Finale Doshi-Velez
Preferential Mixture-of-Experts: Interpretable Models that Rely on Human Expertise as much as Possible
Jan 13, 2021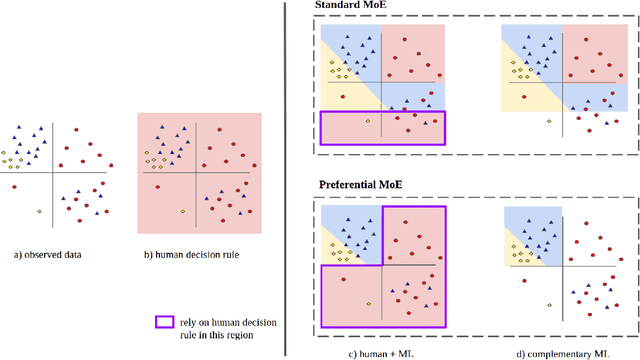
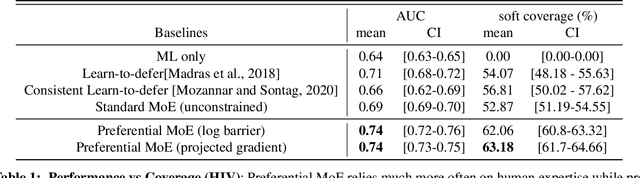
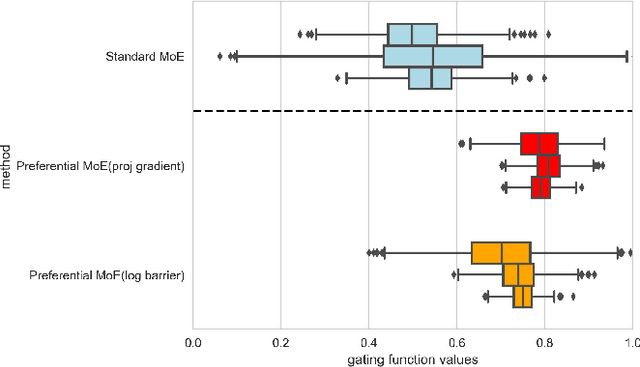
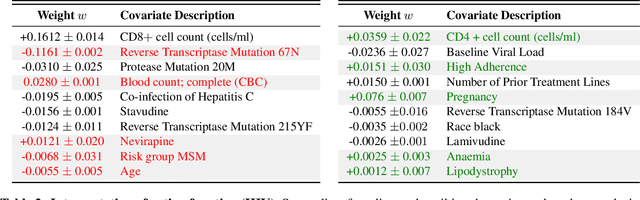
Abstract:We propose Preferential MoE, a novel human-ML mixture-of-experts model that augments human expertise in decision making with a data-based classifier only when necessary for predictive performance. Our model exhibits an interpretable gating function that provides information on when human rules should be followed or avoided. The gating function is maximized for using human-based rules, and classification errors are minimized. We propose solving a coupled multi-objective problem with convex subproblems. We develop approximate algorithms and study their performance and convergence. Finally, we demonstrate the utility of Preferential MoE on two clinical applications for the treatment of Human Immunodeficiency Virus (HIV) and management of Major Depressive Disorder (MDD).
Identifying Decision Points for Safe and Interpretable Reinforcement Learning in Hypotension Treatment
Jan 09, 2021
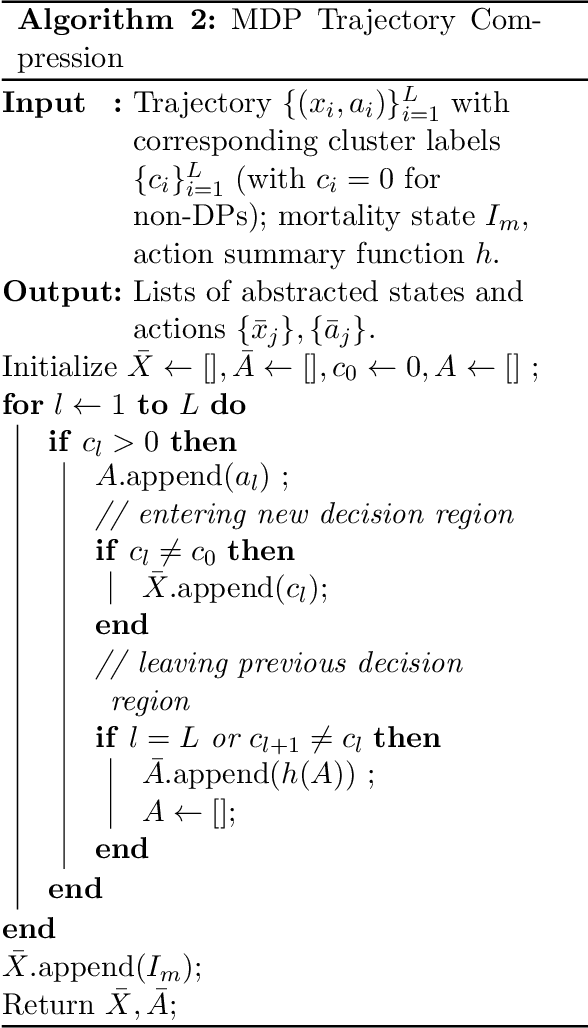
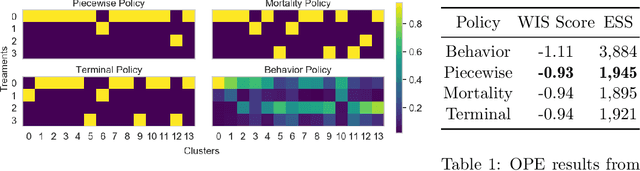

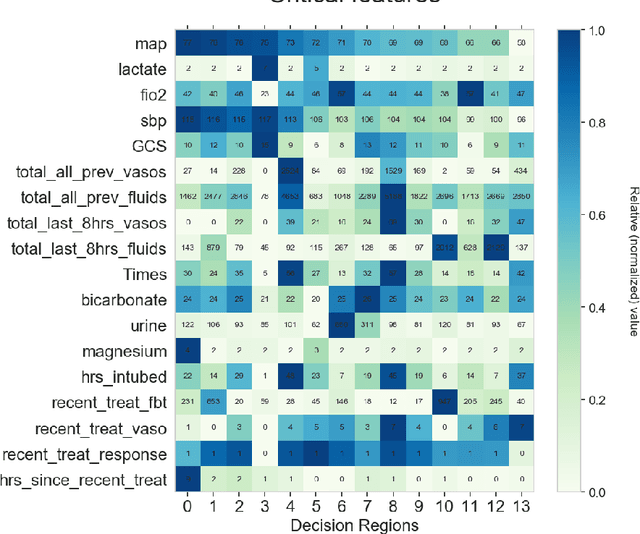
Abstract:Many batch RL health applications first discretize time into fixed intervals. However, this discretization both loses resolution and forces a policy computation at each (potentially fine) interval. In this work, we develop a novel framework to compress continuous trajectories into a few, interpretable decision points --places where the batch data support multiple alternatives. We apply our approach to create recommendations from a cohort of hypotensive patients dataset. Our reduced state space results in faster planning and allows easy inspection by a clinical expert.
Artificial Intelligence & Cooperation
Dec 10, 2020Abstract:The rise of Artificial Intelligence (AI) will bring with it an ever-increasing willingness to cede decision-making to machines. But rather than just giving machines the power to make decisions that affect us, we need ways to work cooperatively with AI systems. There is a vital need for research in "AI and Cooperation" that seeks to understand the ways in which systems of AIs and systems of AIs with people can engender cooperative behavior. Trust in AI is also key: trust that is intrinsic and trust that can only be earned over time. Here we use the term "AI" in its broadest sense, as employed by the recent 20-Year Community Roadmap for AI Research (Gil and Selman, 2019), including but certainly not limited to, recent advances in deep learning. With success, cooperation between humans and AIs can build society just as human-human cooperation has. Whether coming from an intrinsic willingness to be helpful, or driven through self-interest, human societies have grown strong and the human species has found success through cooperation. We cooperate "in the small" -- as family units, with neighbors, with co-workers, with strangers -- and "in the large" as a global community that seeks cooperative outcomes around questions of commerce, climate change, and disarmament. Cooperation has evolved in nature also, in cells and among animals. While many cases involving cooperation between humans and AIs will be asymmetric, with the human ultimately in control, AI systems are growing so complex that, even today, it is impossible for the human to fully comprehend their reasoning, recommendations, and actions when functioning simply as passive observers.
Learning Interpretable Concept-Based Models with Human Feedback
Dec 04, 2020
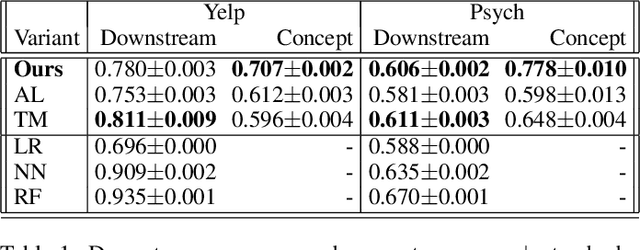
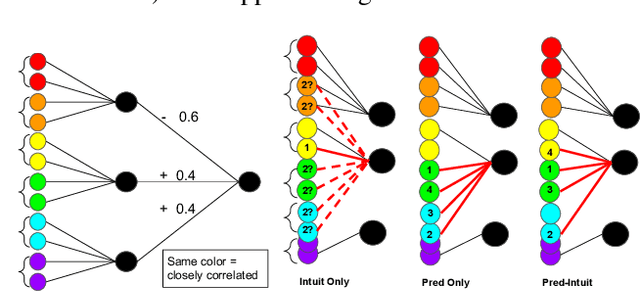
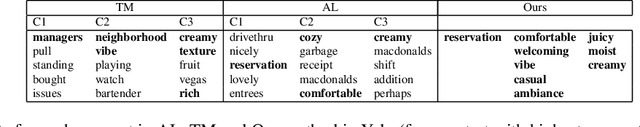
Abstract:Machine learning models that first learn a representation of a domain in terms of human-understandable concepts, then use it to make predictions, have been proposed to facilitate interpretation and interaction with models trained on high-dimensional data. However these methods have important limitations: the way they define concepts are not inherently interpretable, and they assume that concept labels either exist for individual instances or can easily be acquired from users. These limitations are particularly acute for high-dimensional tabular features. We propose an approach for learning a set of transparent concept definitions in high-dimensional tabular data that relies on users labeling concept features instead of individual instances. Our method produces concepts that both align with users' intuitive sense of what a concept means, and facilitate prediction of the downstream label by a transparent machine learning model. This ensures that the full model is transparent and intuitive, and as predictive as possible given this constraint. We demonstrate with simulated user feedback on real prediction problems, including one in a clinical domain, that this kind of direct feedback is much more efficient at learning solutions that align with ground truth concept definitions than alternative transparent approaches that rely on labeling instances or other existing interaction mechanisms, while maintaining similar predictive performance.
Incorporating Interpretable Output Constraints in Bayesian Neural Networks
Oct 21, 2020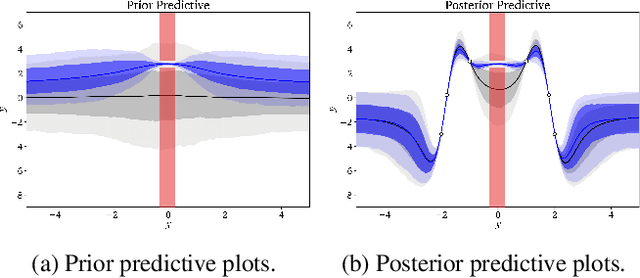
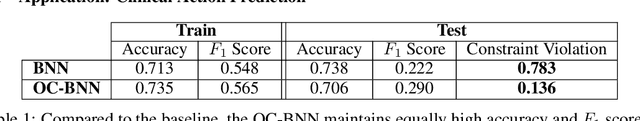
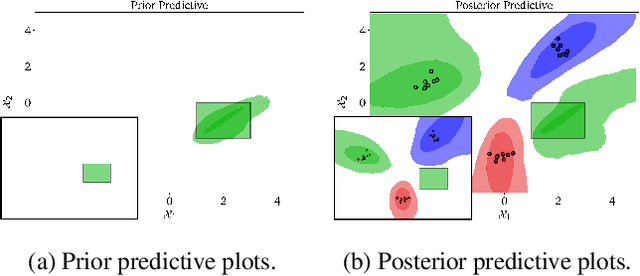
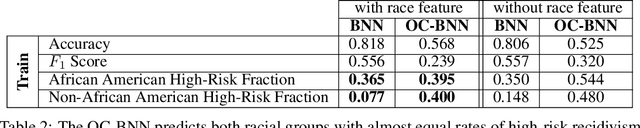
Abstract:Domains where supervised models are deployed often come with task-specific constraints, such as prior expert knowledge on the ground-truth function, or desiderata like safety and fairness. We introduce a novel probabilistic framework for reasoning with such constraints and formulate a prior that enables us to effectively incorporate them into Bayesian neural networks (BNNs), including a variant that can be amortized over tasks. The resulting Output-Constrained BNN (OC-BNN) is fully consistent with the Bayesian framework for uncertainty quantification and is amenable to black-box inference. Unlike typical BNN inference in uninterpretable parameter space, OC-BNNs widen the range of functional knowledge that can be incorporated, especially for model users without expertise in machine learning. We demonstrate the efficacy of OC-BNNs on real-world datasets, spanning multiple domains such as healthcare, criminal justice, and credit scoring.
Failure Modes of Variational Autoencoders and Their Effects on Downstream Tasks
Jul 14, 2020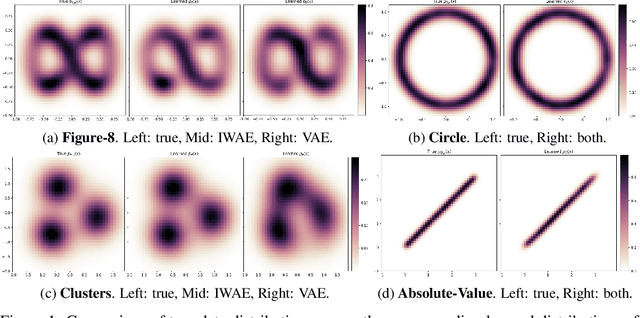
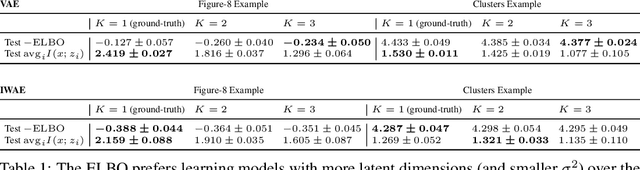
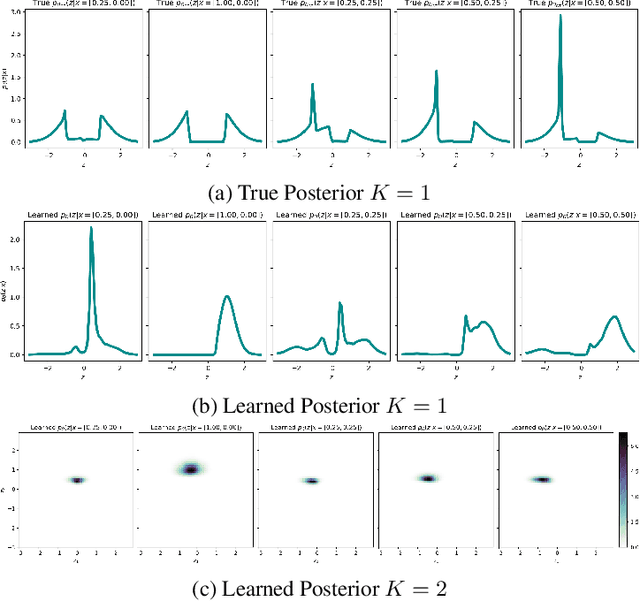
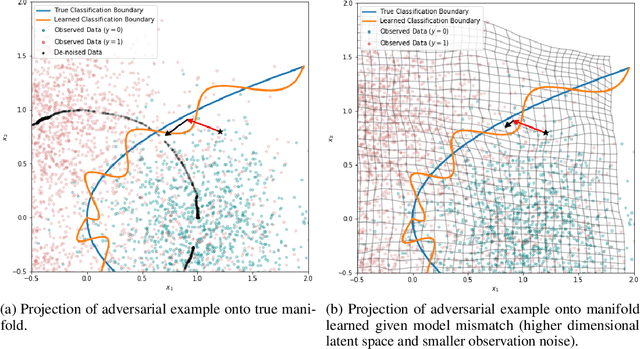
Abstract:Variational Auto-encoders (VAEs) are deep generative latent variable models that are widely used for a number of downstream tasks. While it has been demonstrated that VAE training can suffer from a number of pathologies, existing literature lacks characterizations of exactly when these pathologies occur and how they impact down-stream task performance. In this paper we concretely characterize conditions under which VAE training exhibits pathologies and connect these failure modes to undesirable effects on specific downstream tasks - learning compressed and disentangled representations, adversarial robustness and semi-supervised learning.
BaCOUn: Bayesian Classifers with Out-of-Distribution Uncertainty
Jul 12, 2020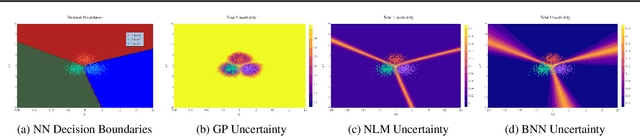
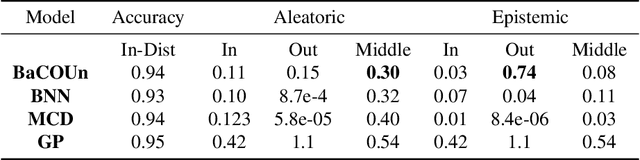
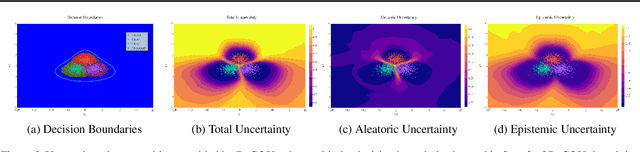
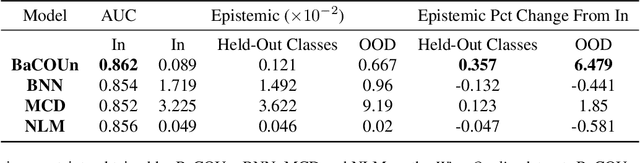
Abstract:Traditional training of deep classifiers yields overconfident models that are not reliable under dataset shift. We propose a Bayesian framework to obtain reliable uncertainty estimates for deep classifiers. Our approach consists of a plug-in "generator" used to augment the data with an additional class of points that lie on the boundary of the training data, followed by Bayesian inference on top of features that are trained to distinguish these "out-of-distribution" points.
Learned Uncertainty-Aware (LUNA) Bases for Bayesian Regression using Multi-Headed Auxiliary Networks
Jul 08, 2020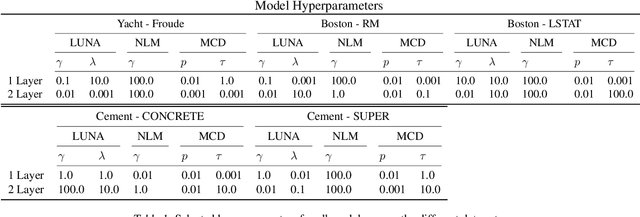
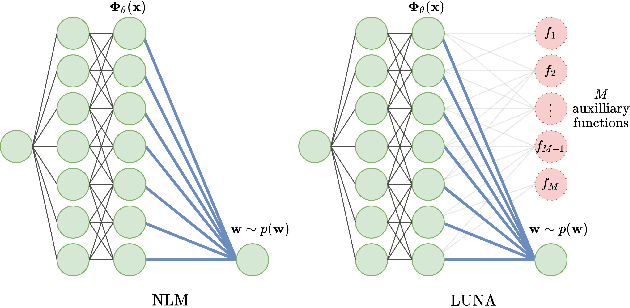
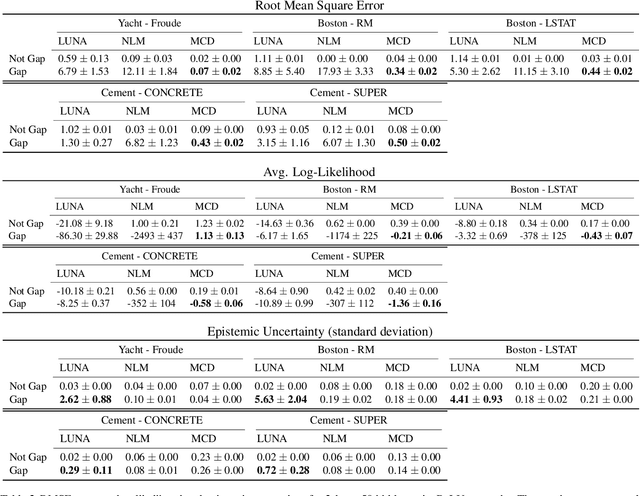
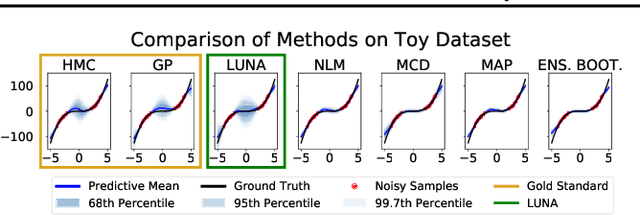
Abstract:Neural Linear Models (NLM) are deep models that produce predictive uncertainty by learning features from the data and then performing Bayesian linear regression over these features. Despite their popularity, few works have focused on formally evaluating the predictive uncertainties of these models. In this work, we show that traditional training procedures for NLMs can drastically underestimate uncertainty in data-scarce regions. We identify the underlying reasons for this behavior and propose a novel training procedure for capturing useful predictive uncertainties.
Model-based Reinforcement Learning for Semi-Markov Decision Processes with Neural ODEs
Jun 29, 2020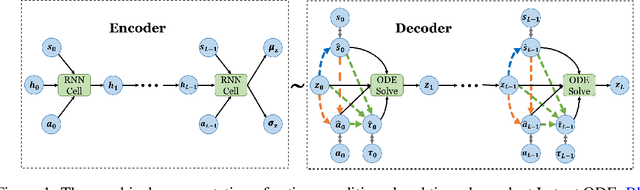

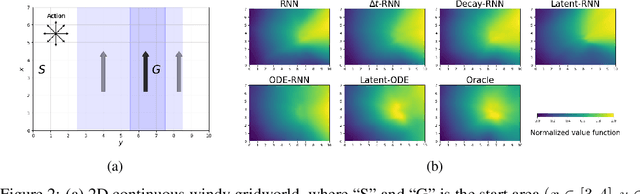
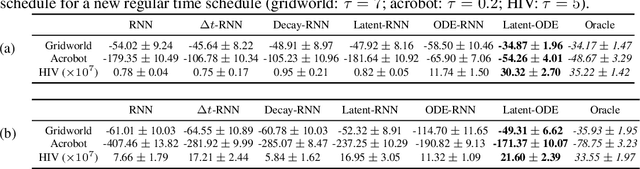
Abstract:We present two elegant solutions for modeling continuous-time dynamics, in a novel model-based reinforcement learning (RL) framework for semi-Markov decision processes (SMDPs), using neural ordinary differential equations (ODEs). Our models accurately characterize continuous-time dynamics and enable us to develop high-performing policies using a small amount of data. We also develop a model-based approach for optimizing time schedules to reduce interaction rates with the environment while maintaining the near-optimal performance, which is not possible for model-free methods. We experimentally demonstrate the efficacy of our methods across various continuous-time domains.
PAC Bounds for Imitation and Model-based Batch Learning of Contextual Markov Decision Processes
Jun 11, 2020Abstract:We consider the problem of batch multi-task reinforcement learning with observed context descriptors, motivated by its application to personalized medical treatment. In particular, we study two general classes of learning algorithms: direct policy learning (DPL), an imitation-learning based approach which learns from expert trajectories, and model-based learning. First, we derive sample complexity bounds for DPL, and then show that model-based learning from expert actions can, even with a finite model class, be impossible. After relaxing the conditions under which the model-based approach is expected to learn by allowing for greater coverage of state-action space, we provide sample complexity bounds for model-based learning with finite model classes, showing that there exist model classes with sample complexity exponential in their statistical complexity. We then derive a sample complexity upper bound for model-based learning based on a measure of concentration of the data distribution. Our results give formal justification for imitation learning over model-based learning in this setting.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge