Mathilde Bateson
Test-Time Adaptation with Shape Moments for Image Segmentation
May 16, 2022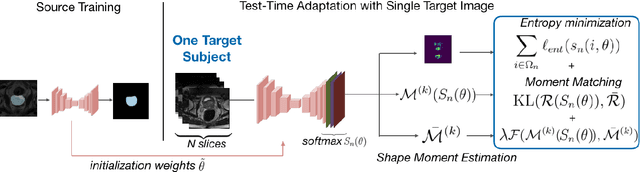
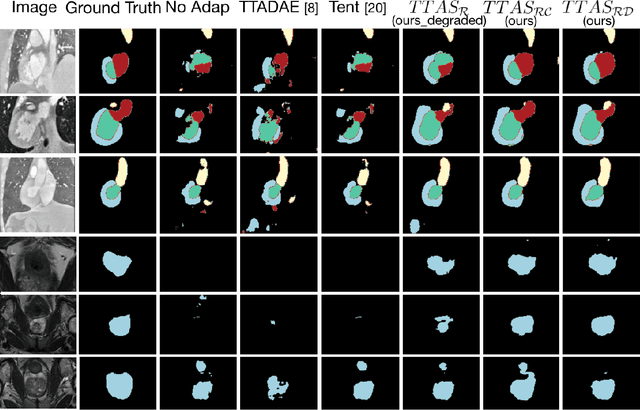
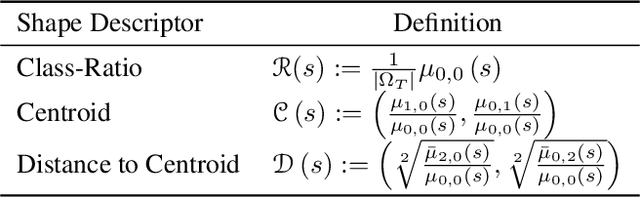
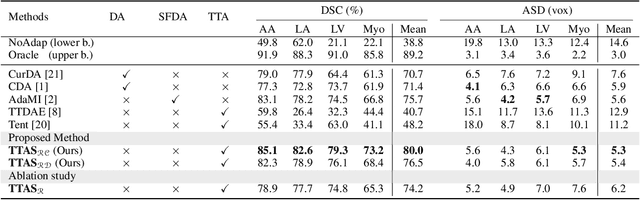

Abstract:Supervised learning is well-known to fail at generalization under distribution shifts. In typical clinical settings, the source data is inaccessible and the target distribution is represented with a handful of samples: adaptation can only happen at test time on a few or even a single subject(s). We investigate test-time single-subject adaptation for segmentation, and propose a Shape-guided Entropy Minimization objective for tackling this task. During inference for a single testing subject, our loss is minimized with respect to the batch normalization's scale and bias parameters. We show the potential of integrating various shape priors to guide adaptation to plausible solutions, and validate our method in two challenging scenarios: MRI-to-CT adaptation of cardiac segmentation and cross-site adaptation of prostate segmentation. Our approach exhibits substantially better performances than the existing test-time adaptation methods. Even more surprisingly, it fares better than state-of-the-art domain adaptation methods, although it forgoes training on additional target data during adaptation. Our results question the usefulness of training on target data in segmentation adaptation, and points to the substantial effect of shape priors on test-time inference. Our framework can be readily used for integrating various priors and for adapting any segmentation network, and our code is available.
Source-Free Domain Adaptation for Image Segmentation
Aug 06, 2021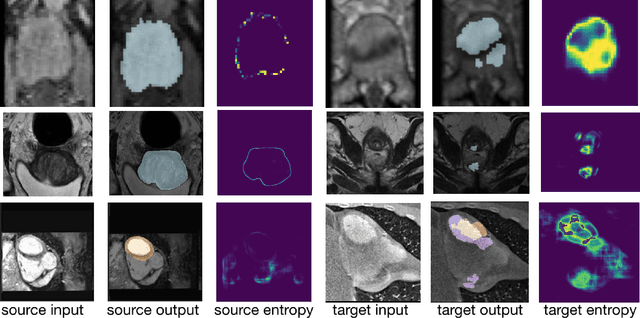
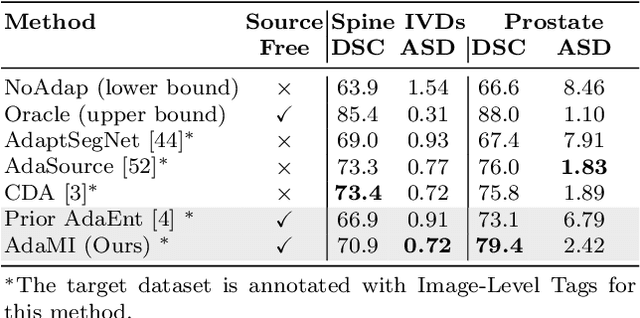
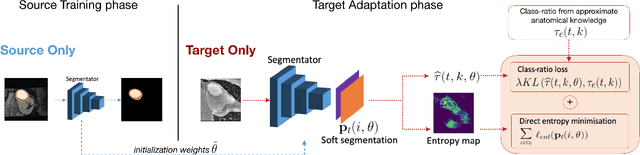
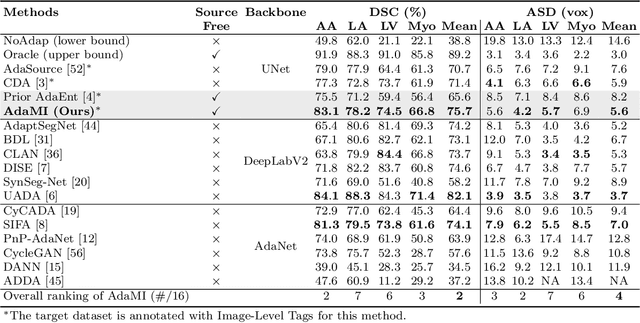
Abstract:Domain adaptation (DA) has drawn high interest for its capacity to adapt a model trained on labeled source data to perform well on unlabeled or weakly labeled target data from a different domain. Most common DA techniques require concurrent access to the input images of both the source and target domains. However, in practice, privacy concerns often impede the availability of source images in the adaptation phase. This is a very frequent DA scenario in medical imaging, where, for instance, the source and target images could come from different clinical sites. We introduce a source-free domain adaptation for image segmentation. Our formulation is based on minimizing a label-free entropy loss defined over target-domain data, which we further guide with a domain-invariant prior on the segmentation regions. Many priors can be derived from anatomical information. Here, a class ratio prior is estimated from anatomical knowledge and integrated in the form of a Kullback Leibler (KL) divergence in our overall loss function. Furthermore, we motivate our overall loss with an interesting link to maximizing the mutual information between the target images and their label predictions. We show the effectiveness of our prior aware entropy minimization in a variety of domain-adaptation scenarios, with different modalities and applications, including spine, prostate, and cardiac segmentation. Our method yields comparable results to several state of the art adaptation techniques, despite having access to much less information, as the source images are entirely absent in our adaptation phase. Our straightforward adaptation strategy uses only one network, contrary to popular adversarial techniques, which are not applicable to a source-free DA setting. Our framework can be readily used in a breadth of segmentation problems, and our code is publicly available: https://github.com/mathilde-b/SFDA
Source-Relaxed Domain Adaptation for Image Segmentation
May 07, 2020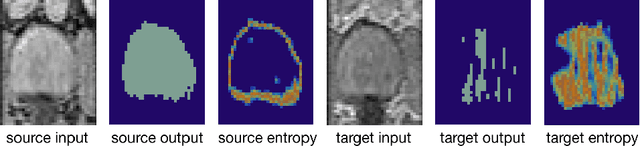
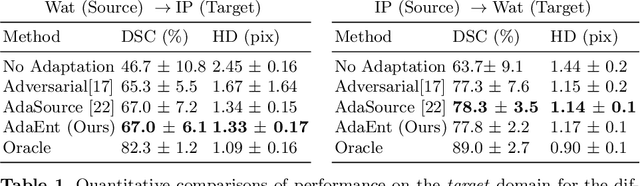

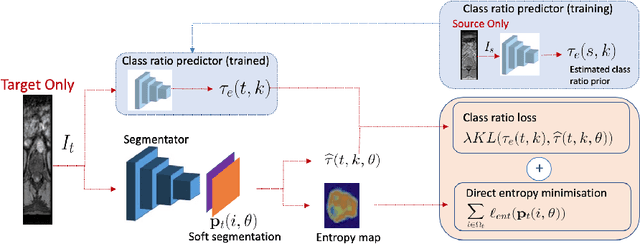
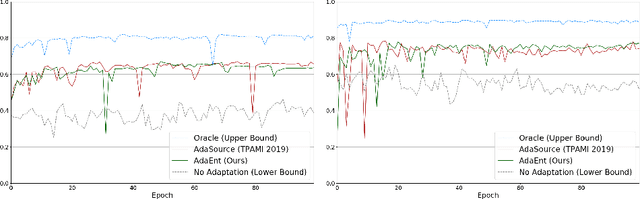
Abstract:Domain adaptation (DA) has drawn high interests for its capacity to adapt a model trained on labeled source data to perform well on unlabeled or weakly labeled target data from a different domain. Most common DA techniques require the concurrent access to the input images of both the source and target domains. However, in practice, it is common that the source images are not available in the adaptation phase. This is a very frequent DA scenario in medical imaging, for instance, when the source and target images come from different clinical sites. We propose a novel formulation for adapting segmentation networks, which relaxes such a constraint. Our formulation is based on minimizing a label-free entropy loss defined over target-domain data, which we further guide with a domain invariant prior on the segmentation regions. Many priors can be used, derived from anatomical information. Here, a class-ratio prior is learned via an auxiliary network and integrated in the form of a Kullback-Leibler (KL) divergence in our overall loss function. We show the effectiveness of our prior-aware entropy minimization in adapting spine segmentation across different MRI modalities. Our method yields comparable results to several state-of-the-art adaptation techniques, even though is has access to less information, the source images being absent in the adaptation phase. Our straight-forward adaptation strategy only uses one network, contrary to popular adversarial techniques, which cannot perform without the presence of the source images. Our framework can be readily used with various priors and segmentation problems.
Constrained domain adaptation for segmentation
Aug 08, 2019
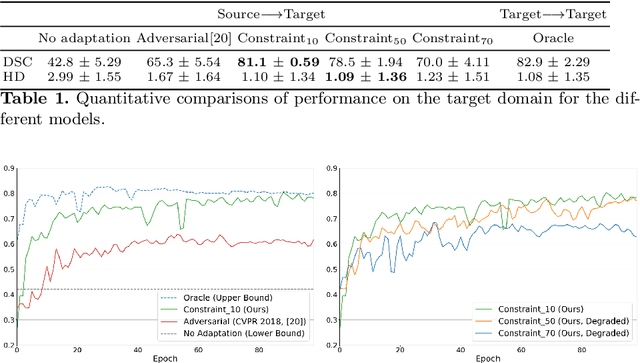
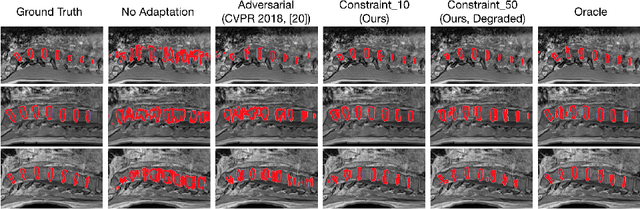
Abstract:We propose to adapt segmentation networks with a constrained formulation, which embeds domain-invariant prior knowledge about the segmentation regions. Such knowledge may take the form of simple anatomical information, e.g., structure size or shape, estimated from source samples or known a priori. Our method imposes domain-invariant inequality constraints on the network outputs of unlabeled target samples. It implicitly matches prediction statistics between target and source domains with permitted uncertainty of prior knowledge. We address our constrained problem with a differentiable penalty, fully suited for standard stochastic gradient descent approaches, removing the need for computationally expensive Lagrangian optimization with dual projections. Unlike current two-step adversarial training, our formulation is based on a single loss in a single network, which simplifies adaptation by avoiding extra adversarial steps, while improving convergence and quality of training. The comparison of our approach with state-of-the-art adversarial methods reveals substantially better performance on the challenging task of adapting spine segmentation across different MRI modalities. Our results also show a robustness to imprecision of size priors, approaching the accuracy of a fully supervised model trained directly in a target domain.Our method can be readily used for various constraints and segmentation problems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge