Heng Wu
Linear-Nonlinear Fusion Neural Operator for Partial Differential Equations
Mar 25, 2026Abstract:Neural operator learning directly constructs the mapping relationship from the equation parameter space to the solution space, enabling efficient direct inference in practical applications without the need for repeated solution of partial differential equations (PDEs) - an advantage that is difficult to achieve with traditional numerical methods. In this work, we find that explicitly decoupling linear and nonlinear effects within such operator mappings leads to markedly improved learning efficiency. This yields a novel network structure, namely the Linear-Nonlinear Fusion Neural Operator (LNF-NO), which models operator mappings via the multiplicative fusion of a linear component and a nonlinear component, thus achieving a lightweight and interpretable representation. This linear-nonlinear decoupling enables efficient capture of complex solution features at the operator level while maintaining stability and generality. LNF-NO naturally supports multiple functional inputs and is applicable to both regular grids and irregular geometries. Across a diverse suite of PDE operator-learning benchmarks, including nonlinear Poisson-Boltzmann equations and multi-physics coupled systems, LNF-NO is typically substantially faster to train than Deep Operator Networks (DeepONet) and Fourier Neural Operators (FNO), while achieving comparable or better accuracy in most cases. On the tested 3D Poisson-Boltzmann case, LNF-NO attains the best accuracy among the compared models and trains approximately 2.7x faster than a 3D FNO baseline.
Operator learning on domain boundary through combining fundamental solution-based artificial data and boundary integral techniques
Jan 16, 2026Abstract:For linear partial differential equations with known fundamental solutions, this work introduces a novel operator learning framework that relies exclusively on domain boundary data, including solution values and normal derivatives, rather than full-domain sampling. By integrating the previously developed Mathematical Artificial Data (MAD) method, which enforces physical consistency, all training data are synthesized directly from the fundamental solutions of the target problems, resulting in a fully data-driven pipeline without the need for external measurements or numerical simulations. We refer to this approach as the Mathematical Artificial Data Boundary Neural Operator (MAD-BNO), which learns boundary-to-boundary mappings using MAD-generated Dirichlet-Neumann data pairs. Once trained, the interior solution at arbitrary locations can be efficiently recovered through boundary integral formulations, supporting Dirichlet, Neumann, and mixed boundary conditions as well as general source terms. The proposed method is validated on benchmark operator learning tasks for two-dimensional Laplace, Poisson, and Helmholtz equations, where it achieves accuracy comparable to or better than existing neural operator approaches while significantly reducing training time. The framework is naturally extensible to three-dimensional problems and complex geometries.
Mathematical artificial data for operator learning
Jul 09, 2025Abstract:Machine learning has emerged as a transformative tool for solving differential equations (DEs), yet prevailing methodologies remain constrained by dual limitations: data-driven methods demand costly labeled datasets while model-driven techniques face efficiency-accuracy trade-offs. We present the Mathematical Artificial Data (MAD) framework, a new paradigm that integrates physical laws with data-driven learning to facilitate large-scale operator discovery. By exploiting DEs' intrinsic mathematical structure to generate physics-embedded analytical solutions and associated synthetic data, MAD fundamentally eliminates dependence on experimental or simulated training data. This enables computationally efficient operator learning across multi-parameter systems while maintaining mathematical rigor. Through numerical demonstrations spanning 2D parametric problems where both the boundary values and source term are functions, we showcase MAD's generalizability and superior efficiency/accuracy across various DE scenarios. This physics-embedded-data-driven framework and its capacity to handle complex parameter spaces gives it the potential to become a universal paradigm for physics-informed machine intelligence in scientific computing.
Hyperspectral image reconstruction for spectral camera based on ghost imaging via sparsity constraints using V-DUnet
Jun 28, 2022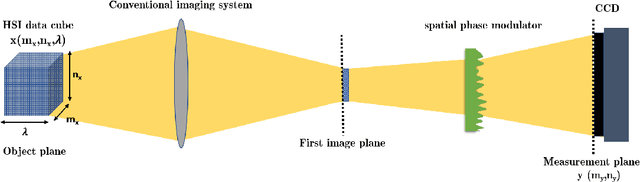
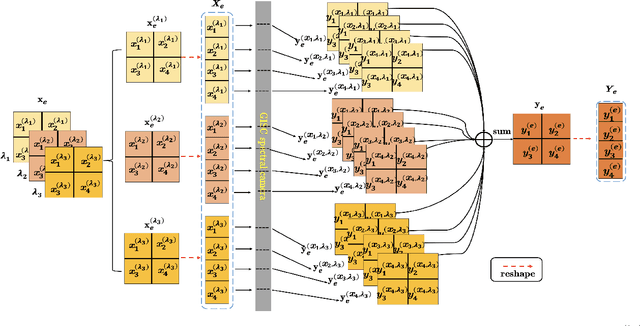
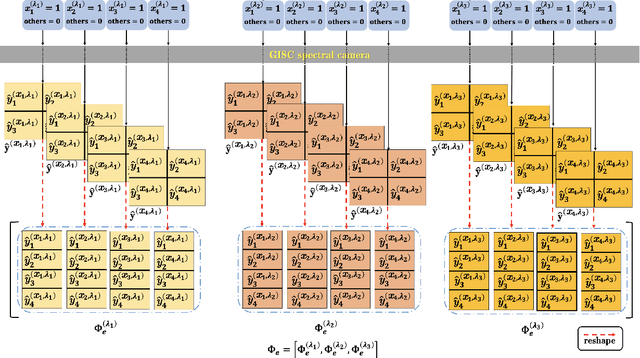
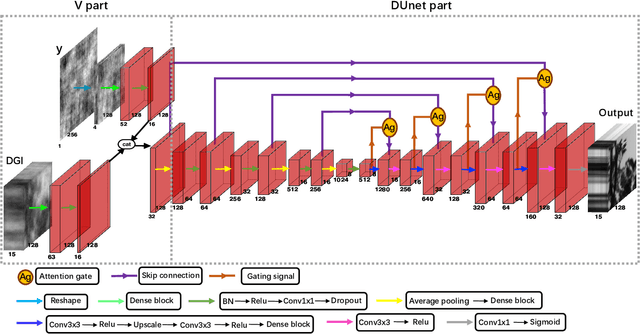
Abstract:Spectral camera based on ghost imaging via sparsity constraints (GISC spectral camera) obtains three-dimensional (3D) hyperspectral information with two-dimensional (2D) compressive measurements in a single shot, which has attracted much attention in recent years. However, its imaging quality and real-time performance of reconstruction still need to be further improved. Recently, deep learning has shown great potential in improving the reconstruction quality and reconstruction speed for computational imaging. When applying deep learning into GISC spectral camera, there are several challenges need to be solved: 1) how to deal with the large amount of 3D hyperspectral data, 2) how to reduce the influence caused by the uncertainty of the random reference measurements, 3) how to improve the reconstructed image quality as far as possible. In this paper, we present an end-to-end V-DUnet for the reconstruction of 3D hyperspectral data in GISC spectral camera. To reduce the influence caused by the uncertainty of the measurement matrix and enhance the reconstructed image quality, both differential ghost imaging results and the detected measurements are sent into the network's inputs. Compared with compressive sensing algorithm, such as PICHCS and TwIST, it not only significantly improves the imaging quality with high noise immunity, but also speeds up the reconstruction time by more than two orders of magnitude.
A Rigid Registration Method in TEVAR
May 19, 2021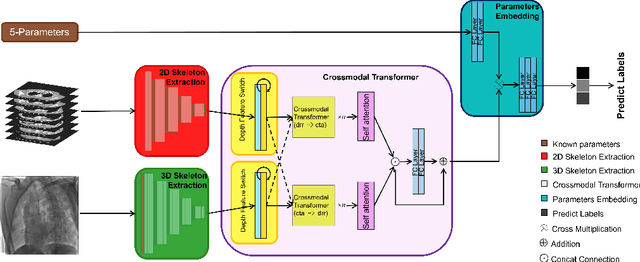

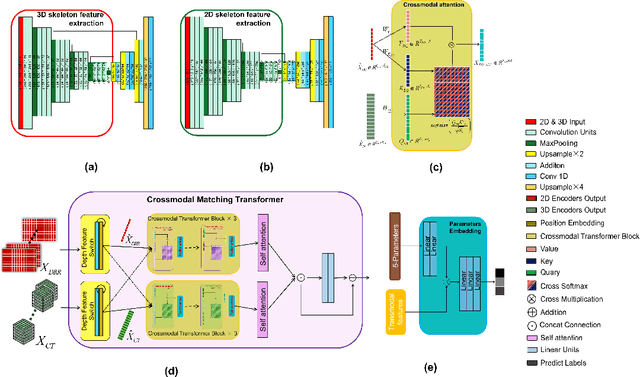
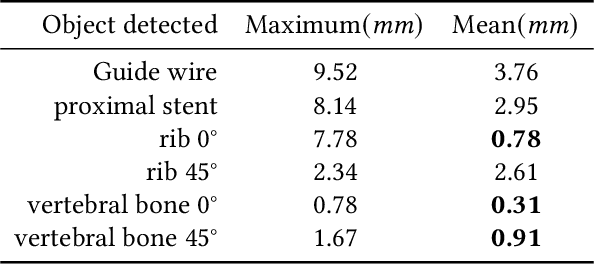
Abstract:Since the mapping relationship between definitized intra-interventional 2D X-ray and undefined pre-interventional 3D Computed Tomography(CT) is uncertain, auxiliary positioning devices or body markers, such as medical implants, are commonly used to determine this relationship. However, such approaches can not be widely used in clinical due to the complex realities. To determine the mapping relationship, and achieve a initializtion post estimation of human body without auxiliary equipment or markers, proposed method applies image segmentation and deep feature matching to directly match the 2D X-ray and 3D CT images. As a result, the well-trained network can directly predict the spatial correspondence between arbitrary 2D X-ray and 3D CT. The experimental results show that when combining our approach with the conventional approach, the achieved accuracy and speed can meet the basic clinical intervention needs, and it provides a new direction for intra-interventional registration.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge