Henkjan Huisman
and on behalf of the UNICORN consortium
Fully Automatic Deep Learning Framework for Pancreatic Ductal Adenocarcinoma Detection on Computed Tomography
Dec 02, 2021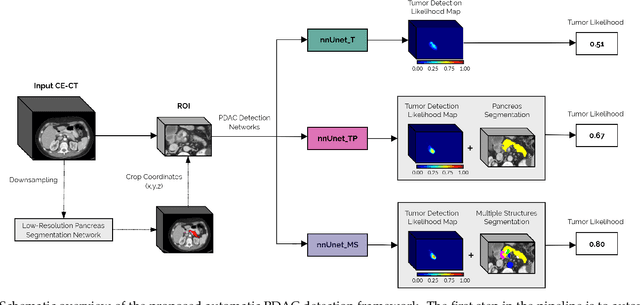
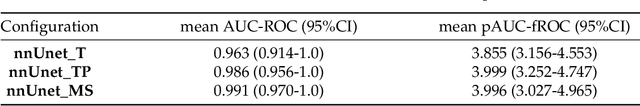

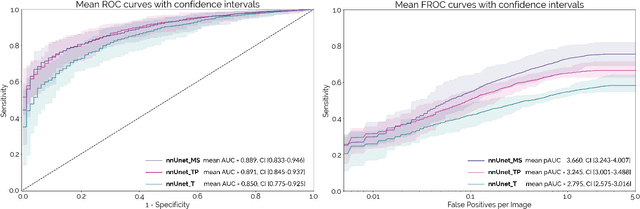
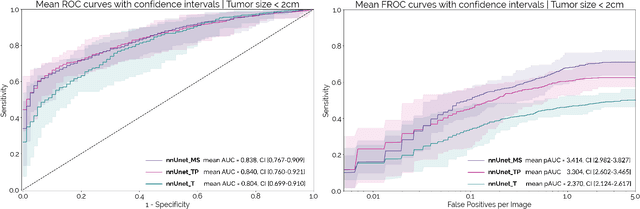
Abstract:Early detection improves prognosis in pancreatic ductal adenocarcinoma (PDAC) but is challenging as lesions are often small and poorly defined on contrast-enhanced computed tomography scans (CE-CT). Deep learning can facilitate PDAC diagnosis, however current models still fail to identify small (<2cm) lesions. In this study, state-of-the-art deep learning models were used to develop an automatic framework for PDAC detection, focusing on small lesions. Additionally, the impact of integrating surrounding anatomy was investigated. CE-CT scans from a cohort of 119 pathology-proven PDAC patients and a cohort of 123 patients without PDAC were used to train a nnUnet for automatic lesion detection and segmentation (nnUnet_T). Two additional nnUnets were trained to investigate the impact of anatomy integration: (1) segmenting the pancreas and tumor (nnUnet_TP), (2) segmenting the pancreas, tumor, and multiple surrounding anatomical structures (nnUnet_MS). An external, publicly available test set was used to compare the performance of the three networks. The nnUnet_MS achieved the best performance, with an area under the receiver operating characteristic curve of 0.91 for the whole test set and 0.88 for tumors <2cm, showing that state-of-the-art deep learning can detect small PDAC and benefits from anatomy information.
Anatomical and Diagnostic Bayesian Segmentation in Prostate MRI $-$Should Different Clinical Objectives Mandate Different Loss Functions?
Oct 25, 2021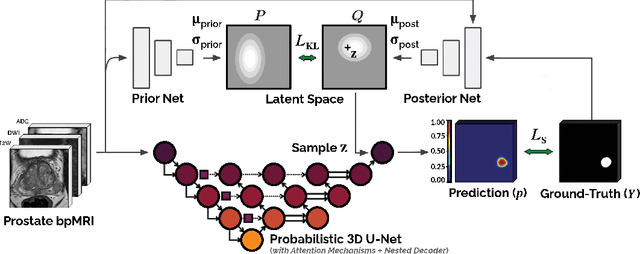
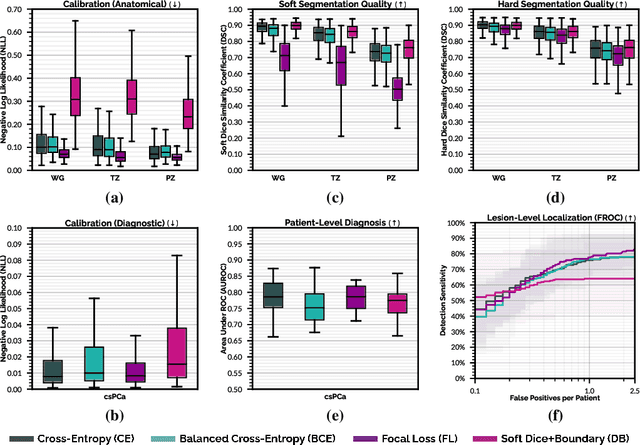
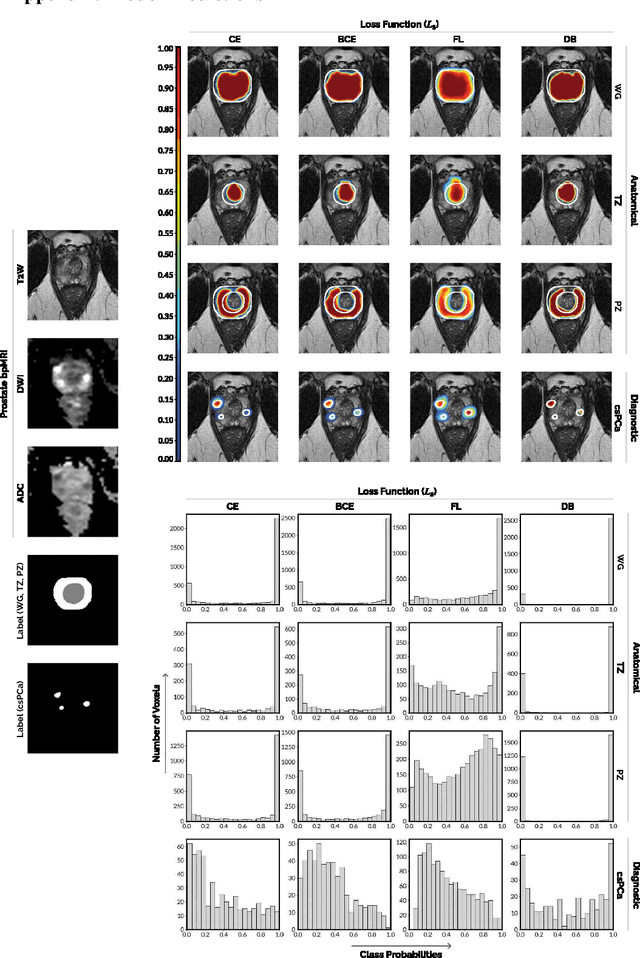
Abstract:We hypothesize that probabilistic voxel-level classification of anatomy and malignancy in prostate MRI, although typically posed as near-identical segmentation tasks via U-Nets, require different loss functions for optimal performance due to inherent differences in their clinical objectives. We investigate distribution, region and boundary-based loss functions for both tasks across 200 patient exams from the publicly-available ProstateX dataset. For evaluation, we conduct a thorough comparative analysis of model predictions and calibration, measured with respect to multi-class volume segmentation of the prostate anatomy (whole-gland, transitional zone, peripheral zone), as well as, patient-level diagnosis and lesion-level detection of clinically significant prostate cancer. Notably, we find that distribution-based loss functions (in particular, focal loss) are well-suited for diagnostic or panoptic segmentation tasks such as lesion detection, primarily due to their implicit property of inducing better calibration. Meanwhile, (with the exception of focal loss) both distribution and region/boundary-based loss functions perform equally well for anatomical or semantic segmentation tasks, such as quantification of organ shape, size and boundaries.
Cine-MRI detection of abdominal adhesions with spatio-temporal deep learning
Jun 15, 2021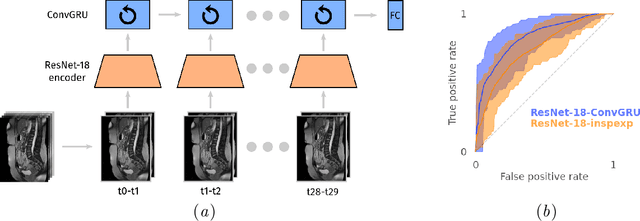
Abstract:Adhesions are an important cause of chronic pain following abdominal surgery. Recent developments in abdominal cine-MRI have enabled the non-invasive diagnosis of adhesions. Adhesions are identified on cine-MRI by the absence of sliding motion during movement. Diagnosis and mapping of adhesions improves the management of patients with pain. Detection of abdominal adhesions on cine-MRI is challenging from both a radiological and deep learning perspective. We focus on classifying presence or absence of adhesions in sagittal abdominal cine-MRI series. We experimented with spatio-temporal deep learning architectures centered around a ConvGRU architecture. A hybrid architecture comprising a ResNet followed by a ConvGRU model allows to classify a whole time-series. Compared to a stand-alone ResNet with a two time-point (inspiration/expiration) input, we show an increase in classification performance (AUROC) from 0.74 to 0.83 ($p<0.05$). Our full temporal classification approach adds only a small amount (5%) of parameters to the entire architecture, which may be useful for other medical imaging problems with a temporal dimension.
The Medical Segmentation Decathlon
Jun 10, 2021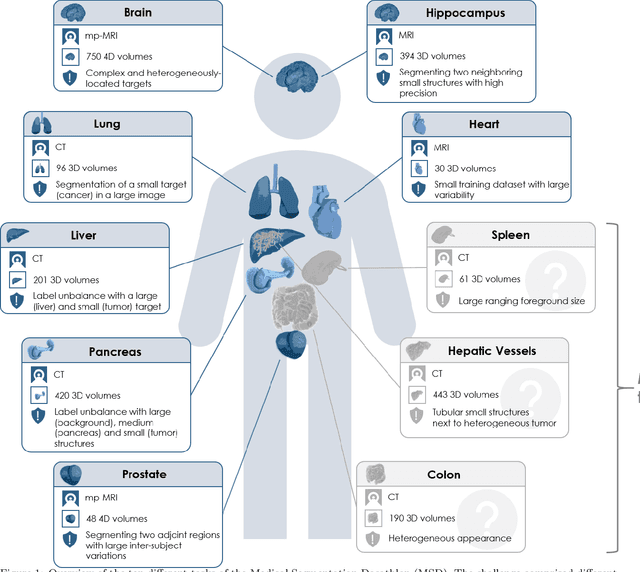
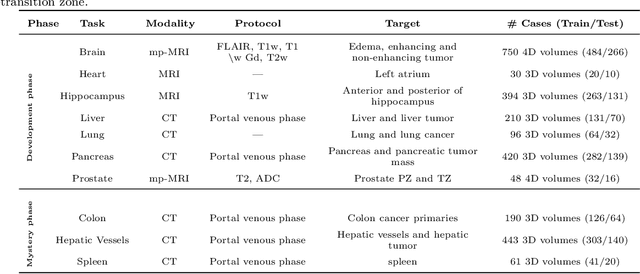
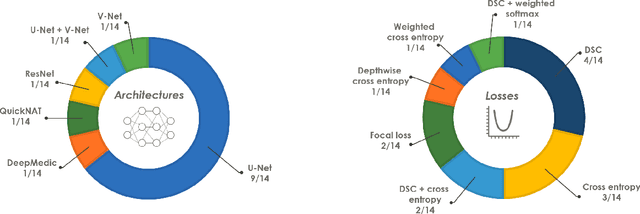
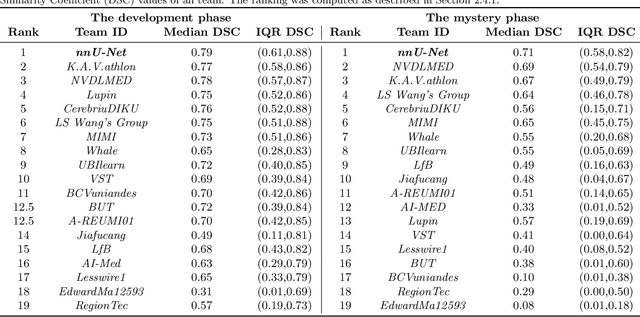
Abstract:International challenges have become the de facto standard for comparative assessment of image analysis algorithms given a specific task. Segmentation is so far the most widely investigated medical image processing task, but the various segmentation challenges have typically been organized in isolation, such that algorithm development was driven by the need to tackle a single specific clinical problem. We hypothesized that a method capable of performing well on multiple tasks will generalize well to a previously unseen task and potentially outperform a custom-designed solution. To investigate the hypothesis, we organized the Medical Segmentation Decathlon (MSD) - a biomedical image analysis challenge, in which algorithms compete in a multitude of both tasks and modalities. The underlying data set was designed to explore the axis of difficulties typically encountered when dealing with medical images, such as small data sets, unbalanced labels, multi-site data and small objects. The MSD challenge confirmed that algorithms with a consistent good performance on a set of tasks preserved their good average performance on a different set of previously unseen tasks. Moreover, by monitoring the MSD winner for two years, we found that this algorithm continued generalizing well to a wide range of other clinical problems, further confirming our hypothesis. Three main conclusions can be drawn from this study: (1) state-of-the-art image segmentation algorithms are mature, accurate, and generalize well when retrained on unseen tasks; (2) consistent algorithmic performance across multiple tasks is a strong surrogate of algorithmic generalizability; (3) the training of accurate AI segmentation models is now commoditized to non AI experts.
End-to-end Prostate Cancer Detection in bpMRI via 3D CNNs: Effect of Attention Mechanisms, Clinical Priori and Decoupled False Positive Reduction
Jan 28, 2021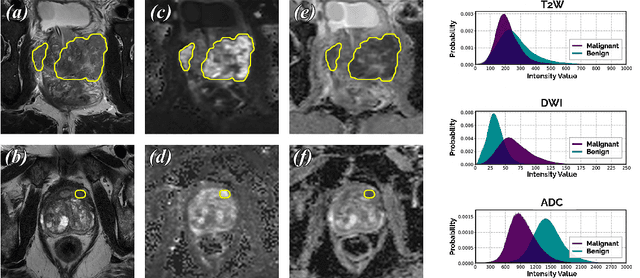
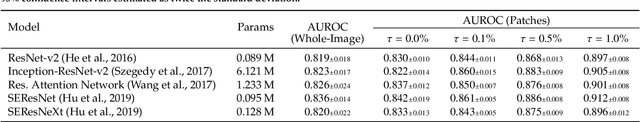
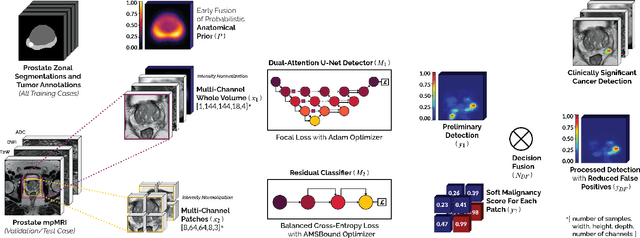
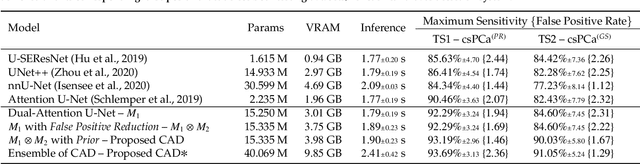
Abstract:We present a novel multi-stage 3D computer-aided detection and diagnosis (CAD) model for automated localization of clinically significant prostate cancer (csPCa) in bi-parametric MR imaging (bpMRI). Deep attention mechanisms drive its detection network, targeting multi-resolution, salient structures and highly discriminative feature dimensions, in order to accurately identify csPCa lesions from indolent cancer and the wide range of benign pathology that can afflict the prostate gland. In parallel, a decoupled residual classifier is used to achieve consistent false positive reduction, without sacrificing high sensitivity or computational efficiency. Furthermore, a probabilistic anatomical prior, which captures the spatial prevalence of csPCa as well as its zonal distinction, is computed and encoded into the CNN architecture to guide model generalization with domain-specific clinical knowledge. For 486 institutional testing scans, the 3D CAD system achieves $83.69\pm5.22\%$ and $93.19\pm2.96\%$ detection sensitivity at 0.50 and 1.46 false positive(s) per patient, respectively, along with $0.882$ AUROC in patient-based diagnosis $-$significantly outperforming four state-of-the-art baseline architectures (U-SEResNet, UNet++, nnU-Net, Attention U-Net) from recent literature. For 296 external testing scans, the ensembled CAD system shares moderate agreement with a consensus of expert radiologists ($76.69\%$; $kappa=0.511$) and independent pathologists ($81.08\%$; $kappa=0.559$); demonstrating strong generalization to histologically-confirmed malignancies, despite using 1950 training-validation cases with radiologically-estimated annotations only.
Encoding Clinical Priori in 3D Convolutional Neural Networks for Prostate Cancer Detection in bpMRI
Nov 03, 2020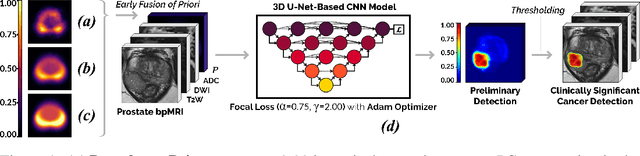
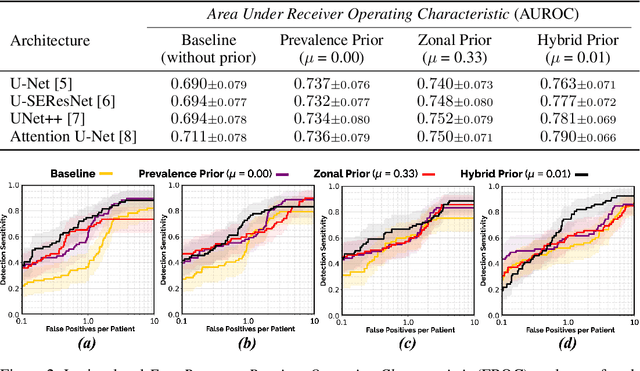
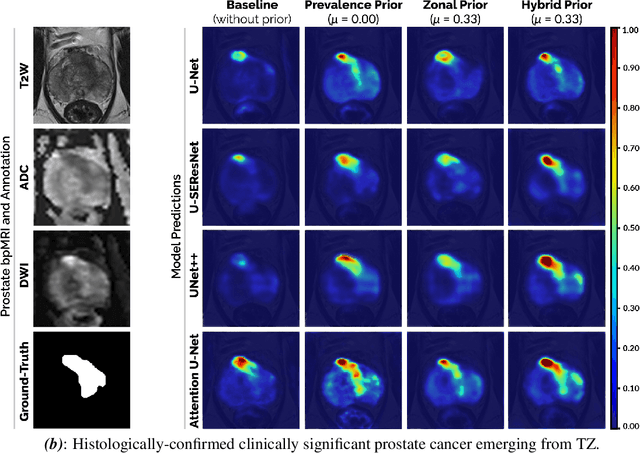
Abstract:We hypothesize that anatomical priors can be viable mediums to infuse domain-specific clinical knowledge into state-of-the-art convolutional neural networks (CNN) based on the U-Net architecture. We introduce a probabilistic population prior which captures the spatial prevalence and zonal distinction of clinically significant prostate cancer (csPCa), in order to improve its computer-aided detection (CAD) in bi-parametric MR imaging (bpMRI). To evaluate performance, we train 3D adaptations of the U-Net, U-SEResNet, UNet++ and Attention U-Net using 800 institutional training-validation scans, paired with radiologically-estimated annotations and our computed prior. For 200 independent testing bpMRI scans with histologically-confirmed delineations of csPCa, our proposed method of encoding clinical priori demonstrates a strong ability to improve patient-based diagnosis (upto 8.70% increase in AUROC) and lesion-level detection (average increase of 1.08 pAUC between 0.1-1.0 false positive per patient) across all four architectures.
Supervised Uncertainty Quantification for Segmentation with Multiple Annotations
Jul 03, 2019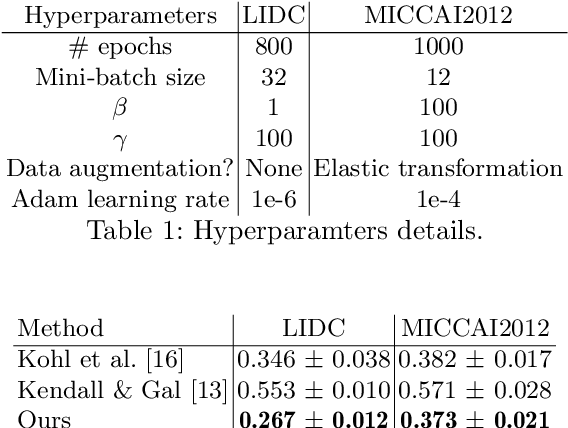

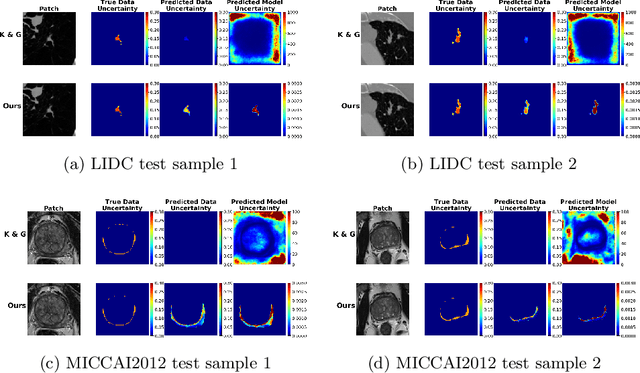
Abstract:The accurate estimation of predictive uncertainty carries importance in medical scenarios such as lung node segmentation. Unfortunately, most existing works on predictive uncertainty do not return calibrated uncertainty estimates, which could be used in practice. In this work we exploit multi-grader annotation variability as a source of 'groundtruth' aleatoric uncertainty, which can be treated as a target in a supervised learning problem. We combine this groundtruth uncertainty with a Probabilistic U-Net and test on the LIDC-IDRI lung nodule CT dataset and MICCAI2012 prostate MRI dataset. We find that we are able to improve predictive uncertainty estimates. We also find that we can improve sample accuracy and sample diversity.
Autoencoders for Multi-Label Prostate MR Segmentation
Jun 26, 2018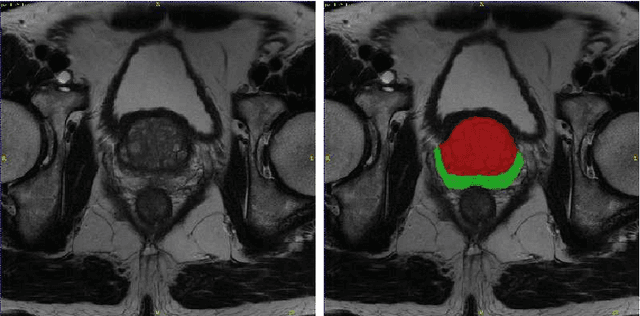

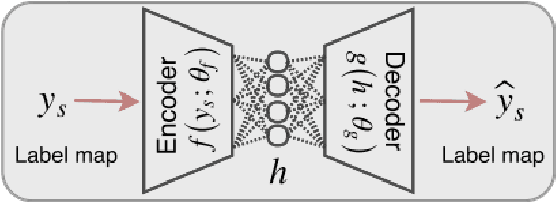

Abstract:Organ image segmentation can be improved by implementing prior knowledge about the anatomy. One way of doing this is by training an autoencoder to learn a lowdimensional representation of the segmentation. In this paper, this is applied in multi-label prostate MR segmentation, with some positive results.
Automatic segmentation of prostate zones
Jun 19, 2018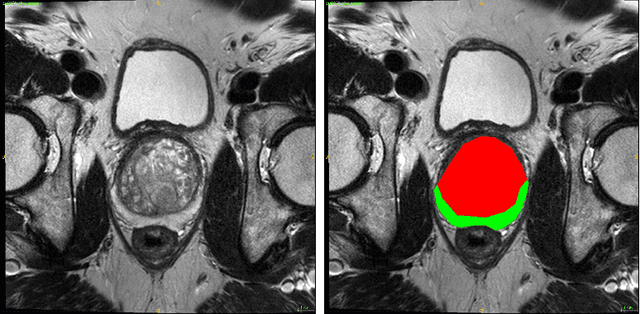
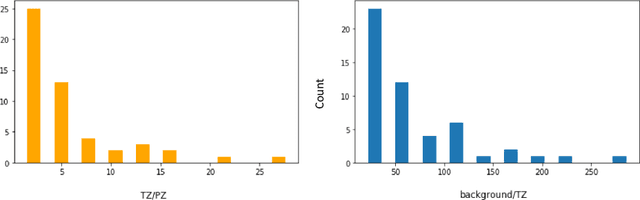
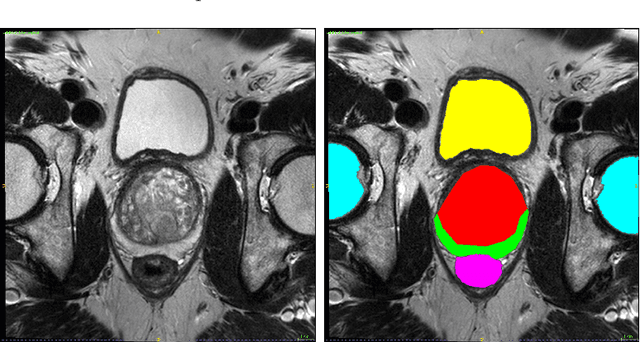
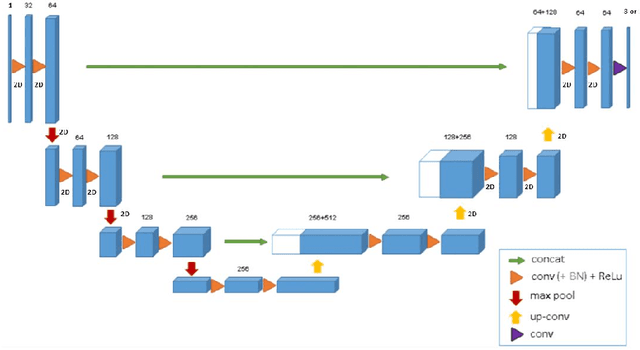
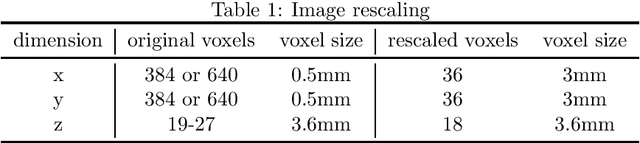
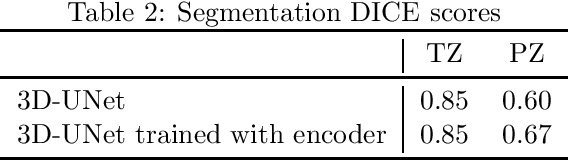
Abstract:Convolutional networks have become state-of-the-art techniques for automatic medical image analysis, with the U-net architecture being the most popular at this moment. In this article we report the application of a 3D version of U-net to the automatic segmentation of prostate peripheral and transition zones in 3D MRI images. Our results are slightly better than recent studies that used 2D U-net and handcrafted feature approaches. In addition, we test ideas for improving the 3D U-net setup, by 1) letting the network segment surrounding tissues, making use of the fixed anatomy, and 2) adjusting the network architecture to reflect the anisotropy in the dimensions of the MRI image volumes. While the latter adjustment gave a marginal improvement, the former adjustment showed a significant deterioration of the network performance. We were able to explain this deterioration by inspecting feature map activations in all layers of the network. We show that to segment more tissues the network replaces feature maps that were dedicated to detecting prostate peripheral zones, by feature maps detecting the surrounding tissues.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge