Georg Langs
Department of Biomedical Imaging and Image-guided Therapy, Computational Imaging Research Lab, Medical University Vienna, Austria
The Effects of Skin Lesion Segmentation on the Performance of Dermatoscopic Image Classification
Aug 28, 2020
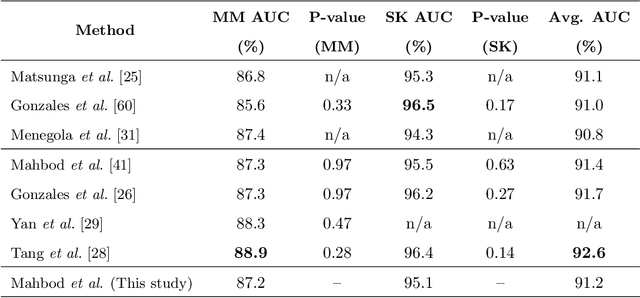
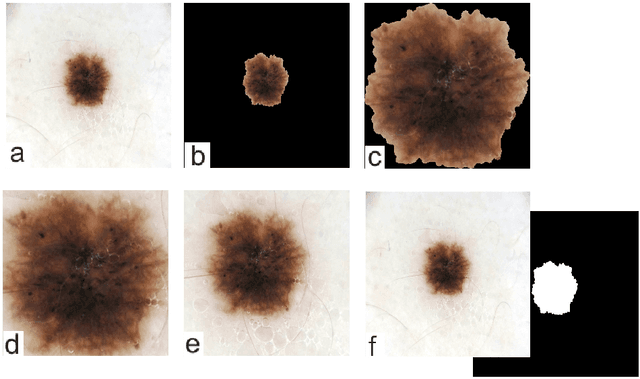
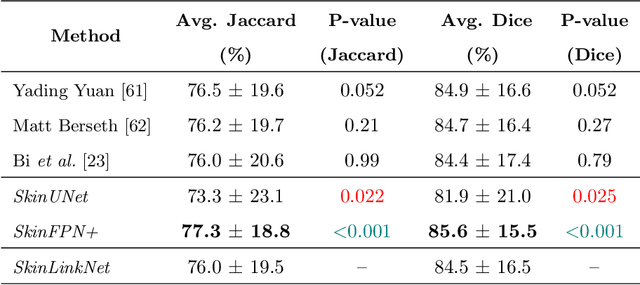
Abstract:Malignant melanoma (MM) is one of the deadliest types of skin cancer. Analysing dermatoscopic images plays an important role in the early detection of MM and other pigmented skin lesions. Among different computer-based methods, deep learning-based approaches and in particular convolutional neural networks have shown excellent classification and segmentation performances for dermatoscopic skin lesion images. These models can be trained end-to-end without requiring any hand-crafted features. However, the effect of using lesion segmentation information on classification performance has remained an open question. In this study, we explicitly investigated the impact of using skin lesion segmentation masks on the performance of dermatoscopic image classification. To do this, first, we developed a baseline classifier as the reference model without using any segmentation masks. Then, we used either manually or automatically created segmentation masks in both training and test phases in different scenarios and investigated the classification performances. Evaluated on the ISIC 2017 challenge dataset which contained two binary classification tasks (i.e. MM vs. all and seborrheic keratosis (SK) vs. all) and based on the derived area under the receiver operating characteristic curve scores, we observed four main outcomes. Our results show that 1) using segmentation masks did not significantly improve the MM classification performance in any scenario, 2) in one of the scenarios (using segmentation masks for dilated cropping), SK classification performance was significantly improved, 3) removing all background information by the segmentation masks significantly degraded the overall classification performance, and 4) in case of using the appropriate scenario (using segmentation for dilated cropping), there is no significant difference of using manually or automatically created segmentation masks.
Dynamic memory to alleviate catastrophic forgetting in continuous learning settings
Jul 07, 2020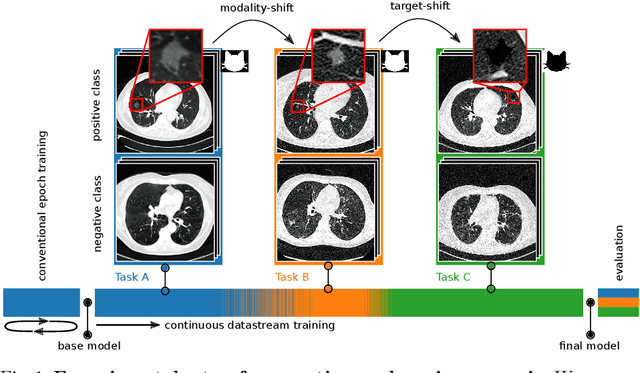
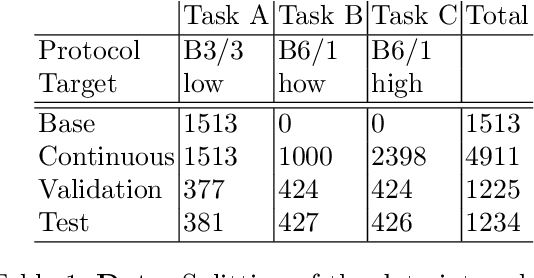
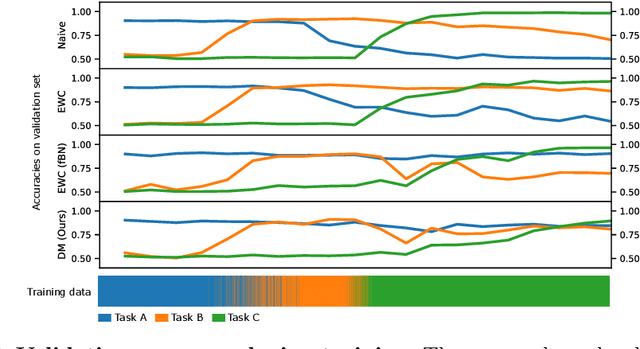
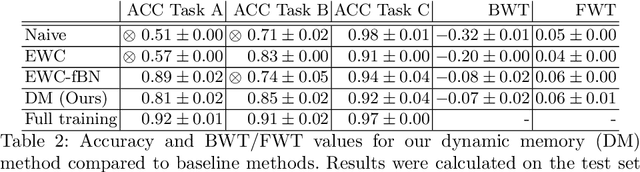
Abstract:In medical imaging, technical progress or changes in diagnostic procedures lead to a continuous change in image appearance. Scanner manufacturer, reconstruction kernel, dose, other protocol specific settings or administering of contrast agents are examples that influence image content independent of the scanned biology. Such domain and task shifts limit the applicability of machine learning algorithms in the clinical routine by rendering models obsolete over time. Here, we address the problem of data shifts in a continuous learning scenario by adapting a model to unseen variations in the source domain while counteracting catastrophic forgetting effects. Our method uses a dynamic memory to facilitate rehearsal of a diverse training data subset to mitigate forgetting. We evaluated our approach on routine clinical CT data obtained with two different scanner protocols and synthetic classification tasks. Experiments show that dynamic memory counters catastrophic forgetting in a setting with multiple data shifts without the necessity for explicit knowledge about when these shifts occur.
Separation of target anatomical structure and occlusions in chest radiographs
Feb 03, 2020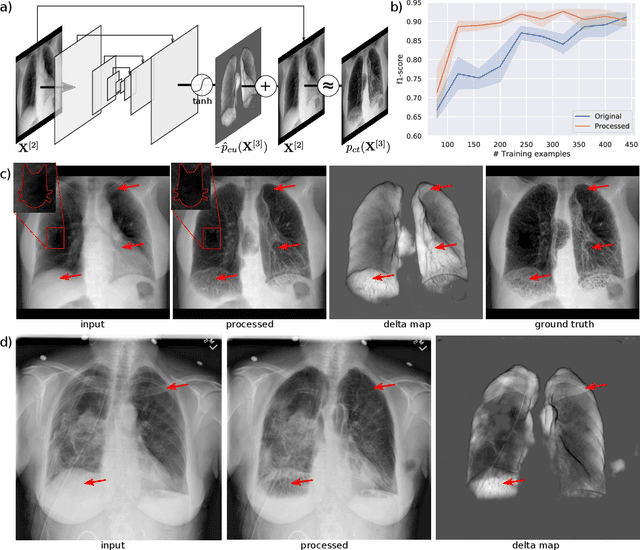
Abstract:Chest radiographs are commonly performed low-cost exams for screening and diagnosis. However, radiographs are 2D representations of 3D structures causing considerable clutter impeding visual inspection and automated image analysis. Here, we propose a Fully Convolutional Network to suppress, for a specific task, undesired visual structure from radiographs while retaining the relevant image information such as lung-parenchyma. The proposed algorithm creates reconstructed radiographs and ground-truth data from high resolution CT-scans. Results show that removing visual variation that is irrelevant for a classification task improves the performance of a classifier when only limited training data are available. This is particularly relevant because a low number of ground-truth cases is common in medical imaging.
Automatic lung segmentation in routine imaging is a data diversity problem, not a methodology problem
Jan 31, 2020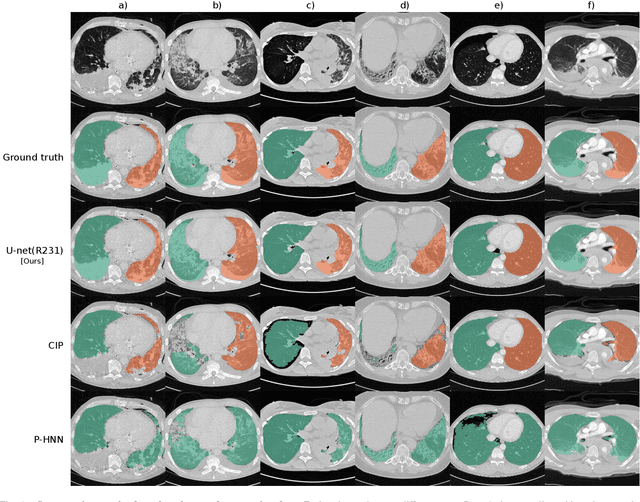
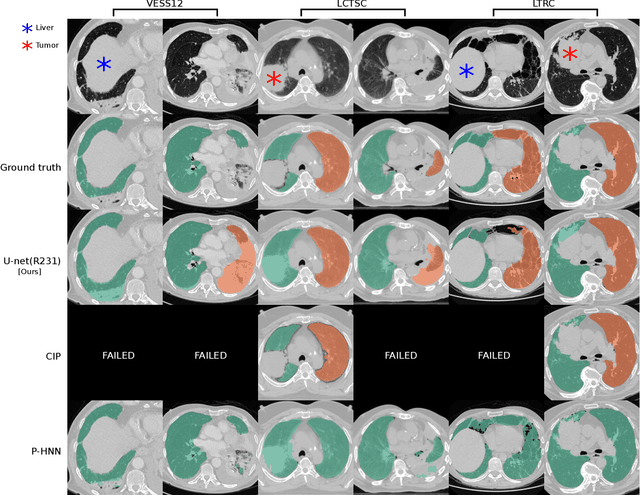
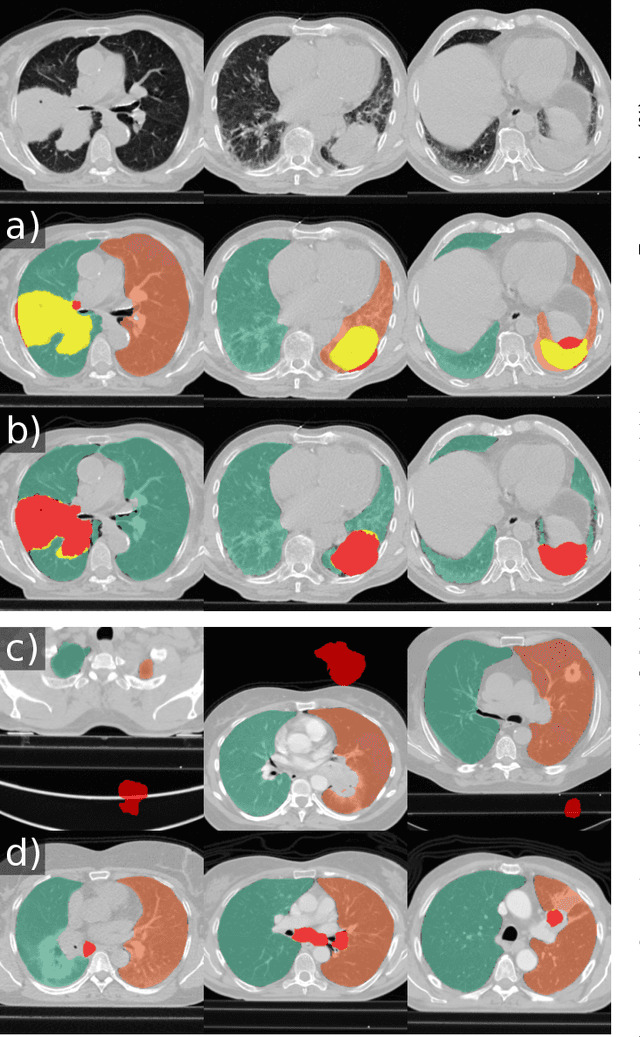
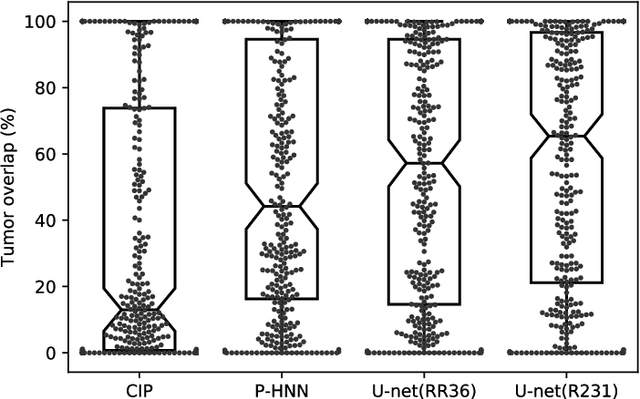
Abstract:Automated segmentation of anatomical structures is a crucial step in many medical image analysis tasks. For lung segmentation, a variety of approaches exist, involving sophisticated pipelines trained and validated on a range of different data sets. However, during translation to clinical routine the applicability of these approaches across diseases remains limited. Here, we show that the accuracy and reliability of lung segmentation algorithms on demanding cases primarily does not depend on methodology, but on the diversity of training data. We compare 4 generic deep learning approaches and 2 published lung segmentation algorithms on routine imaging data with more than 6 different disease patterns and 3 published data sets. We show that a basic approach - U-net - performs either better, or competitively with other approaches on both routine data and published data sets, and outperforms published approaches once trained on a diverse data set covering multiple diseases. Training data composition consistently has a bigger impact than algorithm choice on accuracy across test data sets. We carefully analyse the impact of data diversity, and the specifications of annotations on both training and validation sets to provide a reference for algorithms, training data, and annotation. Results on a seemingly well understood task of lung segmentation suggest the critical importance of training data diversity compared to model choice.
Unsupervised deep clustering for predictive texture pattern discovery in medical images
Jan 31, 2020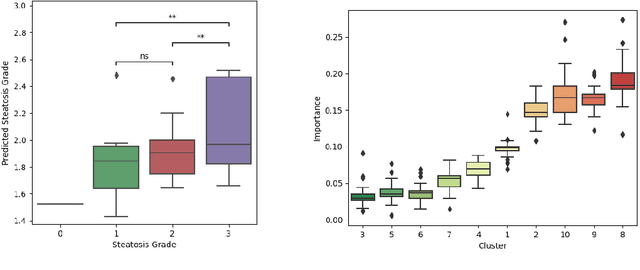
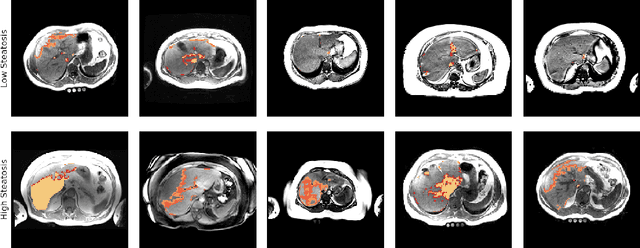
Abstract:Predictive marker patterns in imaging data are a means to quantify disease and progression, but their identification is challenging, if the underlying biology is poorly understood. Here, we present a method to identify predictive texture patterns in medical images in an unsupervised way. Based on deep clustering networks, we simultaneously encode and cluster medical image patches in a low-dimensional latent space. The resulting clusters serve as features for disease staging, linking them to the underlying disease. We evaluate the method on 70 T1-weighted magnetic resonance images of patients with different stages of liver steatosis. The deep clustering approach is able to find predictive clusters with a stable ranking, differentiating between low and high steatosis with an F1-Score of 0.78.
Exploiting Epistemic Uncertainty of Anatomy Segmentation for Anomaly Detection in Retinal OCT
May 29, 2019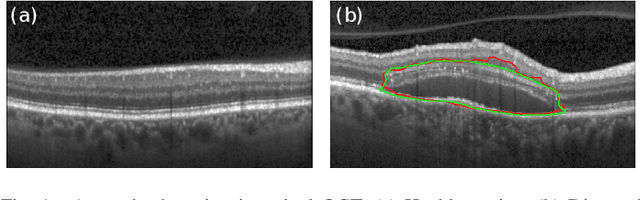
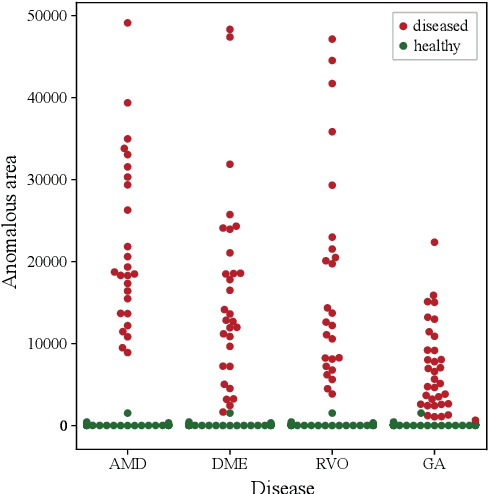
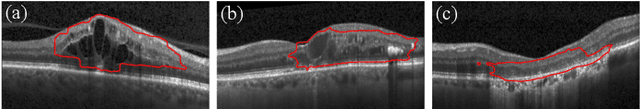
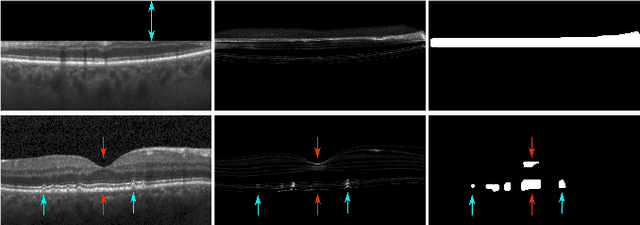
Abstract:Diagnosis and treatment guidance are aided by detecting relevant biomarkers in medical images. Although supervised deep learning can perform accurate segmentation of pathological areas, it is limited by requiring a-priori definitions of these regions, large-scale annotations, and a representative patient cohort in the training set. In contrast, anomaly detection is not limited to specific definitions of pathologies and allows for training on healthy samples without annotation. Anomalous regions can then serve as candidates for biomarker discovery. Knowledge about normal anatomical structure brings implicit information for detecting anomalies. We propose to take advantage of this property using bayesian deep learning, based on the assumption that epistemic uncertainties will correlate with anatomical deviations from a normal training set. A Bayesian U-Net is trained on a well-defined healthy environment using weak labels of healthy anatomy produced by existing methods. At test time, we capture epistemic uncertainty estimates of our model using Monte Carlo dropout. A novel post-processing technique is then applied to exploit these estimates and transfer their layered appearance to smooth blob-shaped segmentations of the anomalies. We experimentally validated this approach in retinal optical coherence tomography (OCT) images, using weak labels of retinal layers. Our method achieved a Dice index of 0.789 in an independent anomaly test set of age-related macular degeneration (AMD) cases. The resulting segmentations allowed very high accuracy for separating healthy and diseased cases with late wet AMD, dry geographic atrophy (GA), diabetic macular edema (DME) and retinal vein occlusion (RVO). Finally, we qualitatively observed that our approach can also detect other deviations in normal scans such as cut edge artifacts.
Using CycleGANs for effectively reducing image variability across OCT devices and improving retinal fluid segmentation
Jan 25, 2019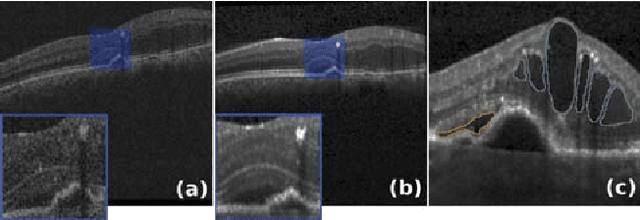
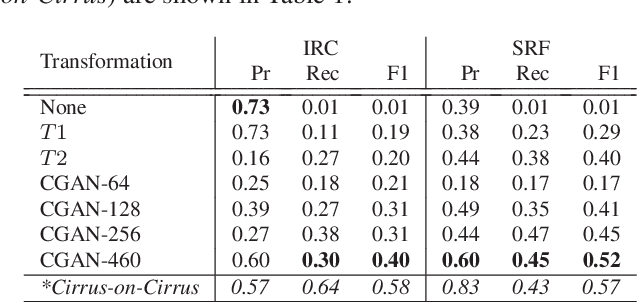
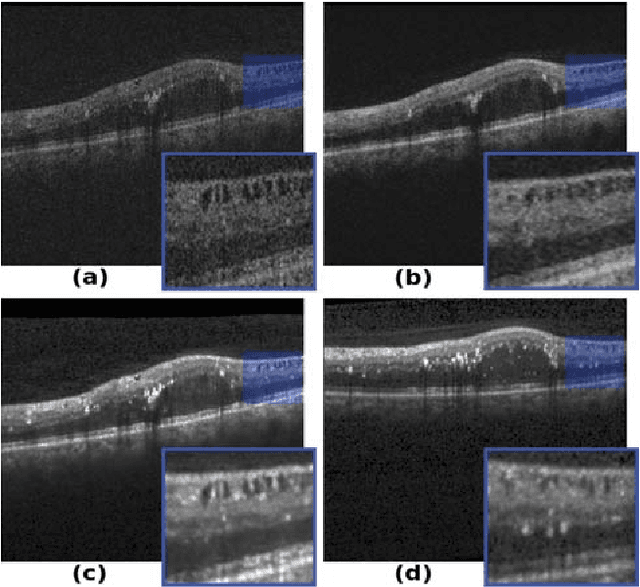
Abstract:Optical coherence tomography (OCT) has become the most important imaging modality in ophthalmology. A substantial amount of research has recently been devoted to the development of machine learning (ML) models for the identification and quantification of pathological features in OCT images. Among the several sources of variability the ML models have to deal with, a major factor is the acquisition device, which can limit the ML model's generalizability. In this paper, we propose to reduce the image variability across different OCT devices (Spectralis and Cirrus) by using CycleGAN, an unsupervised unpaired image transformation algorithm. The usefulness of this approach is evaluated in the setting of retinal fluid segmentation, namely intraretinal cystoid fluid (IRC) and subretinal fluid (SRF). First, we train a segmentation model on images acquired with a source OCT device. Then we evaluate the model on (1) source, (2) target and (3) transformed versions of the target OCT images. The presented transformation strategy shows an F1 score of 0.4 (0.51) for IRC (SRF) segmentations. Compared with traditional transformation approaches, this means an F1 score gain of 0.2 (0.12).
Unsupervised Identification of Disease Marker Candidates in Retinal OCT Imaging Data
Oct 31, 2018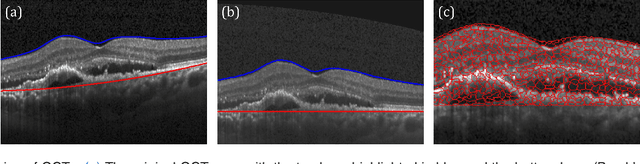
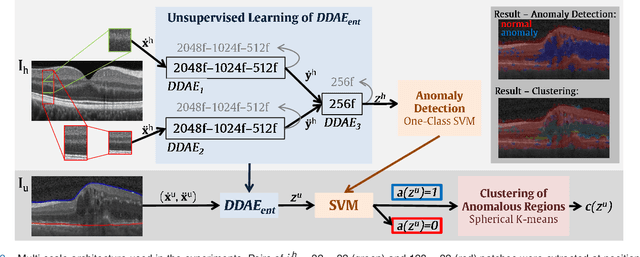
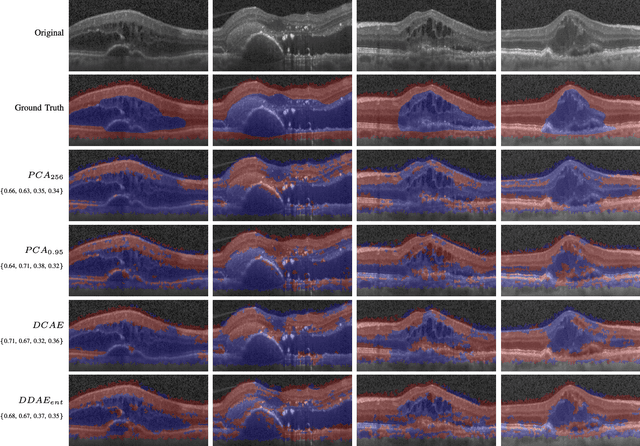
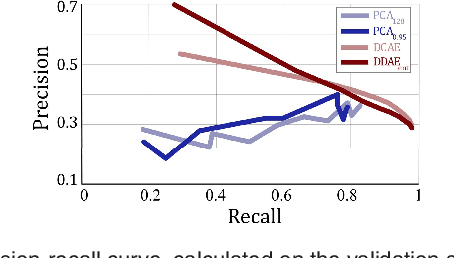
Abstract:The identification and quantification of markers in medical images is critical for diagnosis, prognosis, and disease management. Supervised machine learning enables the detection and exploitation of findings that are known a priori after annotation of training examples by experts. However, supervision does not scale well, due to the amount of necessary training examples, and the limitation of the marker vocabulary to known entities. In this proof-of-concept study, we propose unsupervised identification of anomalies as candidates for markers in retinal Optical Coherence Tomography (OCT) imaging data without a constraint to a priori definitions. We identify and categorize marker candidates occurring frequently in the data, and demonstrate that these markers show predictive value in the task of detecting disease. A careful qualitative analysis of the identified data driven markers reveals how their quantifiable occurrence aligns with our current understanding of disease course, in early- and late age-related macular degeneration (AMD) patients. A multi-scale deep denoising autoencoder is trained on healthy images, and a one-class support vector machine identifies anomalies in new data. Clustering in the anomalies identifies stable categories. Using these markers to classify healthy-, early AMD- and late AMD cases yields an accuracy of 81.40%. In a second binary classification experiment on a publicly available data set (healthy vs. intermediate AMD) the model achieves an area under the ROC curve of 0.944.
Keypoint Transfer for Fast Whole-Body Segmentation
Jun 22, 2018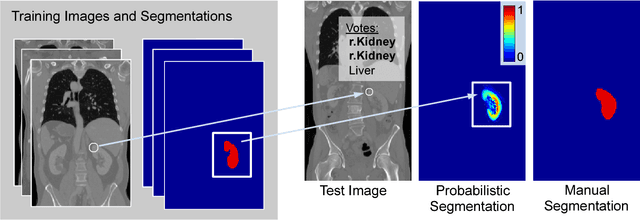
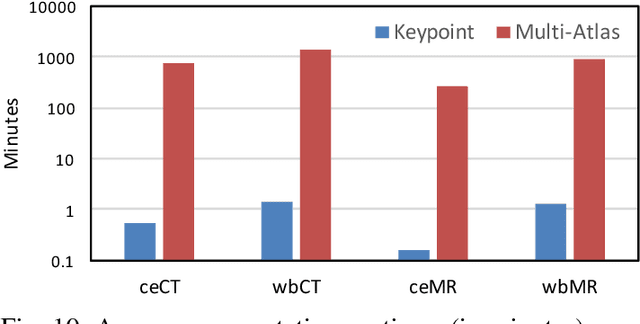
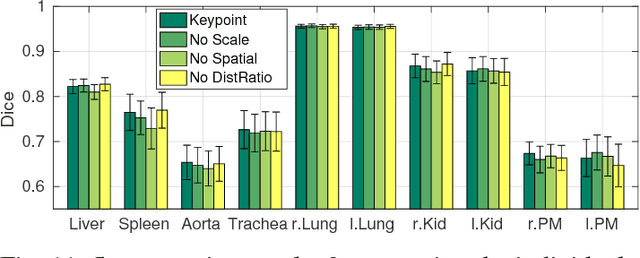
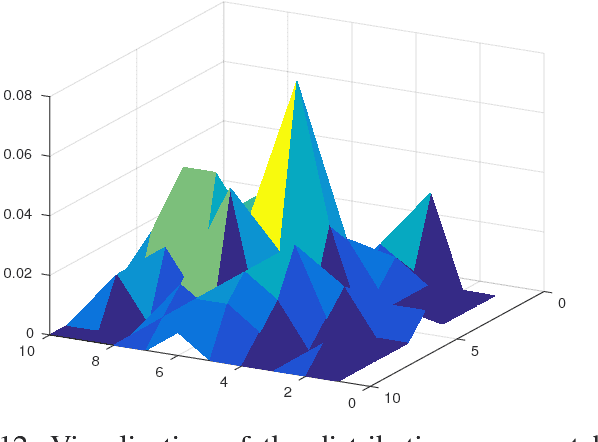
Abstract:We introduce an approach for image segmentation based on sparse correspondences between keypoints in testing and training images. Keypoints represent automatically identified distinctive image locations, where each keypoint correspondence suggests a transformation between images. We use these correspondences to transfer label maps of entire organs from the training images to the test image. The keypoint transfer algorithm includes three steps: (i) keypoint matching, (ii) voting-based keypoint labeling, and (iii) keypoint-based probabilistic transfer of organ segmentations. We report segmentation results for abdominal organs in whole-body CT and MRI, as well as in contrast-enhanced CT and MRI. Our method offers a speed-up of about three orders of magnitude in comparison to common multi-atlas segmentation, while achieving an accuracy that compares favorably. Moreover, keypoint transfer does not require the registration to an atlas or a training phase. Finally, the method allows for the segmentation of scans with highly variable field-of-view.
Fully Automated Segmentation of Hyperreflective Foci in Optical Coherence Tomography Images
May 08, 2018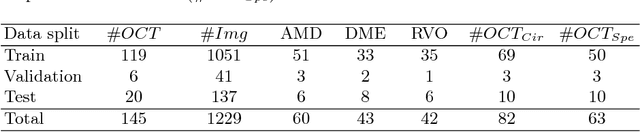
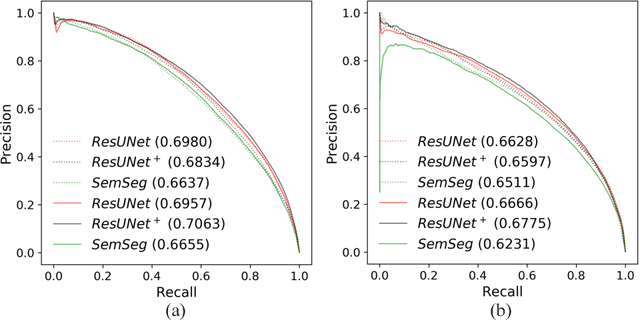
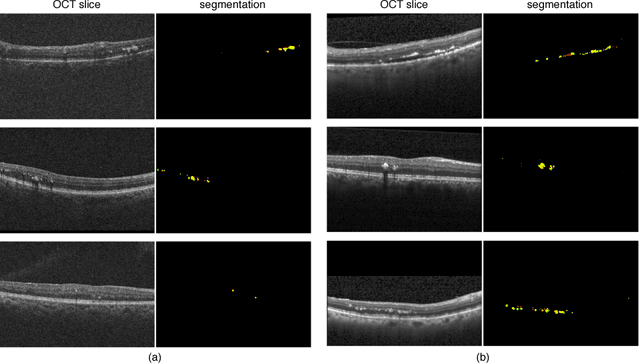
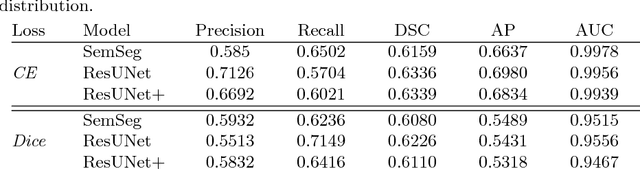
Abstract:The automatic detection of disease related entities in retinal imaging data is relevant for disease- and treatment monitoring. It enables the quantitative assessment of large amounts of data and the corresponding study of disease characteristics. The presence of hyperreflective foci (HRF) is related to disease progression in various retinal diseases. Manual identification of HRF in spectral-domain optical coherence tomography (SD-OCT) scans is error-prone and tedious. We present a fully automated machine learning approach for segmenting HRF in SD-OCT scans. Evaluation on annotated OCT images of the retina demonstrates that a residual U-Net allows to segment HRF with high accuracy. As our dataset comprised data from different retinal diseases including age-related macular degeneration, diabetic macular edema and retinal vein occlusion, the algorithm can safely be applied in all of them though different pathophysiological origins are known.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge