Jan Egger
Medical Deep Learning -- A systematic Meta-Review
Oct 28, 2020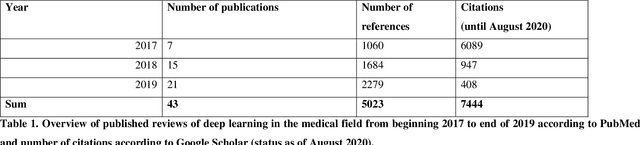
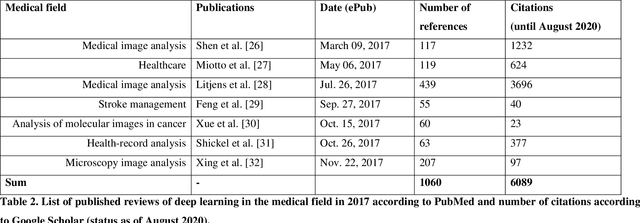
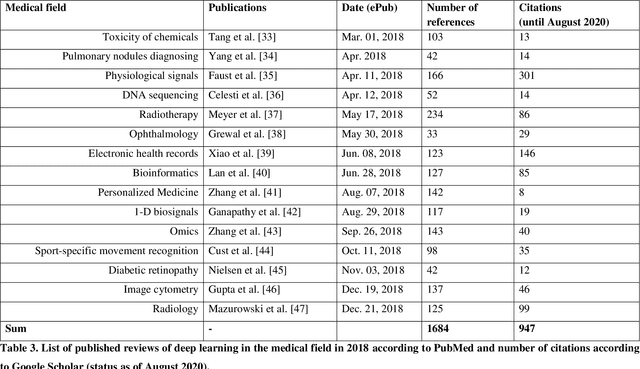

Abstract:Deep learning had a remarkable impact in different scientific disciplines during the last years. This was demonstrated in numerous tasks, where deep learning algorithms were able to outperform the state-of-art methods, also in image processing and analysis. Moreover, deep learning delivers good results in tasks like autonomous driving, which could not have been performed automatically before. There are even applications where deep learning outperformed humans, like object recognition or games. Another field in which this development is showing a huge potential is the medical domain. With the collection of large quantities of patient records and data, and a trend towards personalized treatments, there is a great need for an automatic and reliable processing and analysis of this information. Patient data is not only collected in clinical centres, like hospitals, but it relates also to data coming from general practitioners, healthcare smartphone apps or online websites, just to name a few. This trend resulted in new, massive research efforts during the last years. In Q2/2020, the search engine PubMed returns already over 11.000 results for the search term $'$deep learning$'$, and around 90% of these publications are from the last three years. Hence, a complete overview of the field of $'$medical deep learning$'$ is almost impossible to obtain and getting a full overview of medical sub-fields gets increasingly more difficult. Nevertheless, several review and survey articles about medical deep learning have been presented within the last years. They focused, in general, on specific medical scenarios, like the analysis of medical images containing specific pathologies. With these surveys as foundation, the aim of this contribution is to provide a very first high-level, systematic meta-review of medical deep learning surveys.
A Baseline Approach for AutoImplant: the MICCAI 2020 Cranial Implant Design Challenge
Jun 24, 2020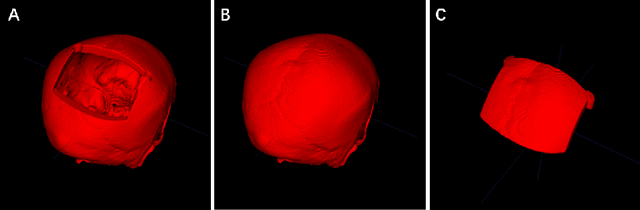
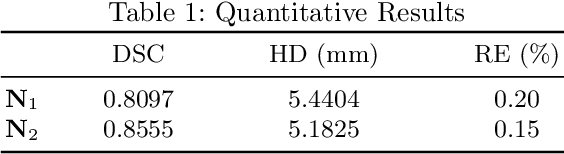
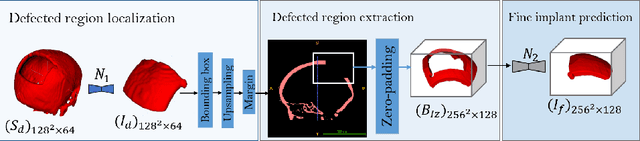

Abstract:In this study, we present a baseline approach for AutoImplant (https://autoimplant.grand-challenge.org/) - the cranial implant design challenge, which, as suggested by the organizers, can be formulated as a volumetric shape learning task. In this task, the defective skull, the complete skull and the cranial implant are represented as binary voxel grids. To accomplish this task, the implant can be either reconstructed directly from the defective skull or obtained by taking the difference between a defective skull and a complete skull. In the latter case, a complete skull has to be reconstructed given a defective skull, which defines a volumetric shape completion problem. Our baseline approach for this task is based on the former formulation, i.e., a deep neural network is trained to predict the implants directly from the defective skulls. The approach generates high-quality implants in two steps: First, an encoder-decoder network learns a coarse representation of the implant from down-sampled, defective skulls; The coarse implant is only used to generate the bounding box of the defected region in the original high-resolution skull. Second, another encoder-decoder network is trained to generate a fine implant from the bounded area. On the test set, the proposed approach achieves an average dice similarity score (DSC) of 0.8555 and Hausdorff distance (HD) of 5.1825 mm. The code is publicly available at https://github.com/Jianningli/autoimplant.
An Online Platform for Automatic Skull Defect Restoration and Cranial Implant Design
Jun 01, 2020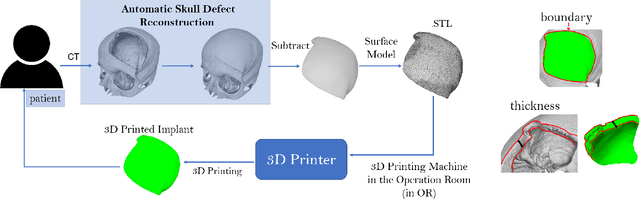
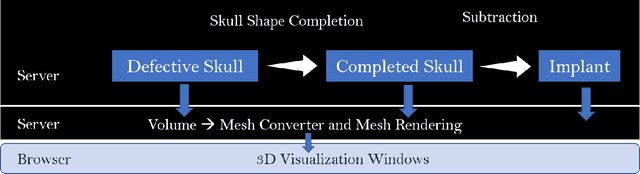
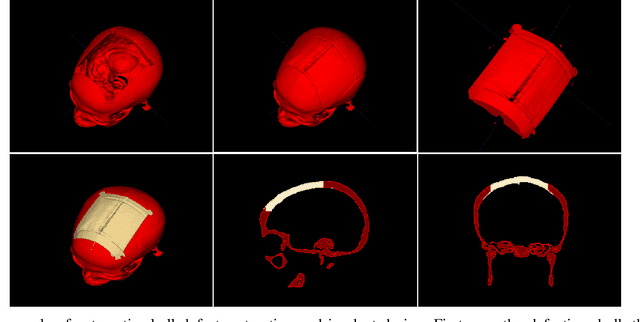
Abstract:We introduce a fully automatic system for cranial implant design, a common task in cranioplasty operations. The system is currently integrated in Studierfenster (http://studierfenster.tugraz.at/), an online, cloud-based medical image processing platform for medical imaging applications. Enhanced by deep learning algorithms, the system automatically restores the missing part of a skull (i.e., skull shape completion) and generates the desired implant by subtracting the defective skull from the completed skull. The generated implant can be downloaded in the STereoLithography (.stl) format directly via the browser interface of the system. The implant model can then be sent to a 3D printer for in loco implant manufacturing. Furthermore, thanks to the standard format, the user can thereafter load the model into another application for post-processing whenever necessary. Such an automatic cranial implant design system can be integrated into the clinical practice to improve the current routine for surgeries related to skull defect repair (e.g., cranioplasty). Our system, although currently intended for educational and research use only, can be seen as an application of additive manufacturing for fast, patient-specific implant design.
Design and Development of a Web-based Tool for Inpainting of Dissected Aortae in Angiography Images
May 06, 2020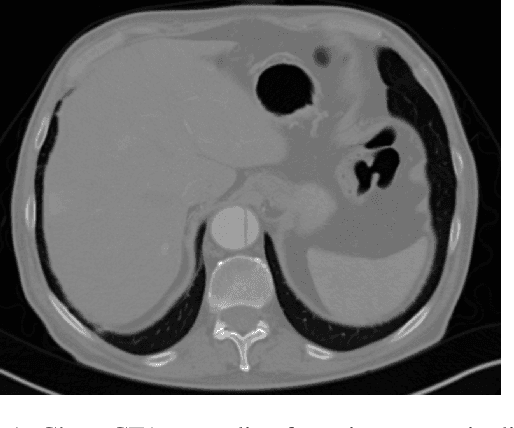
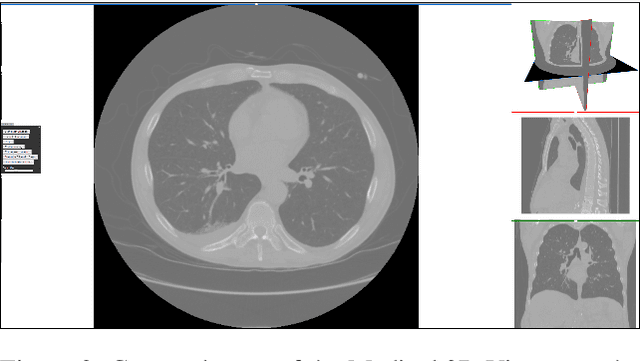

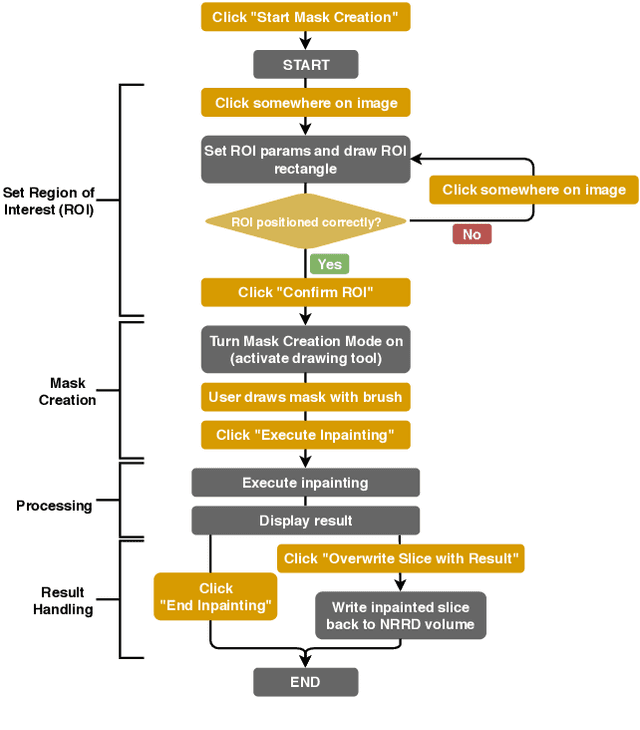
Abstract:Medical imaging is an important tool for the diagnosis and the evaluation of an aortic dissection (AD); a serious condition of the aorta, which could lead to a life-threatening aortic rupture. AD patients need life-long medical monitoring of the aortic enlargement and of the disease progression, subsequent to the diagnosis of the aortic dissection. Since there is a lack of 'healthy-dissected' image pairs from medical studies, the application of inpainting techniques offers an alternative source for generating them by doing a virtual regression from dissected aortae to healthy aortae; an indirect way to study the origin of the disease. The proposed inpainting tool combines a neural network, which was trained on the task of inpainting aortic dissections, with an easy-to-use user interface. To achieve this goal, the inpainting tool has been integrated within the 3D medical image viewer of StudierFenster (www.studierfenster.at). By designing the tool as a web application, we simplify the usage of the neural network and reduce the initial learning curve.
* 9 figures, 14 references
Client/Server Based Online Environment for Manual Segmentation of Medical Images
Apr 18, 2019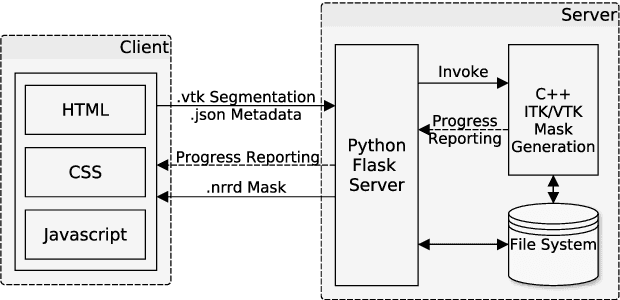
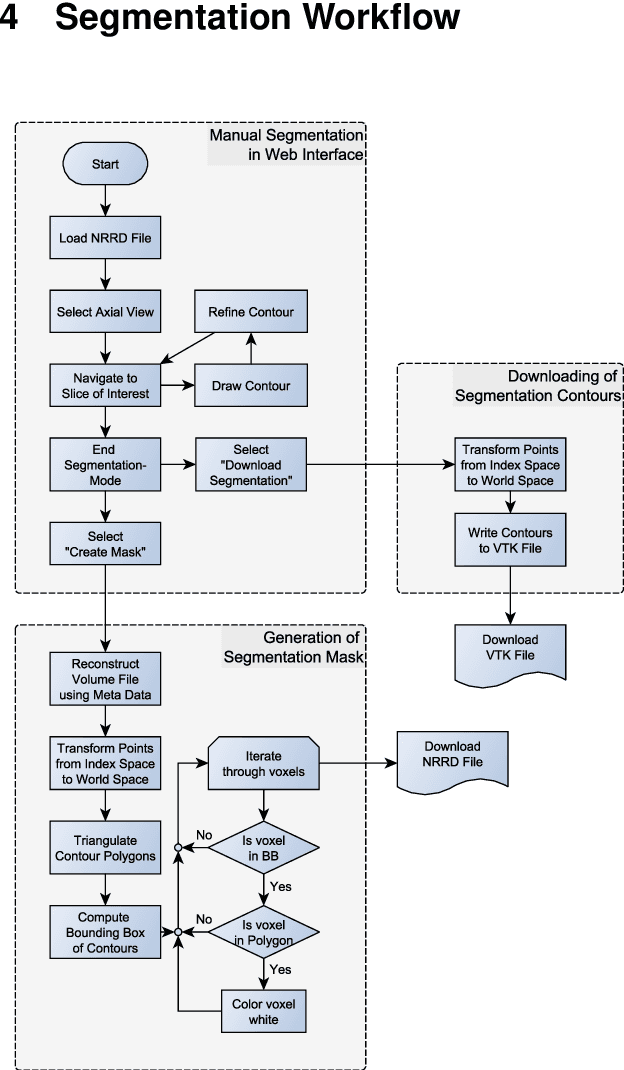
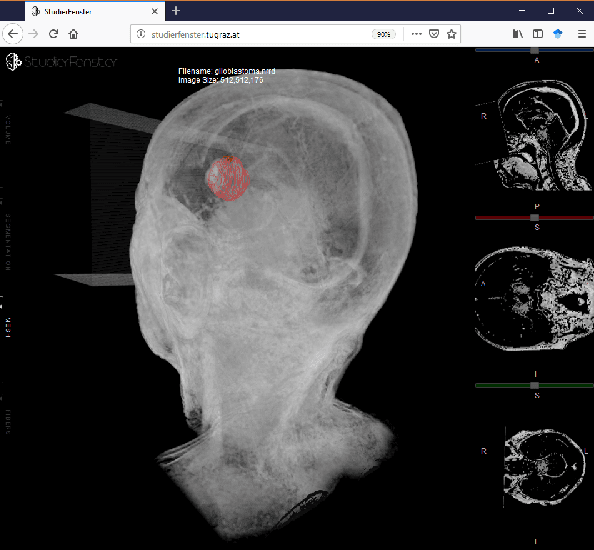
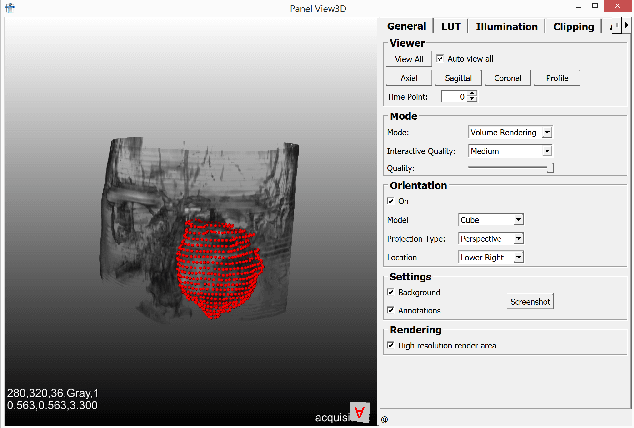
Abstract:Segmentation is a key step in analyzing and processing medical images. Due to the low fault tolerance in medical imaging, manual segmentation remains the de facto standard in this domain. Besides, efforts to automate the segmentation process often rely on large amounts of manually labeled data. While existing software supporting manual segmentation is rich in features and delivers accurate results, the necessary time to set it up and get comfortable using it can pose a hurdle for the collection of large datasets. This work introduces a client/server based online environment, referred to as Studierfenster (studierfenster.at), that can be used to perform manual segmentations directly in a web browser. The aim of providing this functionality in the form of a web application is to ease the collection of ground truth segmentation datasets. Providing a tool that is quickly accessible and usable on a broad range of devices, offers the potential to accelerate this process. The manual segmentation workflow of Studierfenster consists of dragging and dropping the input file into the browser window and slice-by-slice outlining the object under consideration. The final segmentation can then be exported as a file storing its contours and as a binary segmentation mask. In order to evaluate the usability of Studierfenster, a user study was performed. The user study resulted in a mean of 6.3 out of 7.0 possible points given by users, when asked about their overall impression of the tool. The evaluation also provides insights into the results achievable with the tool in practice, by presenting two ground truth segmentations performed by physicians.
* 8 pages
Exploit fully automatic low-level segmented PET data for training high-level deep learning algorithms for the corresponding CT data
Mar 07, 2019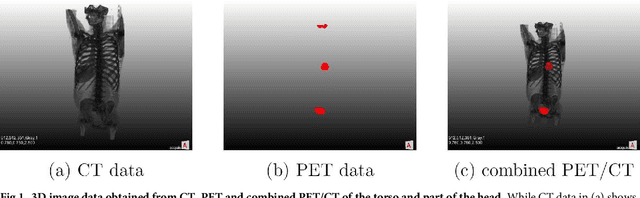
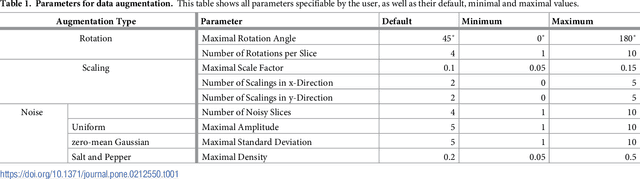
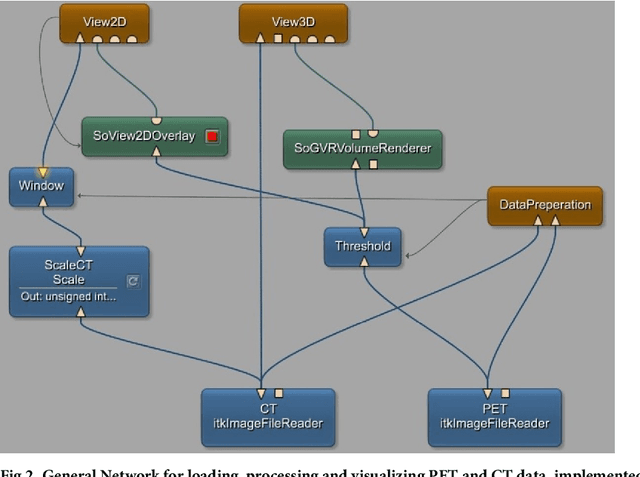
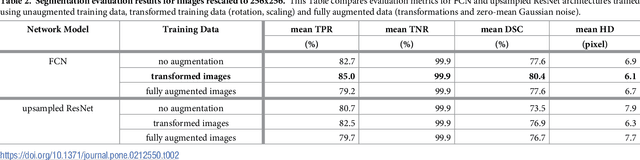
Abstract:We present an approach for fully automatic urinary bladder segmentation in CT images with artificial neural networks in this study. Automatic medical image analysis has become an invaluable tool in the different treatment stages of diseases. Especially medical image segmentation plays a vital role, since segmentation is often the initial step in an image analysis pipeline. Since deep neural networks have made a large impact on the field of image processing in the past years, we use two different deep learning architectures to segment the urinary bladder. Both of these architectures are based on pre-trained classification networks that are adapted to perform semantic segmentation. Since deep neural networks require a large amount of training data, specifically images and corresponding ground truth labels, we furthermore propose a method to generate such a suitable training data set from Positron Emission Tomography/Computed Tomography image data. This is done by applying thresholding to the Positron Emission Tomography data for obtaining a ground truth and by utilizing data augmentation to enlarge the dataset. In this study, we discuss the influence of data augmentation on the segmentation results, and compare and evaluate the proposed architectures in terms of qualitative and quantitative segmentation performance. The results presented in this study allow concluding that deep neural networks can be considered a promising approach to segment the urinary bladder in CT images.
* 20 pages
Computed tomography data collection of the complete human mandible and valid clinical ground truth models
Feb 14, 2019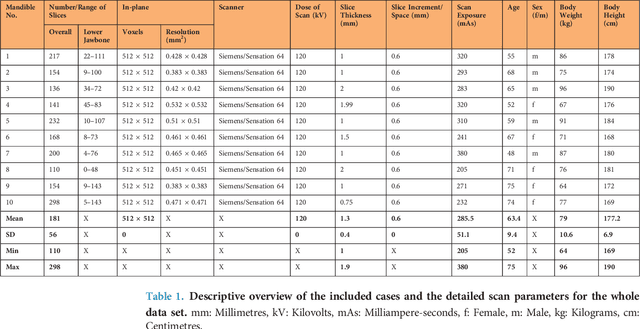
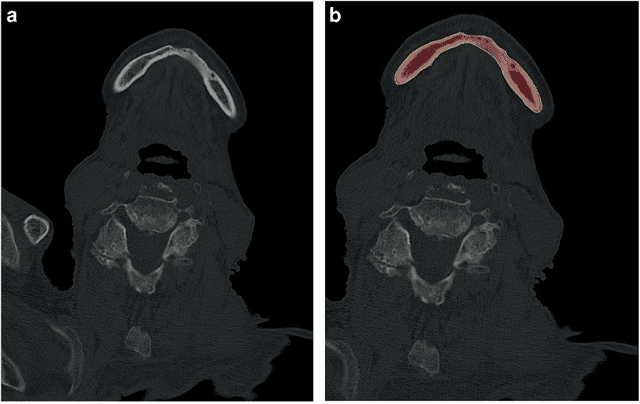
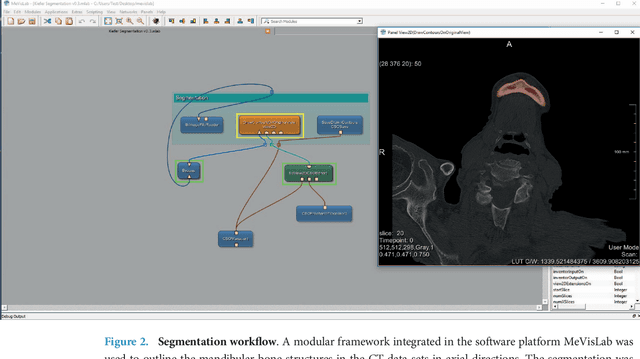
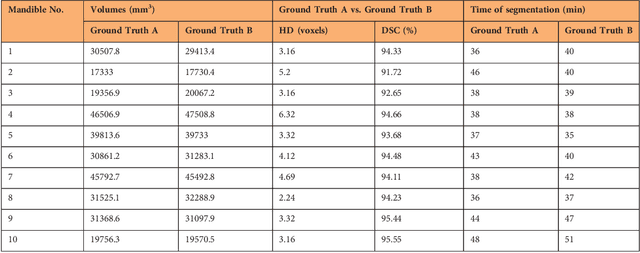
Abstract:Image-based algorithmic software segmentation is an increasingly important topic in many medical fields. Algorithmic segmentation is used for medical three-dimensional visualization, diagnosis or treatment support, especially in complex medical cases. However, accessible medical databases are limited, and valid medical ground truth databases for the evaluation of algorithms are rare and usually comprise only a few images. Inaccuracy or invalidity of medical ground truth data and image-based artefacts also limit the creation of such databases, which is especially relevant for CT data sets of the maxillomandibular complex. This contribution provides a unique and accessible data set of the complete mandible, including 20 valid ground truth segmentation models originating from 10 CT scans from clinical practice without artefacts or faulty slices. From each CT scan, two 3D ground truth models were created by clinical experts through independent manual slice-by-slice segmentation, and the models were statistically compared to prove their validity. These data could be used to conduct serial image studies of the human mandible, evaluating segmentation algorithms and developing adequate image tools.
* 14 pages, 8 Figures, 4 Tables, 55 References
Clinical evaluation of semi-automatic opensource algorithmic software segmentation of the mandibular bone: Practical feasibility and assessment of a new course of action
May 11, 2018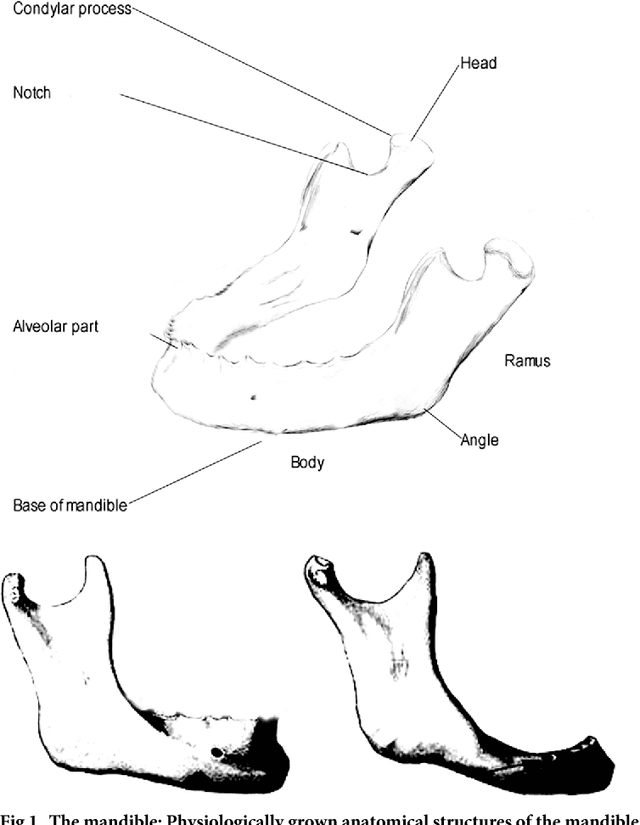
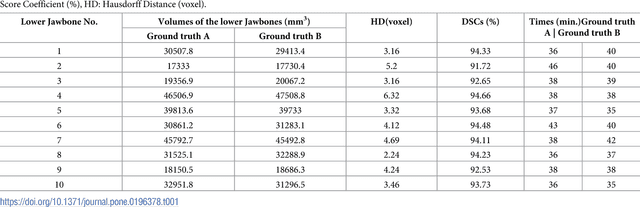
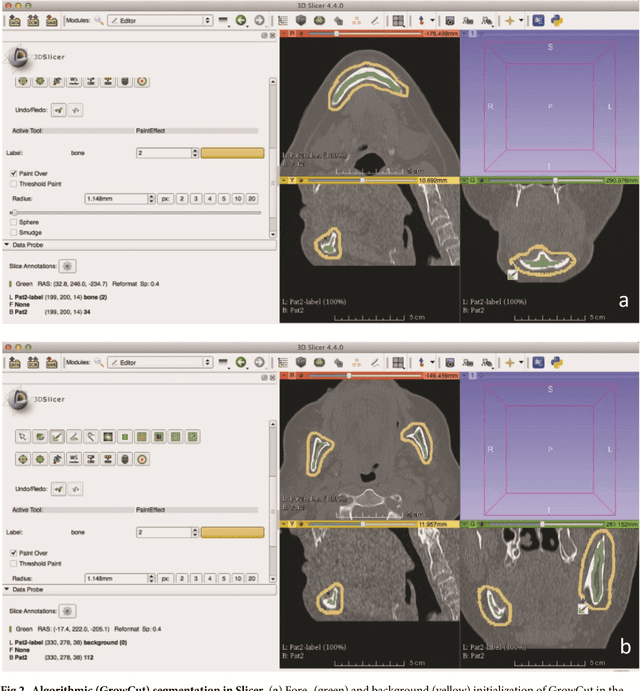
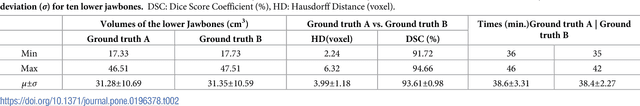
Abstract:Computer assisted technologies based on algorithmic software segmentation are an increasing topic of interest in complex surgical cases. However - due to functional instability, time consuming software processes, personnel resources or licensed-based financial costs many segmentation processes are often outsourced from clinical centers to third parties and the industry. Therefore, the aim of this trial was to assess the practical feasibility of an easy available, functional stable and licensed-free segmentation approach to be used in the clinical practice. In this retrospective, randomized, controlled trail the accuracy and accordance of the open-source based segmentation algorithm GrowCut (GC) was assessed through the comparison to the manually generated ground truth of the same anatomy using 10 CT lower jaw data-sets from the clinical routine. Assessment parameters were the segmentation time, the volume, the voxel number, the Dice Score (DSC) and the Hausdorff distance (HD). Overall segmentation times were about one minute. Mean DSC values of over 85% and HD below 33.5 voxel could be achieved. Statistical differences between the assessment parameters were not significant (p<0.05) and correlation coefficients were close to the value one (r > 0.94). Complete functional stable and time saving segmentations with high accuracy and high positive correlation could be performed by the presented interactive open-source based approach. In the cranio-maxillofacial complex the used method could represent an algorithmic alternative for image-based segmentation in the clinical practice for e.g. surgical treatment planning or visualization of postoperative results and offers several advantages. Systematic comparisons to other segmentation approaches or with a greater data amount are areas of future works.
* 26 pages
In-depth Assessment of an Interactive Graph-based Approach for the Segmentation for Pancreatic Metastasis in Ultrasound Acquisitions of the Liver with two Specialists in Internal Medicine
Mar 12, 2018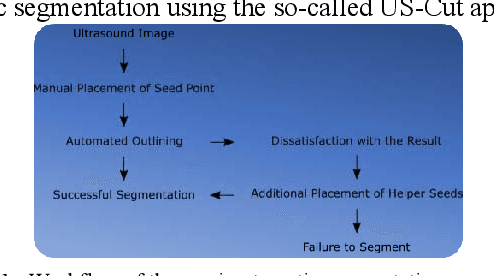
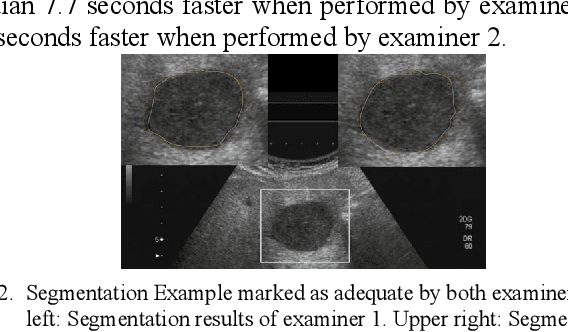
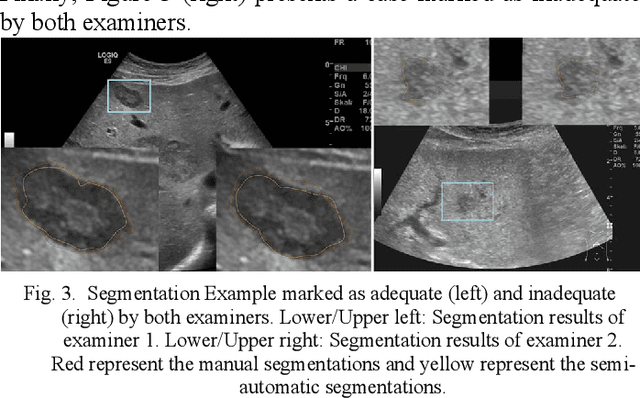
Abstract:The manual outlining of hepatic metastasis in (US) ultrasound acquisitions from patients suffering from pancreatic cancer is common practice. However, such pure manual measurements are often very time consuming, and the results repeatedly differ between the raters. In this contribution, we study the in-depth assessment of an interactive graph-based approach for the segmentation for pancreatic metastasis in US images of the liver with two specialists in Internal Medicine. Thereby, evaluating the approach with over one hundred different acquisitions of metastases. The two physicians or the algorithm had never assessed the acquisitions before the evaluation. In summary, the physicians first performed a pure manual outlining followed by an algorithmic segmentation over one month later. As a result, the experts satisfied in up to ninety percent of algorithmic segmentation results. Furthermore, the algorithmic segmentation was much faster than manual outlining and achieved a median Dice Similarity Coefficient (DSC) of over eighty percent. Ultimately, the algorithm enables a fast and accurate segmentation of liver metastasis in clinical US images, which can support the manual outlining in daily practice.
Vertebral body segmentation with GrowCut: Initial experience, workflow and practical application
Nov 13, 2017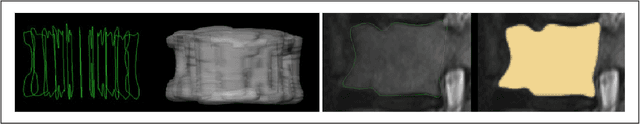
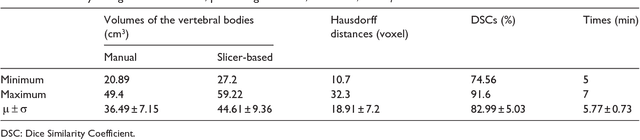
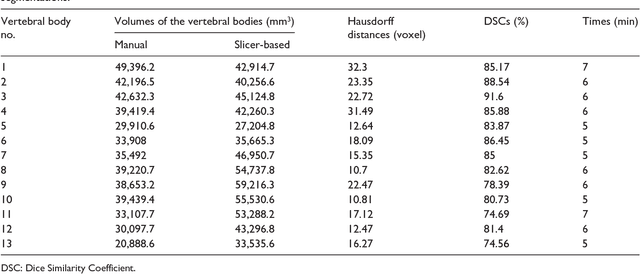
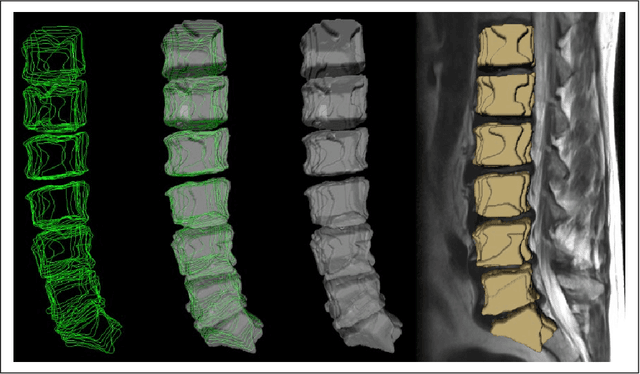

Abstract:In this contribution, we used the GrowCut segmentation algorithm publicly available in three-dimensional Slicer for three-dimensional segmentation of vertebral bodies. To the best of our knowledge, this is the first time that the GrowCut method has been studied for the usage of vertebral body segmentation. In brief, we found that the GrowCut segmentation times were consistently less than the manual segmentation times. Hence, GrowCut provides an alternative to a manual slice-by-slice segmentation process.
* 10 pages
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge