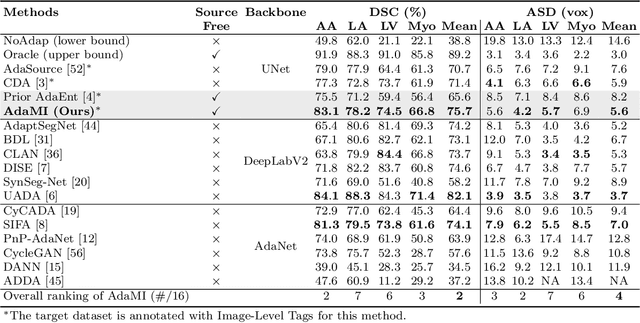
Abstract:Domain adaptation (DA) has drawn high interest for its capacity to adapt a model trained on labeled source data to perform well on unlabeled or weakly labeled target data from a different domain. Most common DA techniques require concurrent access to the input images of both the source and target domains. However, in practice, privacy concerns often impede the availability of source images in the adaptation phase. This is a very frequent DA scenario in medical imaging, where, for instance, the source and target images could come from different clinical sites. We introduce a source-free domain adaptation for image segmentation. Our formulation is based on minimizing a label-free entropy loss defined over target-domain data, which we further guide with a domain-invariant prior on the segmentation regions. Many priors can be derived from anatomical information. Here, a class ratio prior is estimated from anatomical knowledge and integrated in the form of a Kullback Leibler (KL) divergence in our overall loss function. Furthermore, we motivate our overall loss with an interesting link to maximizing the mutual information between the target images and their label predictions. We show the effectiveness of our prior aware entropy minimization in a variety of domain-adaptation scenarios, with different modalities and applications, including spine, prostate, and cardiac segmentation. Our method yields comparable results to several state of the art adaptation techniques, despite having access to much less information, as the source images are entirely absent in our adaptation phase. Our straightforward adaptation strategy uses only one network, contrary to popular adversarial techniques, which are not applicable to a source-free DA setting. Our framework can be readily used in a breadth of segmentation problems, and our code is publicly available: https://github.com/mathilde-b/SFDA
Mutual-Information Based Few-Shot Classification
Jun 23, 2021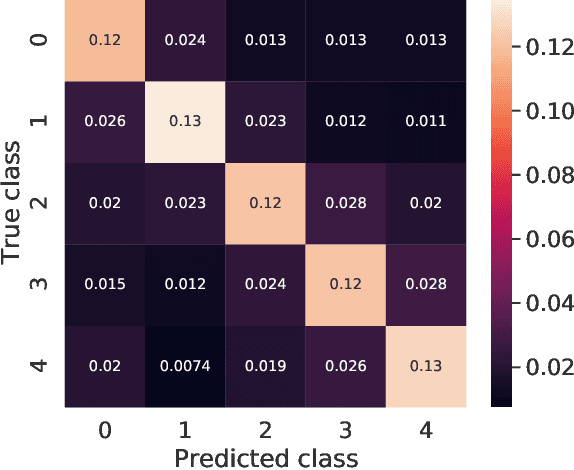
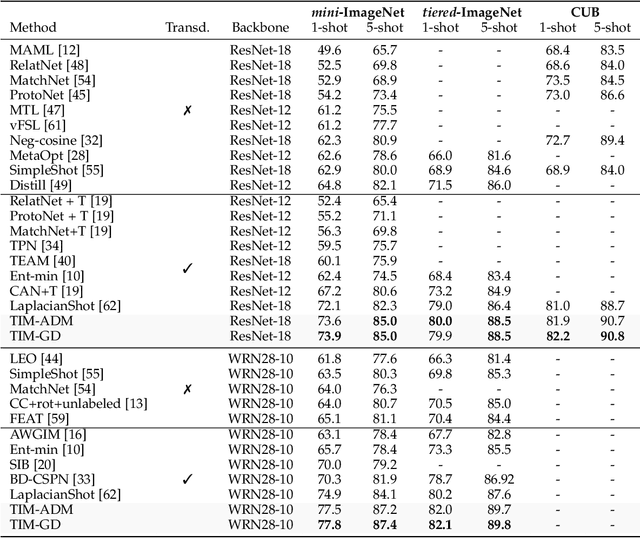
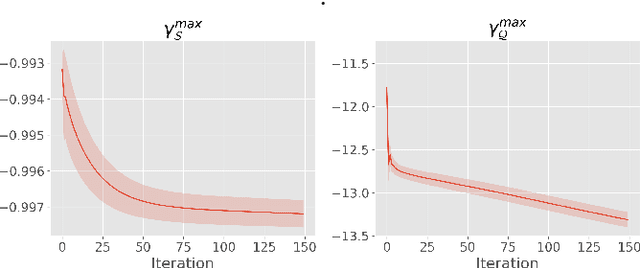
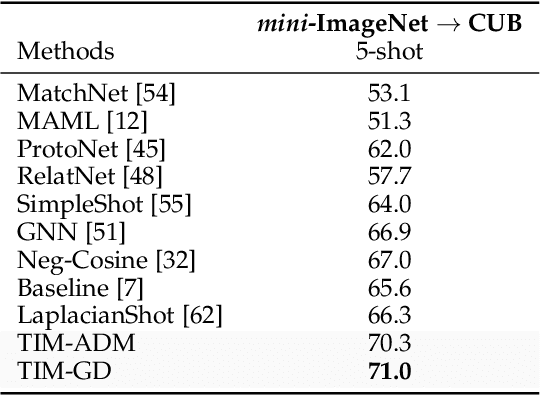
Abstract:We introduce Transductive Infomation Maximization (TIM) for few-shot learning. Our method maximizes the mutual information between the query features and their label predictions for a given few-shot task, in conjunction with a supervision loss based on the support set. We motivate our transductive loss by deriving a formal relation between the classification accuracy and mutual-information maximization. Furthermore, we propose a new alternating-direction solver, which substantially speeds up transductive inference over gradient-based optimization, while yielding competitive accuracy. We also provide a convergence analysis of our solver based on Zangwill's theory and bound-optimization arguments. TIM inference is modular: it can be used on top of any base-training feature extractor. Following standard transductive few-shot settings, our comprehensive experiments demonstrate that TIM outperforms state-of-the-art methods significantly across various datasets and networks, while used on top of a fixed feature extractor trained with simple cross-entropy on the base classes, without resorting to complex meta-learning schemes. It consistently brings between 2 % and 5 % improvement in accuracy over the best performing method, not only on all the well-established few-shot benchmarks but also on more challenging scenarios, with random tasks, domain shift and larger numbers of classes, as in the recently introduced META-DATASET. Our code is publicly available at https://github.com/mboudiaf/TIM. We also publicly release a standalone PyTorch implementation of META-DATASET, along with additional benchmarking results, at https://github.com/mboudiaf/pytorch-meta-dataset.
Transductive Few-Shot Learning: Clustering is All You Need?
Jun 16, 2021
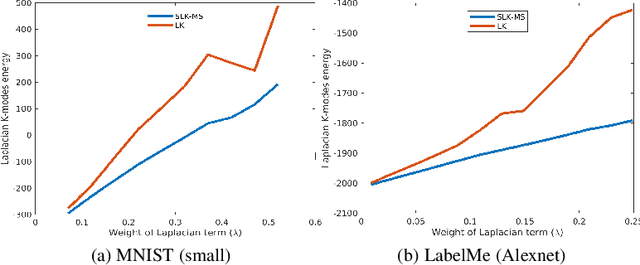
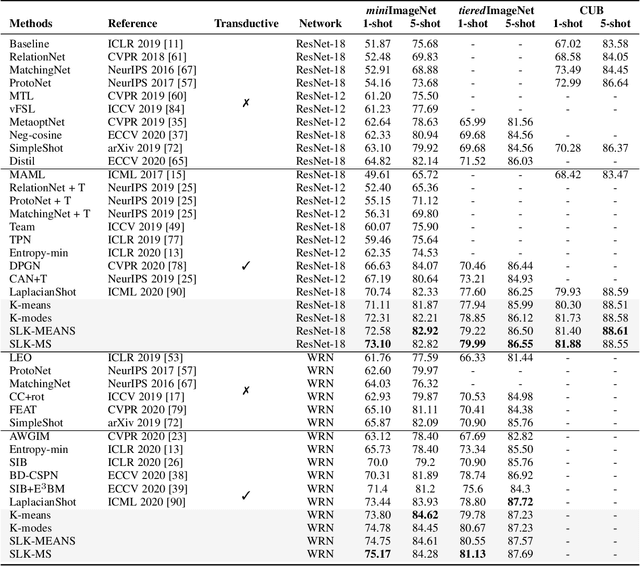
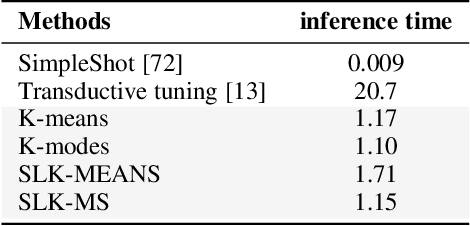
Abstract:We investigate a general formulation for clustering and transductive few-shot learning, which integrates prototype-based objectives, Laplacian regularization and supervision constraints from a few labeled data points. We propose a concave-convex relaxation of the problem, and derive a computationally efficient block-coordinate bound optimizer, with convergence guarantee. At each iteration,our optimizer computes independent (parallel) updates for each point-to-cluster assignment. Therefore, it could be trivially distributed for large-scale clustering and few-shot tasks. Furthermore, we provides a thorough convergence analysis based on point-to-set maps. Were port comprehensive clustering and few-shot learning experiments over various data sets, showing that our method yields competitive performances, in term of accuracy and optimization quality, while scaling up to large problems. Using standard training on the base classes, without resorting to complex meta-learning and episodic-training strategies, our approach outperforms state-of-the-art few-shot methods by significant margins, across various models, settings and data sets. Surprisingly, we found that even standard clustering procedures (e.g., K-means), which correspond to particular, non-regularized cases of our general model, already achieve competitive performances in comparison to the state-of-the-art in few-shot learning. These surprising results point to the limitations of the current few-shot benchmarks, and question the viability of a large body of convoluted few-shot learning techniques in the recent literature.
Orthogonal Ensemble Networks for Biomedical Image Segmentation
May 22, 2021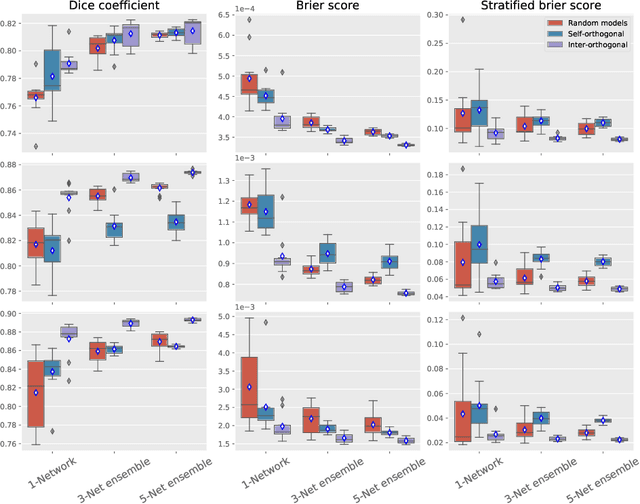
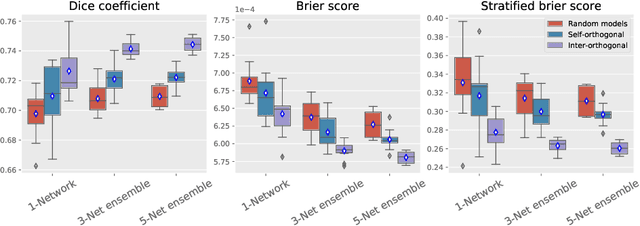
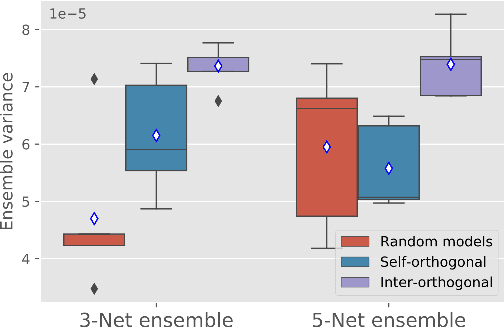
Abstract:Despite the astonishing performance of deep-learning based approaches for visual tasks such as semantic segmentation, they are known to produce miscalibrated predictions, which could be harmful for critical decision-making processes. Ensemble learning has shown to not only boost the performance of individual models but also reduce their miscalibration by averaging independent predictions. In this scenario, model diversity has become a key factor, which facilitates individual models converging to different functional solutions. In this work, we introduce Orthogonal Ensemble Networks (OEN), a novel framework to explicitly enforce model diversity by means of orthogonal constraints. The proposed method is based on the hypothesis that inducing orthogonality among the constituents of the ensemble will increase the overall model diversity. We resort to a new pairwise orthogonality constraint which can be used to regularize a sequential ensemble training process, resulting on improved predictive performance and better calibrated model outputs. We benchmark the proposed framework in two challenging brain lesion segmentation tasks --brain tumor and white matter hyper-intensity segmentation in MR images. The experimental results show that our approach produces more robust and well-calibrated ensemble models and can deal with challenging tasks in the context of biomedical image segmentation.
Self-learning for weakly supervised Gleason grading of local patterns
May 21, 2021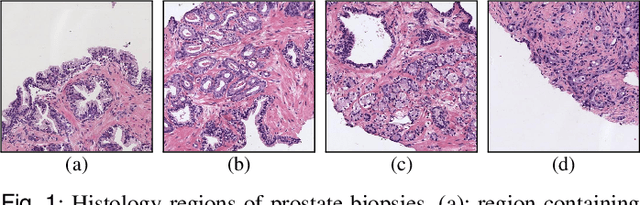
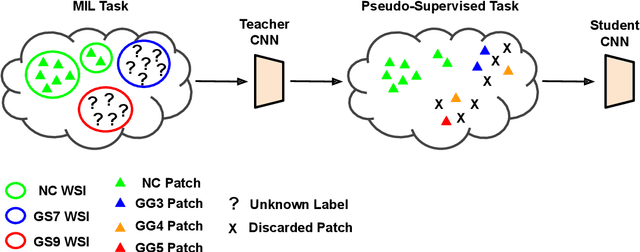
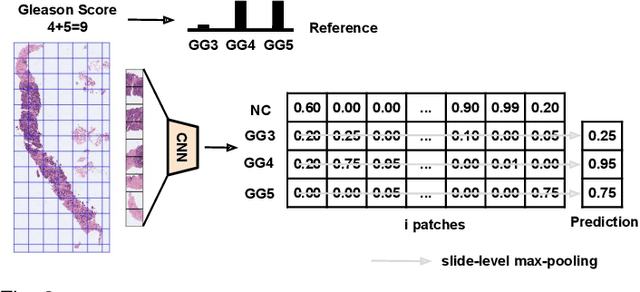
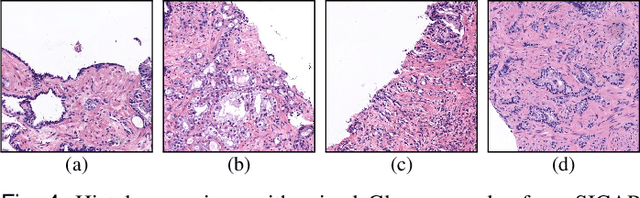
Abstract:Prostate cancer is one of the main diseases affecting men worldwide. The gold standard for diagnosis and prognosis is the Gleason grading system. In this process, pathologists manually analyze prostate histology slides under microscope, in a high time-consuming and subjective task. In the last years, computer-aided-diagnosis (CAD) systems have emerged as a promising tool that could support pathologists in the daily clinical practice. Nevertheless, these systems are usually trained using tedious and prone-to-error pixel-level annotations of Gleason grades in the tissue. To alleviate the need of manual pixel-wise labeling, just a handful of works have been presented in the literature. Motivated by this, we propose a novel weakly-supervised deep-learning model, based on self-learning CNNs, that leverages only the global Gleason score of gigapixel whole slide images during training to accurately perform both, grading of patch-level patterns and biopsy-level scoring. To evaluate the performance of the proposed method, we perform extensive experiments on three different external datasets for the patch-level Gleason grading, and on two different test sets for global Grade Group prediction. We empirically demonstrate that our approach outperforms its supervised counterpart on patch-level Gleason grading by a large margin, as well as state-of-the-art methods on global biopsy-level scoring. Particularly, the proposed model brings an average improvement on the Cohen's quadratic kappa (k) score of nearly 18% compared to full-supervision for the patch-level Gleason grading task.
Beyond pixel-wise supervision for segmentation: A few global shape descriptors might be surprisingly good!
May 03, 2021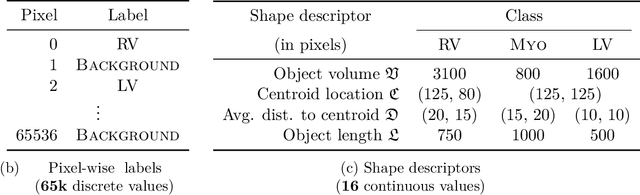

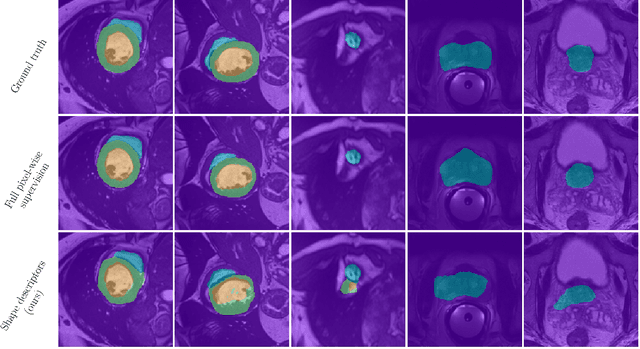
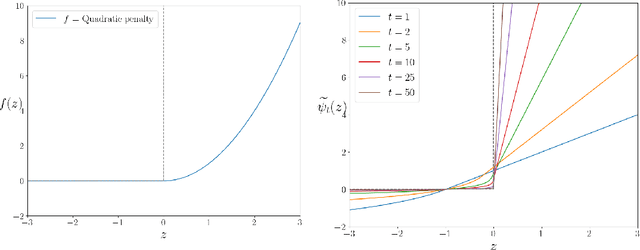
Abstract:Standard losses for training deep segmentation networks could be seen as individual classifications of pixels, instead of supervising the global shape of the predicted segmentations. While effective, they require exact knowledge of the label of each pixel in an image. This study investigates how effective global geometric shape descriptors could be, when used on their own as segmentation losses for training deep networks. Not only interesting theoretically, there exist deeper motivations to posing segmentation problems as a reconstruction of shape descriptors: Annotations to obtain approximations of low-order shape moments could be much less cumbersome than their full-mask counterparts, and anatomical priors could be readily encoded into invariant shape descriptions, which might alleviate the annotation burden. Also, and most importantly, we hypothesize that, given a task, certain shape descriptions might be invariant across image acquisition protocols/modalities and subject populations, which might open interesting research avenues for generalization in medical image segmentation. We introduce and formulate a few shape descriptors in the context of deep segmentation, and evaluate their potential as standalone losses on two different challenging tasks. Inspired by recent works in constrained optimization for deep networks, we propose a way to use those descriptors to supervise segmentation, without any pixel-level label. Very surprisingly, as little as 4 descriptors values per class can approach the performance of a segmentation mask with 65k individual discrete labels. We also found that shape descriptors can be a valid way to encode anatomical priors about the task, enabling to leverage expert knowledge without additional annotations. Our implementation is publicly available and can be easily extended to other tasks and descriptors: https://github.com/hkervadec/shape_descriptors
Incremental Multi-Target Domain Adaptation for Object Detection with Efficient Domain Transfer
Apr 19, 2021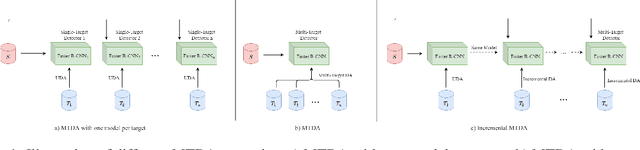
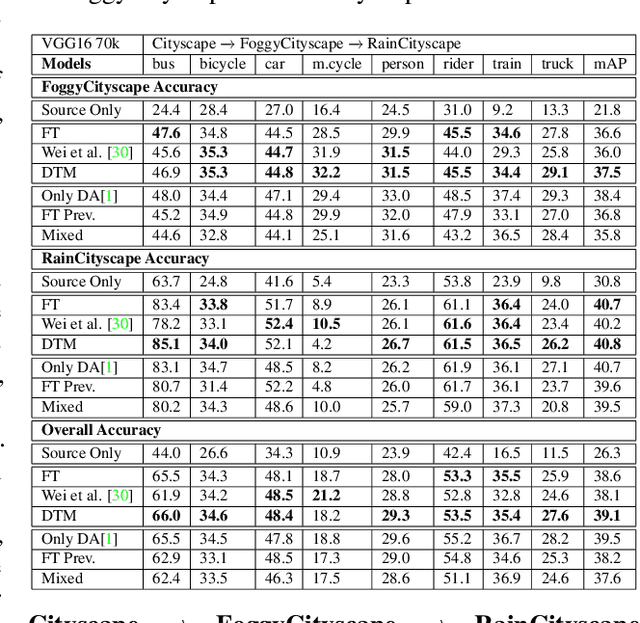
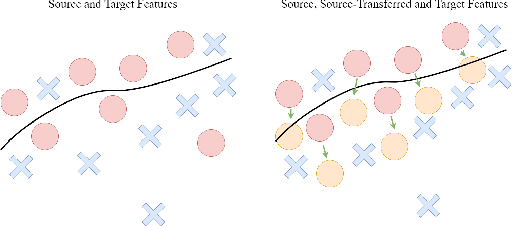
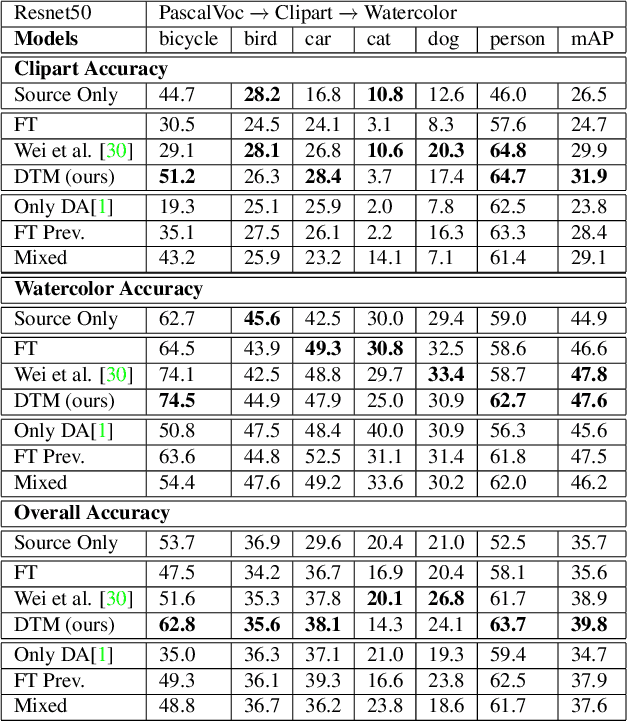
Abstract:Techniques for multi-target domain adaptation (MTDA) seek to adapt a recognition model such that it can generalize well across multiple target domains. While several successful techniques have been proposed for unsupervised single-target domain adaptation (STDA) in object detection, adapting a model to multiple target domains using unlabeled image data remains a challenging and largely unexplored problem. Key challenges include the lack of bounding box annotations for target data, knowledge corruption, and the growing resource requirements needed to train accurate deep detection models. The later requirements are augmented by the need to retraining a model with previous-learned target data when adapting to each new target domain. Currently, the only MTDA technique in literature for object detection relies on distillation with a duplicated model to avoid knowledge corruption but does not leverage the source-target feature alignment after UDA. To address these challenges, we propose a new Incremental MTDA technique for object detection that can adapt a detector to multiple target domains, one at a time, without having to retain data of previously-learned target domains. Instead of distillation, our technique efficiently transfers source images to a joint target domains' space, on the fly, thereby preserving knowledge during incremental MTDA. Using adversarial training, our Domain Transfer Module (DTM) is optimized to trick the domain classifiers into classifying source images as though transferred into the target domain, thus allowing the DTM to generate samples close to a joint distribution of target domains. Our proposed technique is validated on different MTDA detection benchmarks, and results show it improving accuracy across multiple domains, despite the considerable reduction in complexity.
The hidden label-marginal biases of segmentation losses
Apr 18, 2021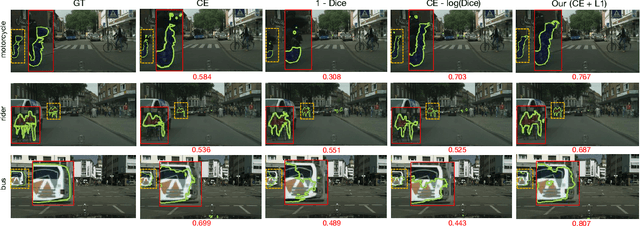
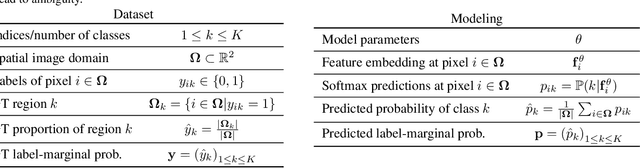
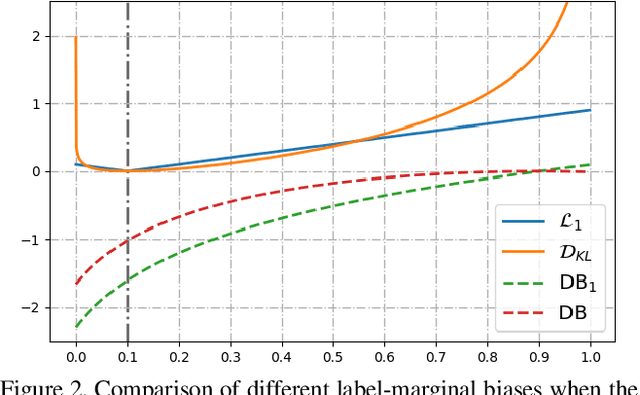
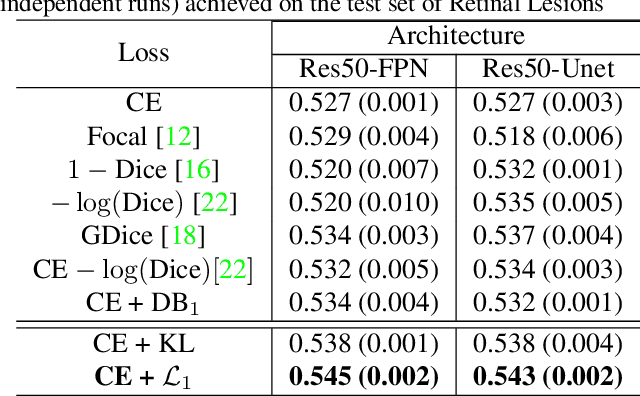
Abstract:Most segmentation losses are arguably variants of the Cross-Entropy (CE) or Dice loss. In the literature, there is no clear consensus as to which of these losses is a better choice, with varying performances for each across different benchmarks and applications. We develop a theoretical analysis that links these two types of losses, exposing their advantages and weaknesses. First, we explicitly demonstrate that CE and Dice share a much deeper connection than previously thought: CE is an upper bound on both logarithmic and linear Dice losses. Furthermore, we provide an information-theoretic analysis, which highlights hidden label-marginal biases : Dice has an intrinsic bias towards imbalanced solutions, whereas CE implicitly encourages the ground-truth region proportions. Our theoretical results explain the wide experimental evidence in the medical-imaging literature, whereby Dice losses bring improvements for imbalanced segmentation. It also explains why CE dominates natural-image problems with diverse class proportions, in which case Dice might have difficulty adapting to different label-marginal distributions. Based on our theoretical analysis, we propose a principled and simple solution, which enables to control explicitly the label-marginal bias. Our loss integrates CE with explicit ${\cal L}_1$ regularization, which encourages label marginals to match target class proportions, thereby mitigating class imbalance but without losing generality. Comprehensive experiments and ablation studies over different losses and applications validate our theoretical analysis, as well as the effectiveness of our explicit label-marginal regularizers.
Weakly supervised segmentation with cross-modality equivariant constraints
Apr 06, 2021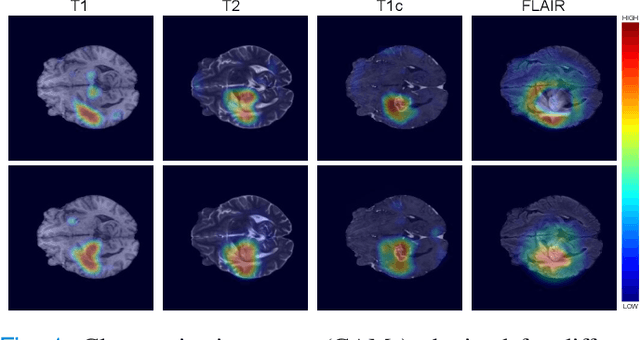
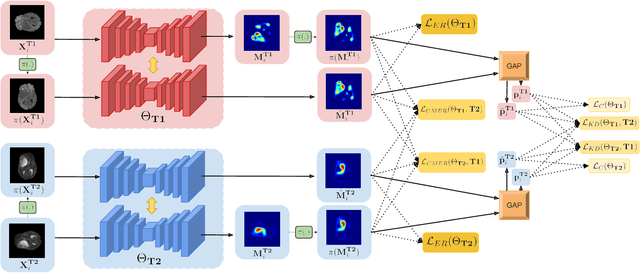
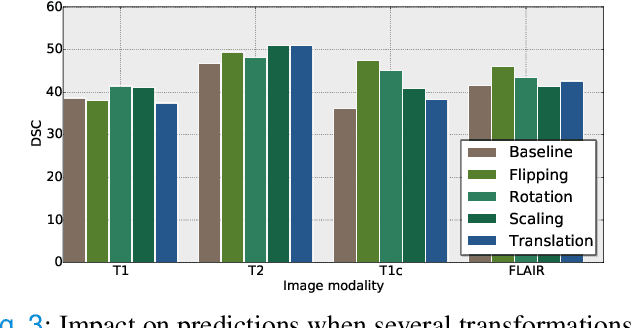
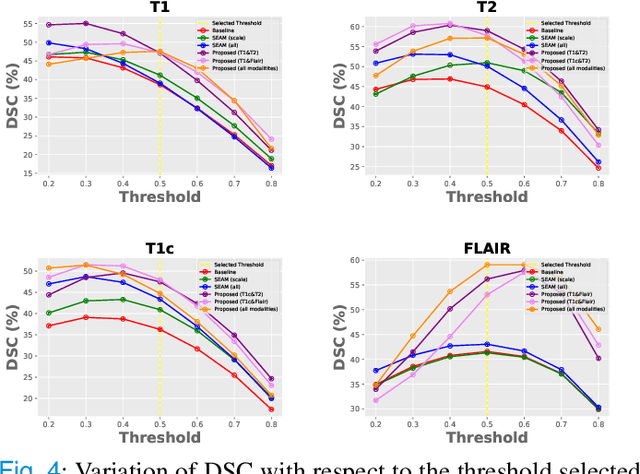
Abstract:Weakly supervised learning has emerged as an appealing alternative to alleviate the need for large labeled datasets in semantic segmentation. Most current approaches exploit class activation maps (CAMs), which can be generated from image-level annotations. Nevertheless, resulting maps have been demonstrated to be highly discriminant, failing to serve as optimal proxy pixel-level labels. We present a novel learning strategy that leverages self-supervision in a multi-modal image scenario to significantly enhance original CAMs. In particular, the proposed method is based on two observations. First, the learning of fully-supervised segmentation networks implicitly imposes equivariance by means of data augmentation, whereas this implicit constraint disappears on CAMs generated with image tags. And second, the commonalities between image modalities can be employed as an efficient self-supervisory signal, correcting the inconsistency shown by CAMs obtained across multiple modalities. To effectively train our model, we integrate a novel loss function that includes a within-modality and a cross-modality equivariant term to explicitly impose these constraints during training. In addition, we add a KL-divergence on the class prediction distributions to facilitate the information exchange between modalities, which, combined with the equivariant regularizers further improves the performance of our model. Exhaustive experiments on the popular multi-modal BRATS dataset demonstrate that our approach outperforms relevant recent literature under the same learning conditions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge