Genevieve B. Melton
Uncertainty-Aware Large Language Models for Explainable Disease Diagnosis
May 06, 2025



Abstract:Explainable disease diagnosis, which leverages patient information (e.g., signs and symptoms) and computational models to generate probable diagnoses and reasonings, offers clear clinical values. However, when clinical notes encompass insufficient evidence for a definite diagnosis, such as the absence of definitive symptoms, diagnostic uncertainty usually arises, increasing the risk of misdiagnosis and adverse outcomes. Although explicitly identifying and explaining diagnostic uncertainties is essential for trustworthy diagnostic systems, it remains under-explored. To fill this gap, we introduce ConfiDx, an uncertainty-aware large language model (LLM) created by fine-tuning open-source LLMs with diagnostic criteria. We formalized the task and assembled richly annotated datasets that capture varying degrees of diagnostic ambiguity. Evaluating ConfiDx on real-world datasets demonstrated that it excelled in identifying diagnostic uncertainties, achieving superior diagnostic performance, and generating trustworthy explanations for diagnoses and uncertainties. To our knowledge, this is the first study to jointly address diagnostic uncertainty recognition and explanation, substantially enhancing the reliability of automatic diagnostic systems.
Interpretable Differential Diagnosis with Dual-Inference Large Language Models
Jul 10, 2024



Abstract:Methodological advancements to automate the generation of differential diagnosis (DDx) to predict a list of potential diseases as differentials given patients' symptom descriptions are critical to clinical reasoning and applications such as decision support. However, providing reasoning or interpretation for these differential diagnoses is more meaningful. Fortunately, large language models (LLMs) possess powerful language processing abilities and have been proven effective in various related tasks. Motivated by this potential, we investigate the use of LLMs for interpretable DDx. First, we develop a new DDx dataset with expert-derived interpretation on 570 public clinical notes. Second, we propose a novel framework, named Dual-Inf, that enables LLMs to conduct bidirectional inference for interpretation. Both human and automated evaluation demonstrate the effectiveness of Dual-Inf in predicting differentials and diagnosis explanations. Specifically, the performance improvement of Dual-Inf over the baseline methods exceeds 32% w.r.t. BERTScore in DDx interpretation. Furthermore, experiments verify that Dual-Inf (1) makes fewer errors in interpretation, (2) has great generalizability, (3) is promising for rare disease diagnosis and explanation.
An Empirical Study of UMLS Concept Extraction from Clinical Notes using Boolean Combination Ensembles
Aug 04, 2021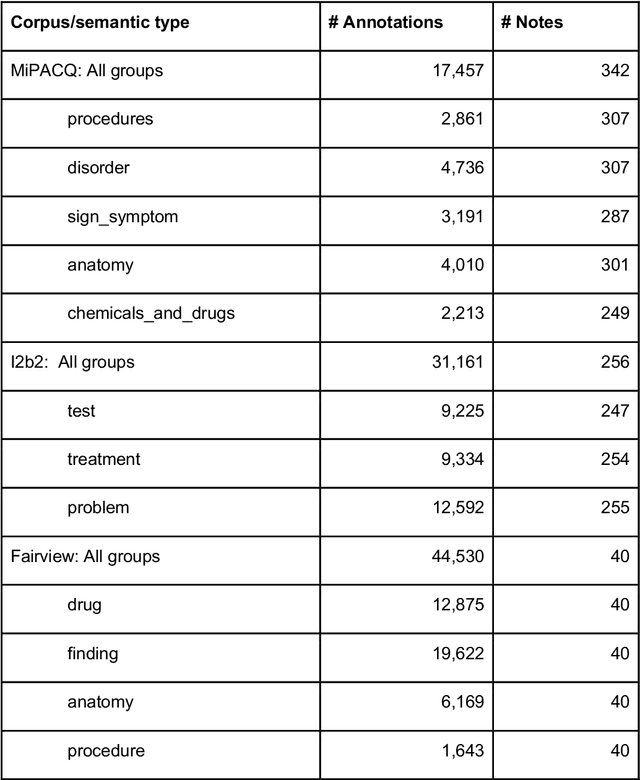
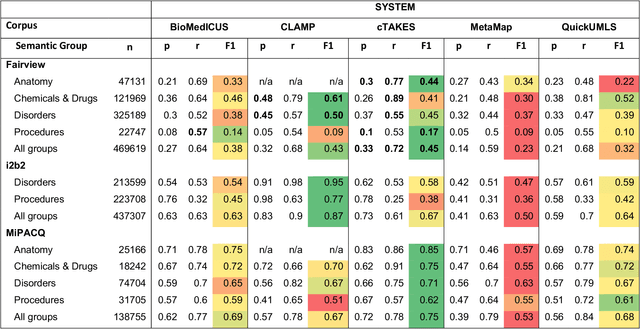
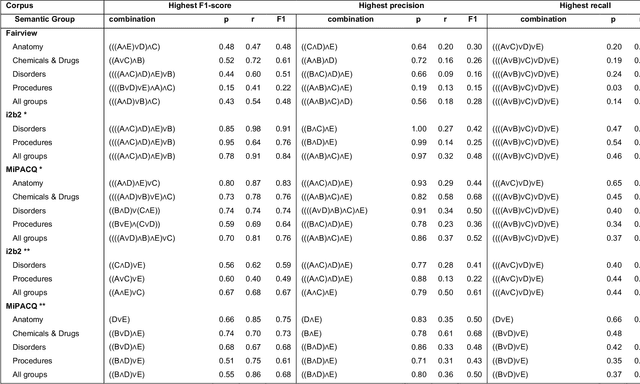
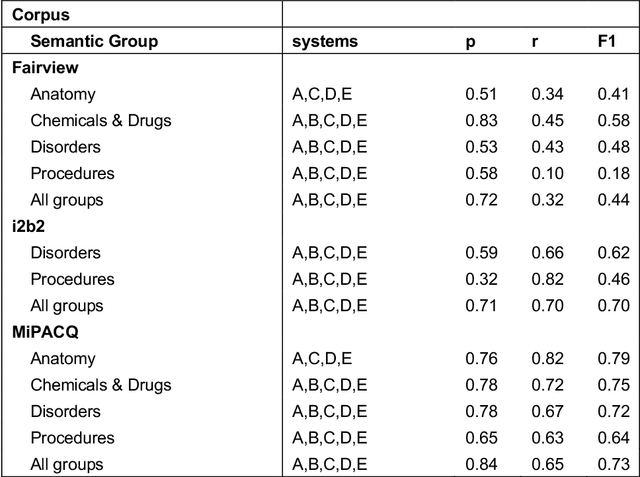
Abstract:Our objective in this study is to investigate the behavior of Boolean operators on combining annotation output from multiple Natural Language Processing (NLP) systems across multiple corpora and to assess how filtering by aggregation of Unified Medical Language System (UMLS) Metathesaurus concepts affects system performance for Named Entity Recognition (NER) of UMLS concepts. We used three corpora annotated for UMLS concepts: 2010 i2b2 VA challenge set (31,161 annotations), Multi-source Integrated Platform for Answering Clinical Questions (MiPACQ) corpus (17,457 annotations including UMLS concept unique identifiers), and Fairview Health Services corpus (44,530 annotations). Our results showed that for UMLS concept matching, Boolean ensembling of the MiPACQ corpus trended towards higher performance over individual systems. Use of an approximate grid-search can help optimize the precision-recall tradeoff and can provide a set of heuristics for choosing an optimal set of ensembles.
A Prospective Observational Study to Investigate Performance of a Chest X-ray Artificial Intelligence Diagnostic Support Tool Across 12 U.S. Hospitals
Jun 07, 2021Abstract:Importance: An artificial intelligence (AI)-based model to predict COVID-19 likelihood from chest x-ray (CXR) findings can serve as an important adjunct to accelerate immediate clinical decision making and improve clinical decision making. Despite significant efforts, many limitations and biases exist in previously developed AI diagnostic models for COVID-19. Utilizing a large set of local and international CXR images, we developed an AI model with high performance on temporal and external validation. Conclusions and Relevance: AI-based diagnostic tools may serve as an adjunct, but not replacement, for clinical decision support of COVID-19 diagnosis, which largely hinges on exposure history, signs, and symptoms. While AI-based tools have not yet reached full diagnostic potential in COVID-19, they may still offer valuable information to clinicians taken into consideration along with clinical signs and symptoms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge