Jacob Wynne
Deep Learning Based Apparent Diffusion Coefficient Map Generation1 from Multi-parametric MR Images for Patients with Diffuse Gliomas
Jul 02, 2024



Abstract:Purpose: Apparent diffusion coefficient (ADC) maps derived from diffusion weighted (DWI) MRI provides functional measurements about the water molecules in tissues. However, DWI is time consuming and very susceptible to image artifacts, leading to inaccurate ADC measurements. This study aims to develop a deep learning framework to synthesize ADC maps from multi-parametric MR images. Methods: We proposed the multiparametric residual vision transformer model (MPR-ViT) that leverages the long-range context of ViT layers along with the precision of convolutional operators. Residual blocks throughout the network significantly increasing the representational power of the model. The MPR-ViT model was applied to T1w and T2- fluid attenuated inversion recovery images of 501 glioma cases from a publicly available dataset including preprocessed ADC maps. Selected patients were divided into training (N=400), validation (N=50) and test (N=51) sets, respectively. Using the preprocessed ADC maps as ground truth, model performance was evaluated and compared against the Vision Convolutional Transformer (VCT) and residual vision transformer (ResViT) models. Results: The results are as follows using T1w + T2-FLAIR MRI as inputs: MPR-ViT - PSNR: 31.0 +/- 2.1, MSE: 0.009 +/- 0.0005, SSIM: 0.950 +/- 0.015. In addition, ablation studies showed the relative impact on performance of each input sequence. Both qualitative and quantitative results indicate that the proposed MR- ViT model performs favorably against the ground truth data. Conclusion: We show that high-quality ADC maps can be synthesized from structural MRI using a MPR- VCT model. Our predicted images show better conformality to the ground truth volume than ResViT and VCT predictions. These high-quality synthetic ADC maps would be particularly useful for disease diagnosis and intervention, especially when ADC maps have artifacts or are unavailable.
Synthetic CT Generation from MRI using 3D Transformer-based Denoising Diffusion Model
May 31, 2023



Abstract:Magnetic resonance imaging (MRI)-based synthetic computed tomography (sCT) simplifies radiation therapy treatment planning by eliminating the need for CT simulation and error-prone image registration, ultimately reducing patient radiation dose and setup uncertainty. We propose an MRI-to-CT transformer-based denoising diffusion probabilistic model (MC-DDPM) to transform MRI into high-quality sCT to facilitate radiation treatment planning. MC-DDPM implements diffusion processes with a shifted-window transformer network to generate sCT from MRI. The proposed model consists of two processes: a forward process which adds Gaussian noise to real CT scans, and a reverse process in which a shifted-window transformer V-net (Swin-Vnet) denoises the noisy CT scans conditioned on the MRI from the same patient to produce noise-free CT scans. With an optimally trained Swin-Vnet, the reverse diffusion process was used to generate sCT scans matching MRI anatomy. We evaluated the proposed method by generating sCT from MRI on a brain dataset and a prostate dataset. Qualitative evaluation was performed using the mean absolute error (MAE) of Hounsfield unit (HU), peak signal to noise ratio (PSNR), multi-scale Structure Similarity index (MS-SSIM) and normalized cross correlation (NCC) indexes between ground truth CTs and sCTs. MC-DDPM generated brain sCTs with state-of-the-art quantitative results with MAE 43.317 HU, PSNR 27.046 dB, SSIM 0.965, and NCC 0.983. For the prostate dataset, MC-DDPM achieved MAE 59.953 HU, PSNR 26.920 dB, SSIM 0.849, and NCC 0.948. In conclusion, we have developed and validated a novel approach for generating CT images from routine MRIs using a transformer-based DDPM. This model effectively captures the complex relationship between CT and MRI images, allowing for robust and high-quality synthetic CT (sCT) images to be generated in minutes.
Cross-Shaped Windows Transformer with Self-supervised Pretraining for Clinically Significant Prostate Cancer Detection in Bi-parametric MRI
Apr 30, 2023



Abstract:Multiparametric magnetic resonance imaging (mpMRI) has demonstrated promising results in prostate cancer (PCa) detection using deep convolutional neural networks (CNNs). Recently, transformers have achieved competitive performance compared to CNNs in computer vision. Large-scale transformers need abundant annotated data for training, which are difficult to obtain in medical imaging. Self-supervised learning can effectively leverage unlabeled data to extract useful semantic representations without annotation and its associated costs. This can improve model performance on downstream tasks with limited labelled data and increase generalizability. We introduce a novel end-to-end Cross-Shaped windows (CSwin) transformer UNet model, CSwin UNet, to detect clinically significant prostate cancer (csPCa) in prostate bi-parametric MR imaging (bpMRI) and demonstrate the effectiveness of our proposed self-supervised pre-training framework. Using a large prostate bpMRI dataset with 1500 patients, we first pre-train CSwin transformer using multi-task self-supervised learning to improve data-efficiency and network generalizability. We then finetuned using lesion annotations to perform csPCa detection. Five-fold cross validation shows that self-supervised CSwin UNet achieves 0.888 AUC and 0.545 Average Precision (AP), significantly outperforming four state-of-the-art models (Swin UNETR, DynUNet, Attention UNet, UNet). Using a separate bpMRI dataset with 158 patients, we evaluated our model robustness to external hold-out data. Self-supervised CSwin UNet achieves 0.79 AUC and 0.45 AP, still outperforming all other comparable methods and demonstrating generalization to a dataset shift.
Deep Learning-based Multi-Organ CT Segmentation with Adversarial Data Augmentation
Feb 25, 2023Abstract:In this work, we propose an adversarial attack-based data augmentation method to improve the deep-learning-based segmentation algorithm for the delineation of Organs-At-Risk (OAR) in abdominal Computed Tomography (CT) to facilitate radiation therapy. We introduce Adversarial Feature Attack for Medical Image (AFA-MI) augmentation, which forces the segmentation network to learn out-of-distribution statistics and improve generalization and robustness to noises. AFA-MI augmentation consists of three steps: 1) generate adversarial noises by Fast Gradient Sign Method (FGSM) on the intermediate features of the segmentation network's encoder; 2) inject the generated adversarial noises into the network, intentionally compromising performance; 3) optimize the network with both clean and adversarial features. Experiments are conducted segmenting the heart, left and right kidney, liver, left and right lung, spinal cord, and stomach. We first evaluate the AFA-MI augmentation using nnUnet and TT-Vnet on the test data from a public abdominal dataset and an institutional dataset. In addition, we validate how AFA-MI affects the networks' robustness to the noisy data by evaluating the networks with added Gaussian noises of varying magnitudes to the institutional dataset. Network performance is quantitatively evaluated using Dice Similarity Coefficient (DSC) for volume-based accuracy. Also, Hausdorff Distance (HD) is applied for surface-based accuracy. On the public dataset, nnUnet with AFA-MI achieves DSC = 0.85 and HD = 6.16 millimeters (mm); and TT-Vnet achieves DSC = 0.86 and HD = 5.62 mm. AFA-MI augmentation further improves all contour accuracies up to 0.217 DSC score when tested on images with Gaussian noises. AFA-MI augmentation is therefore demonstrated to improve segmentation performance and robustness in CT multi-organ segmentation.
Artificial Intelligence in Tumor Subregion Analysis Based on Medical Imaging: A Review
Mar 25, 2021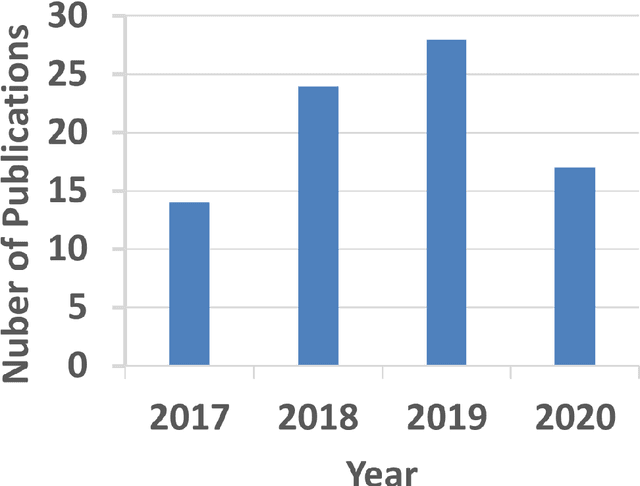
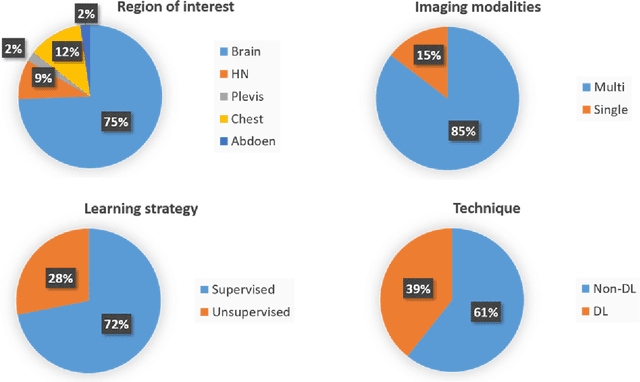

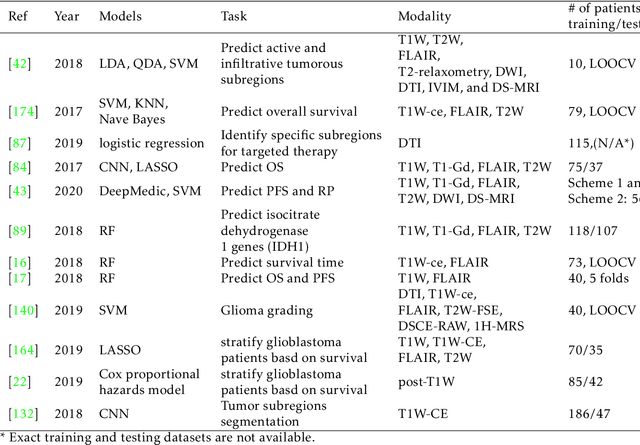
Abstract:Medical imaging is widely used in cancer diagnosis and treatment, and artificial intelligence (AI) has achieved tremendous success in various tasks of medical image analysis. This paper reviews AI-based tumor subregion analysis in medical imaging. We summarize the latest AI-based methods for tumor subregion analysis and their applications. Specifically, we categorize the AI-based methods by training strategy: supervised and unsupervised. A detailed review of each category is presented, highlighting important contributions and achievements. Specific challenges and potential AI applications in tumor subregion analysis are discussed.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge