Hyung-Chul Lee
Coding-Free and Privacy-Preserving MCP Framework for Clinical Agentic Research Intelligence System
Apr 14, 2026Abstract:Clinical research involves labor-intensive processes such as study design, cohort construction, model development, and documentation, requiring domain expertise, programming skills, and access to sensitive patient data. These demands create barriers for clinicians and external researchers conducting data-driven studies. To overcome these limitations, we developed a Clinical Agentic Research Intelligence System (CARIS) that automates the clinical research workflow while preserving data privacy, enabling comprehensive studies without direct access to raw data. CARIS integrates Large Language Models (LLMs) with modular tools via the Model Context Protocol (MCP), enabling natural language-driven orchestration of appropriate tools. Databases remain securely within the MCP server, and users access only the outputs and final research reports. Based on user intent, CARIS automatically executes the full pipeline: research planning, literature search, cohort construction, Institutional Review Board (IRB) documentation, Vibe Machine Learning (ML), and report generation, with iterative human-in-the-loop refinement. We evaluated CARIS on three heterogeneous datasets with distinct clinical tasks. Research plans and IRB documents were finalized within three to four iterations, using evidence from literature and data. The system supported Vibe ML by exploring feature-model combinations, ranking the top ten models, and generating performance visualizations. Final reports showed high completeness based on a checklist derived from the TRIPOD+AI framework, achieving 96% coverage in LLM evaluation and 82% in human evaluation. CARIS demonstrates that agentic AI can transform clinical hypotheses into executable research workflows across heterogeneous datasets. By eliminating the need for coding and direct data access, the system lowers barriers and bridges public and private clinical data environments.
A Multi-faceted Analysis of Cognitive Abilities: Evaluating Prompt Methods with Large Language Models on the CONSORT Checklist
Oct 22, 2025Abstract:Despite the rapid expansion of Large Language Models (LLMs) in healthcare, the ability of these systems to assess clinical trial reporting according to CONSORT standards remains unclear, particularly with respect to their cognitive and reasoning strategies. This study applies a behavioral and metacognitive analytic approach with expert-validated data, systematically comparing two representative LLMs under three prompt conditions. Clear differences emerged in how the models approached various CONSORT items, and prompt types, including shifts in reasoning style, explicit uncertainty, and alternative interpretations shaped response patterns. Our results highlight the current limitations of these systems in clinical compliance automation and underscore the importance of understanding their cognitive adaptations and strategic behavior in developing more explainable and reliable medical AI.
EchoQA: A Large Collection of Instruction Tuning Data for Echocardiogram Reports
Mar 06, 2025Abstract:We introduce a novel question-answering (QA) dataset using echocardiogram reports sourced from the Medical Information Mart for Intensive Care database. This dataset is specifically designed to enhance QA systems in cardiology, consisting of 771,244 QA pairs addressing a wide array of cardiac abnormalities and their severity. We compare large language models (LLMs), including open-source and biomedical-specific models for zero-shot evaluation, and closed-source models for zero-shot and three-shot evaluation. Our results show that fine-tuning LLMs improves performance across various QA metrics, validating the value of our dataset. Clinicians also qualitatively evaluate the best-performing model to assess the LLM responses for correctness. Further, we conduct fine-grained fairness audits to assess the bias-performance trade-off of LLMs across various social determinants of health. Our objective is to propel the field forward by establishing a benchmark for LLM AI agents aimed at supporting clinicians with cardiac differential diagnoses, thereby reducing the documentation burden that contributes to clinician burnout and enabling healthcare professionals to focus more on patient care.
Unmasking Societal Biases in Respiratory Support for ICU Patients through Social Determinants of Health
Feb 23, 2025Abstract:In critical care settings, where precise and timely interventions are crucial for health outcomes, evaluating disparities in patient outcomes is essential. Current approaches often fail to fully capture the impact of respiratory support interventions on individuals affected by social determinants of health. While attributes such as gender, race, and age are commonly assessed and provide valuable insights, they offer only a partial view of the complexities faced by diverse populations. In this study, we focus on two clinically motivated tasks: prolonged mechanical ventilation and successful weaning. Additionally, we conduct fairness audits on the models' predictions across demographic groups and social determinants of health to better understand health inequities in respiratory interventions within the intensive care unit. Furthermore, we release a temporal benchmark dataset, verified by clinical experts, to facilitate benchmarking of clinical respiratory intervention tasks.
Attention mechanisms for physiological signal deep learning: which attention should we take?
Jul 04, 2022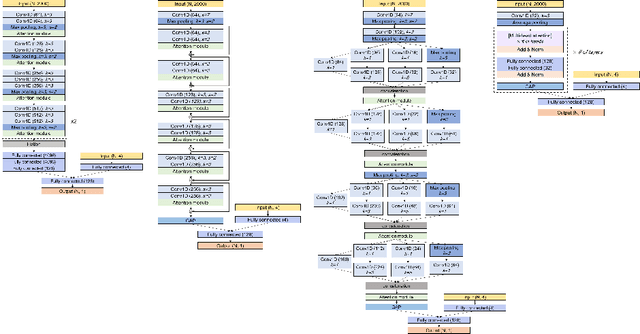
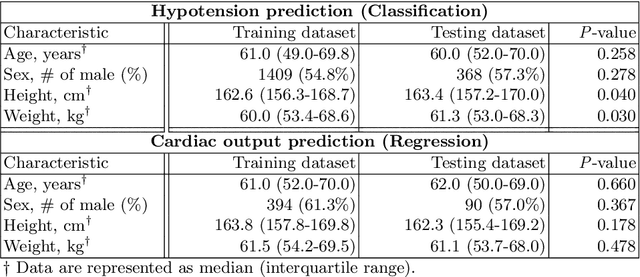
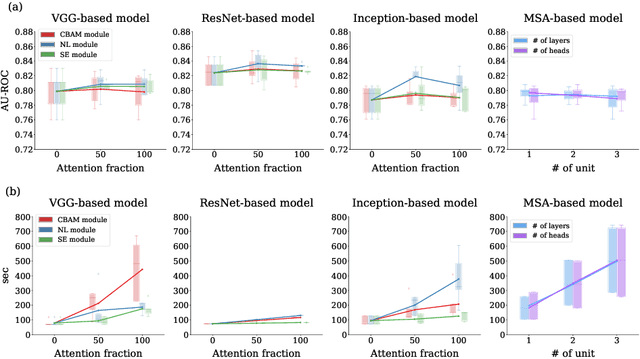
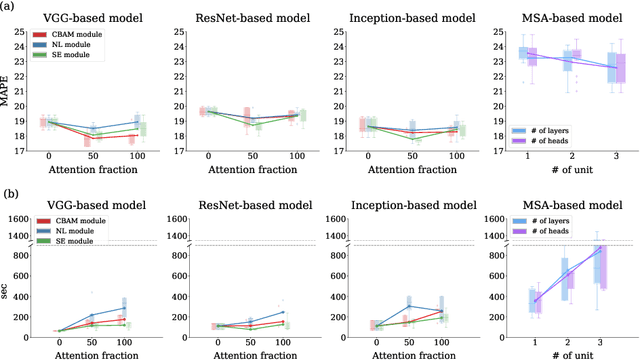
Abstract:Attention mechanisms are widely used to dramatically improve deep learning model performance in various fields. However, their general ability to improve the performance of physiological signal deep learning model is immature. In this study, we experimentally analyze four attention mechanisms (e.g., squeeze-and-excitation, non-local, convolutional block attention module, and multi-head self-attention) and three convolutional neural network (CNN) architectures (e.g., VGG, ResNet, and Inception) for two representative physiological signal prediction tasks: the classification for predicting hypotension and the regression for predicting cardiac output (CO). We evaluated multiple combinations for performance and convergence of physiological signal deep learning model. Accordingly, the CNN models with the spatial attention mechanism showed the best performance in the classification problem, whereas the channel attention mechanism achieved the lowest error in the regression problem. Moreover, the performance and convergence of the CNN models with attention mechanisms were better than stand-alone self-attention models in both problems. Hence, we verified that convolutional operation and attention mechanisms are complementary and provide faster convergence time, despite the stand-alone self-attention models requiring fewer parameters.
Developing and validating multi-modal models for mortality prediction in COVID-19 patients: a multi-center retrospective study
Sep 01, 2021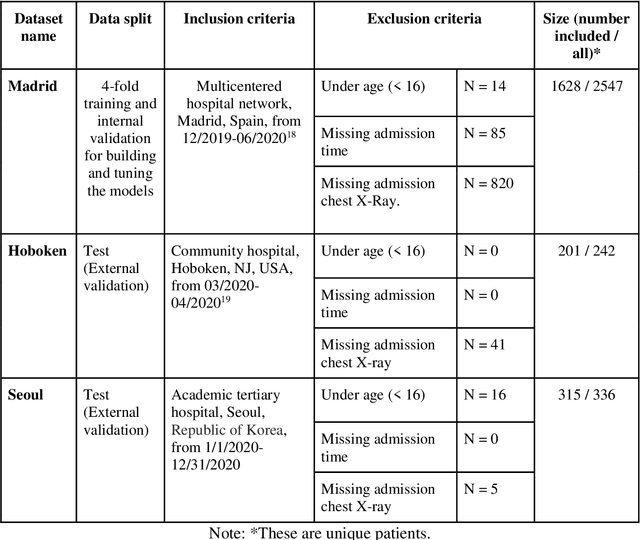
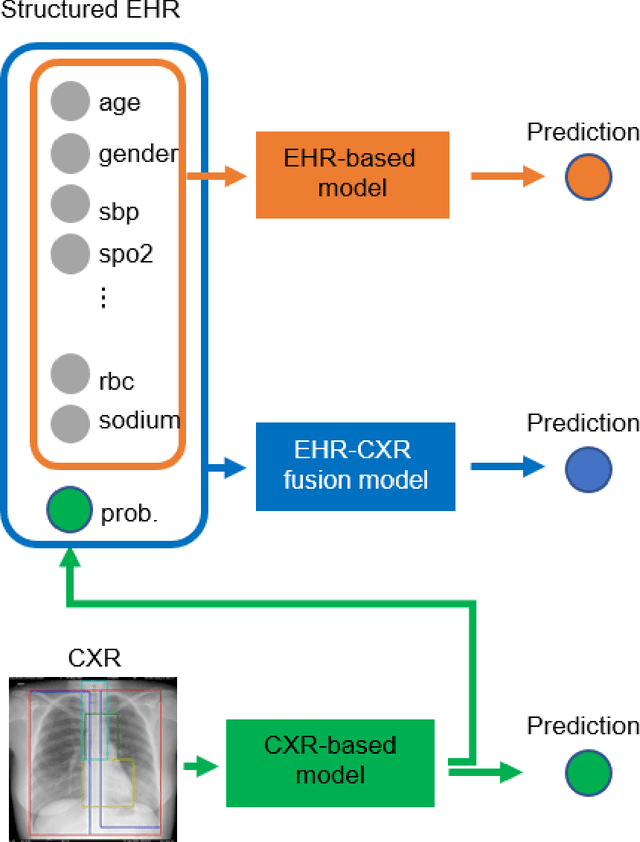
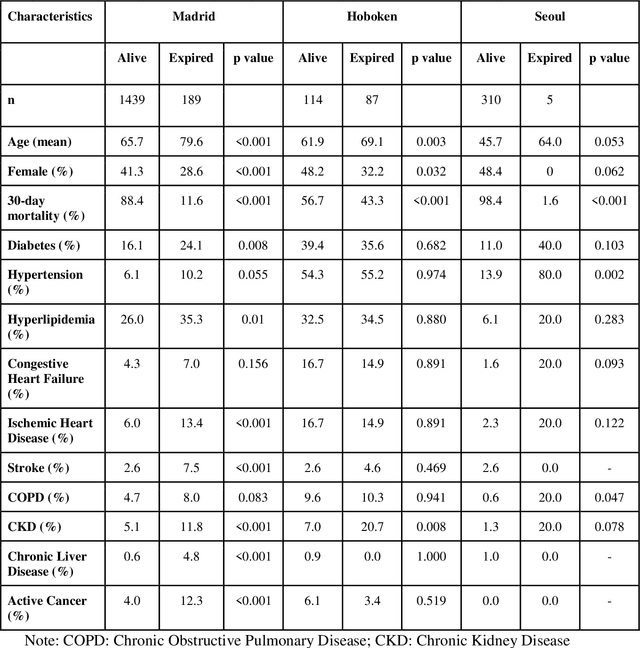
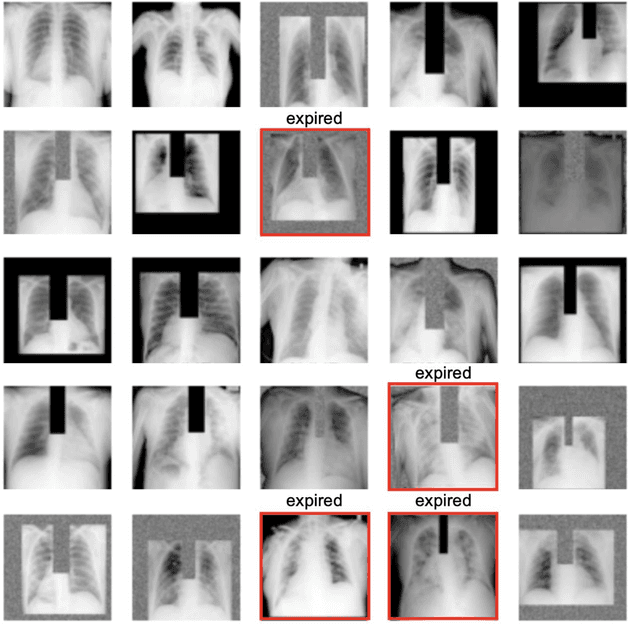
Abstract:The unprecedented global crisis brought about by the COVID-19 pandemic has sparked numerous efforts to create predictive models for the detection and prognostication of SARS-CoV-2 infections with the goal of helping health systems allocate resources. Machine learning models, in particular, hold promise for their ability to leverage patient clinical information and medical images for prediction. However, most of the published COVID-19 prediction models thus far have little clinical utility due to methodological flaws and lack of appropriate validation. In this paper, we describe our methodology to develop and validate multi-modal models for COVID-19 mortality prediction using multi-center patient data. The models for COVID-19 mortality prediction were developed using retrospective data from Madrid, Spain (N=2547) and were externally validated in patient cohorts from a community hospital in New Jersey, USA (N=242) and an academic center in Seoul, Republic of Korea (N=336). The models we developed performed differently across various clinical settings, underscoring the need for a guided strategy when employing machine learning for clinical decision-making. We demonstrated that using features from both the structured electronic health records and chest X-ray imaging data resulted in better 30-day-mortality prediction performance across all three datasets (areas under the receiver operating characteristic curves: 0.85 (95% confidence interval: 0.83-0.87), 0.76 (0.70-0.82), and 0.95 (0.92-0.98)). We discuss the rationale for the decisions made at every step in developing the models and have made our code available to the research community. We employed the best machine learning practices for clinical model development. Our goal is to create a toolkit that would assist investigators and organizations in building multi-modal models for prediction, classification and/or optimization.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge