Huan Tran
PolyGET: Accelerating Polymer Simulations by Accurate and Generalizable Forcefield with Equivariant Transformer
Sep 01, 2023



Abstract:Polymer simulation with both accuracy and efficiency is a challenging task. Machine learning (ML) forcefields have been developed to achieve both the accuracy of ab initio methods and the efficiency of empirical force fields. However, existing ML force fields are usually limited to single-molecule settings, and their simulations are not robust enough. In this paper, we present PolyGET, a new framework for Polymer Forcefields with Generalizable Equivariant Transformers. PolyGET is designed to capture complex quantum interactions between atoms and generalize across various polymer families, using a deep learning model called Equivariant Transformers. We propose a new training paradigm that focuses exclusively on optimizing forces, which is different from existing methods that jointly optimize forces and energy. This simple force-centric objective function avoids competing objectives between energy and forces, thereby allowing for learning a unified forcefield ML model over different polymer families. We evaluated PolyGET on a large-scale dataset of 24 distinct polymer types and demonstrated state-of-the-art performance in force accuracy and robust MD simulations. Furthermore, PolyGET can simulate large polymers with high fidelity to the reference ab initio DFT method while being able to generalize to unseen polymers.
May the Force be with You: Unified Force-Centric Pre-Training for 3D Molecular Conformations
Aug 24, 2023Abstract:Recent works have shown the promise of learning pre-trained models for 3D molecular representation. However, existing pre-training models focus predominantly on equilibrium data and largely overlook off-equilibrium conformations. It is challenging to extend these methods to off-equilibrium data because their training objective relies on assumptions of conformations being the local energy minima. We address this gap by proposing a force-centric pretraining model for 3D molecular conformations covering both equilibrium and off-equilibrium data. For off-equilibrium data, our model learns directly from their atomic forces. For equilibrium data, we introduce zero-force regularization and forced-based denoising techniques to approximate near-equilibrium forces. We obtain a unified pre-trained model for 3D molecular representation with over 15 million diverse conformations. Experiments show that, with our pre-training objective, we increase forces accuracy by around 3 times compared to the un-pre-trained Equivariant Transformer model. By incorporating regularizations on equilibrium data, we solved the problem of unstable MD simulations in vanilla Equivariant Transformers, achieving state-of-the-art simulation performance with 2.45 times faster inference time than NequIP. As a powerful molecular encoder, our pre-trained model achieves on-par performance with state-of-the-art property prediction tasks.
Polymer Informatics with Multi-Task Learning
Oct 28, 2020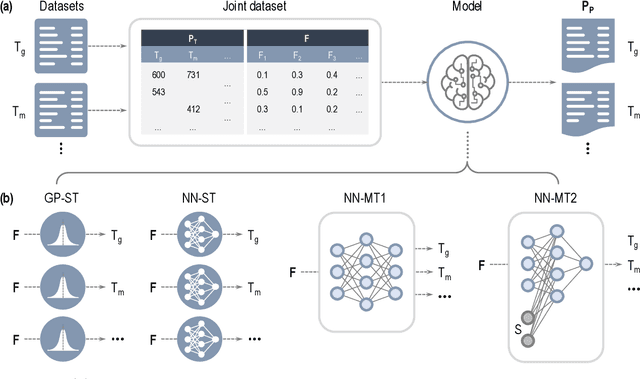
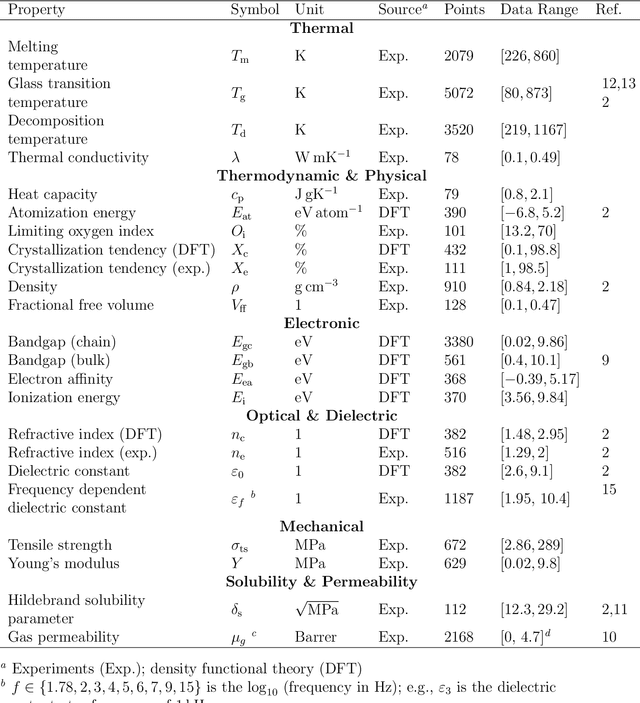
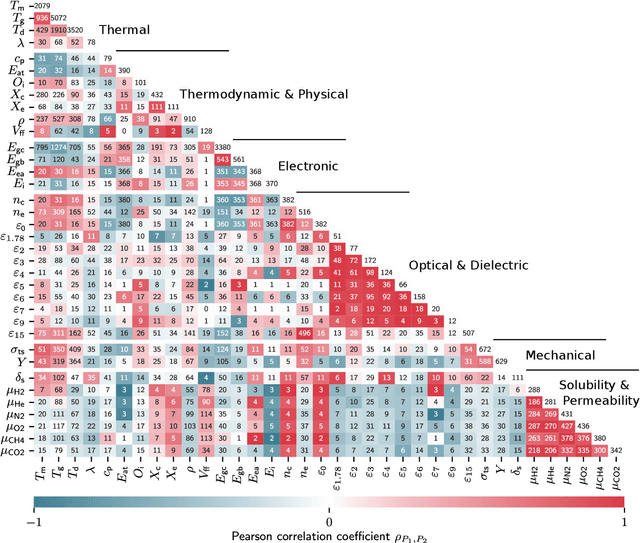
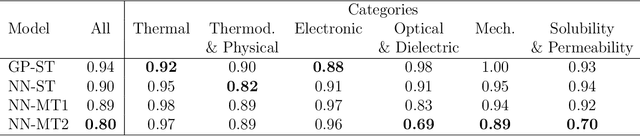
Abstract:Modern data-driven tools are transforming application-specific polymer development cycles. Surrogate models that can be trained to predict the properties of new polymers are becoming commonplace. Nevertheless, these models do not utilize the full breadth of the knowledge available in datasets, which are oftentimes sparse; inherent correlations between different property datasets are disregarded. Here, we demonstrate the potency of multi-task learning approaches that exploit such inherent correlations effectively, particularly when some property dataset sizes are small. Data pertaining to 36 different properties of over $13, 000$ polymers (corresponding to over $23,000$ data points) are coalesced and supplied to deep-learning multi-task architectures. Compared to conventional single-task learning models (that are trained on individual property datasets independently), the multi-task approach is accurate, efficient, scalable, and amenable to transfer learning as more data on the same or different properties become available. Moreover, these models are interpretable. Chemical rules, that explain how certain features control trends in specific property values, emerge from the present work, paving the way for the rational design of application specific polymers meeting desired property or performance objectives.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge