Elisa Ficarra
Handling Missing Modalities in Multimodal Survival Prediction for Non-Small Cell Lung Cancer
Jan 15, 2026Abstract:Accurate survival prediction in Non-Small Cell Lung Cancer (NSCLC) requires the integration of heterogeneous clinical, radiological, and histopathological information. While Multimodal Deep Learning (MDL) offers a promises for precision prognosis and survival prediction, its clinical applicability is severely limited by small cohort sizes and the presence of missing modalities, often forcing complete-case filtering or aggressive imputation. In this work, we present a missing-aware multimodal survival framework that integrates Computed Tomography (CT), Whole-Slide Histopathology (WSI) Images, and structured clinical variables for overall survival modeling in unresectable stage II-III NSCLC. By leveraging Foundation Models (FM) for modality-specific feature extraction and a missing-aware encoding strategy, the proposed approach enables intermediate multimodal fusion under naturally incomplete modality profiles. The proposed architecture is resilient to missing modalities by design, allowing the model to utilize all available data without being forced to drop patients during training or inference. Experimental results demonstrate that intermediate fusion consistently outperforms unimodal baselines as well as early and late fusion strategies, with the strongest performance achieved by the fusion of WSI and clinical modalities (73.30 C-index). Further analyses of modality importance reveal an adaptive behavior in which less informative modalities, i.e., CT modality, are automatically down-weighted and contribute less to the final survival prediction.
Update Your Transformer to the Latest Release: Re-Basin of Task Vectors
May 28, 2025Abstract:Foundation models serve as the backbone for numerous specialized models developed through fine-tuning. However, when the underlying pretrained model is updated or retrained (e.g., on larger and more curated datasets), the fine-tuned model becomes obsolete, losing its utility and requiring retraining. This raises the question: is it possible to transfer fine-tuning to a new release of the model? In this work, we investigate how to transfer fine-tuning to a new checkpoint without having to re-train, in a data-free manner. To do so, we draw principles from model re-basin and provide a recipe based on weight permutations to re-base the modifications made to the original base model, often called task vector. In particular, our approach tailors model re-basin for Transformer models, taking into account the challenges of residual connections and multi-head attention layers. Specifically, we propose a two-level method rooted in spectral theory, initially permuting the attention heads and subsequently adjusting parameters within select pairs of heads. Through extensive experiments on visual and textual tasks, we achieve the seamless transfer of fine-tuned knowledge to new pre-trained backbones without relying on a single training step or datapoint. Code is available at https://github.com/aimagelab/TransFusion.
CUBIC: Concept Embeddings for Unsupervised Bias Identification using VLMs
May 16, 2025Abstract:Deep vision models often rely on biases learned from spurious correlations in datasets. To identify these biases, methods that interpret high-level, human-understandable concepts are more effective than those relying primarily on low-level features like heatmaps. A major challenge for these concept-based methods is the lack of image annotations indicating potentially bias-inducing concepts, since creating such annotations requires detailed labeling for each dataset and concept, which is highly labor-intensive. We present CUBIC (Concept embeddings for Unsupervised Bias IdentifiCation), a novel method that automatically discovers interpretable concepts that may bias classifier behavior. Unlike existing approaches, CUBIC does not rely on predefined bias candidates or examples of model failures tied to specific biases, as such information is not always available. Instead, it leverages image-text latent space and linear classifier probes to examine how the latent representation of a superclass label$\unicode{x2014}$shared by all instances in the dataset$\unicode{x2014}$is influenced by the presence of a given concept. By measuring these shifts against the normal vector to the classifier's decision boundary, CUBIC identifies concepts that significantly influence model predictions. Our experiments demonstrate that CUBIC effectively uncovers previously unknown biases using Vision-Language Models (VLMs) without requiring the samples in the dataset where the classifier underperforms or prior knowledge of potential biases.
Taming Mambas for Voxel Level 3D Medical Image Segmentation
Oct 20, 2024



Abstract:Recently, the field of 3D medical segmentation has been dominated by deep learning models employing Convolutional Neural Networks (CNNs) and Transformer-based architectures, each with their distinctive strengths and limitations. CNNs are constrained by a local receptive field, whereas transformers are hindered by their substantial memory requirements as well as they data hungriness, making them not ideal for processing 3D medical volumes at a fine-grained level. For these reasons, fully convolutional neural networks, as nnUNet, still dominate the scene when segmenting medical structures in 3D large medical volumes. Despite numerous advancements towards developing transformer variants with subquadratic time and memory complexity, these models still fall short in content-based reasoning. A recent breakthrough is Mamba, a Recurrent Neural Network (RNN) based on State Space Models (SSMs) outperforming Transformers in many long-context tasks (million-length sequences) on famous natural language processing and genomic benchmarks while keeping a linear complexity.
Neuro Symbolic Continual Learning: Knowledge, Reasoning Shortcuts and Concept Rehearsal
Feb 02, 2023



Abstract:We introduce Neuro-Symbolic Continual Learning, where a model has to solve a sequence of neuro-symbolic tasks, that is, it has to map sub-symbolic inputs to high-level concepts and compute predictions by reasoning consistently with prior knowledge. Our key observation is that neuro-symbolic tasks, although different, often share concepts whose semantics remains stable over time. Traditional approaches fall short: existing continual strategies ignore knowledge altogether, while stock neuro-symbolic architectures suffer from catastrophic forgetting. We show that leveraging prior knowledge by combining neuro-symbolic architectures with continual strategies does help avoid catastrophic forgetting, but also that doing so can yield models affected by reasoning shortcuts. These undermine the semantics of the acquired concepts, even when detailed prior knowledge is provided upfront and inference is exact, and in turn continual performance. To overcome these issues, we introduce COOL, a COncept-level cOntinual Learning strategy tailored for neuro-symbolic continual problems that acquires high-quality concepts and remembers them over time. Our experiments on three novel benchmarks highlights how COOL attains sustained high performance on neuro-symbolic continual learning tasks in which other strategies fail.
Exploiting generative self-supervised learning for the assessment of biological images with lack of annotations: a COVID-19 case-study
Jul 26, 2021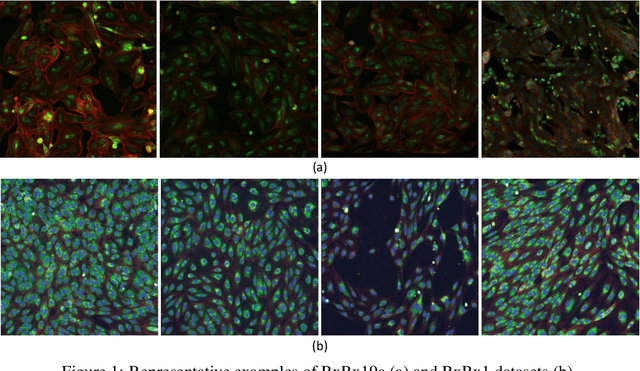

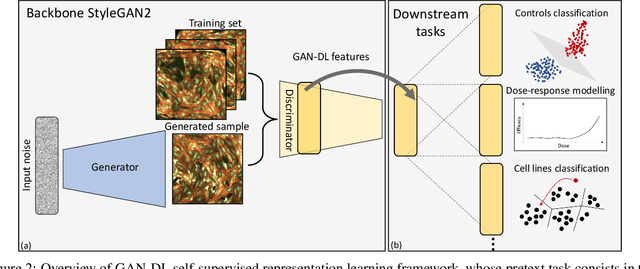
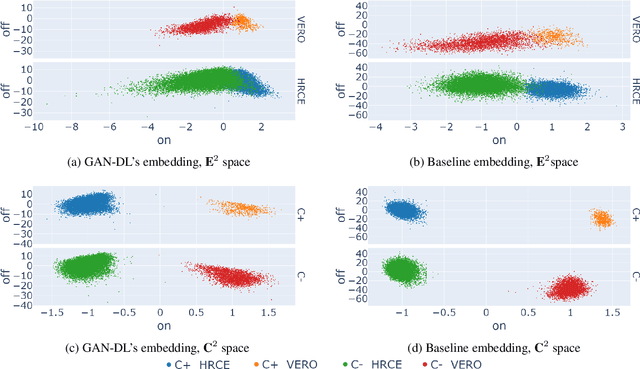
Abstract:Computer-aided analysis of biological images typically requires extensive training on large-scale annotated datasets, which is not viable in many situations. In this paper we present GAN-DL, a Discriminator Learner based on the StyleGAN2 architecture, which we employ for self-supervised image representation learning in the case of fluorescent biological images. We show that Wasserstein Generative Adversarial Networks combined with linear Support Vector Machines enable high-throughput compound screening based on raw images. We demonstrate this by classifying active and inactive compounds tested for the inhibition of SARS-CoV-2 infection in VERO and HRCE cell lines. In contrast to previous methods, our deep learning based approach does not require any annotation besides the one that is normally collected during the sample preparation process. We test our technique on the RxRx19a Sars-CoV-2 image collection. The dataset consists of fluorescent images that were generated to assess the ability of regulatory-approved or in late-stage clinical trials compound to modulate the in vitro infection from SARS-CoV-2 in both VERO and HRCE cell lines. We show that our technique can be exploited not only for classification tasks, but also to effectively derive a dose response curve for the tested treatments, in a self-supervised manner. Lastly, we demonstrate its generalization capabilities by successfully addressing a zero-shot learning task, consisting in the categorization of four different cell types of the RxRx1 fluorescent images collection.
W2WNet: a two-module probabilistic Convolutional Neural Network with embedded data cleansing functionality
Mar 24, 2021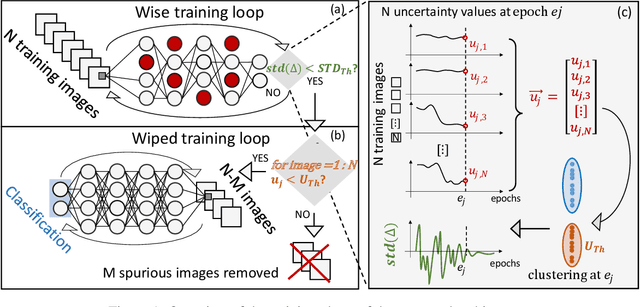
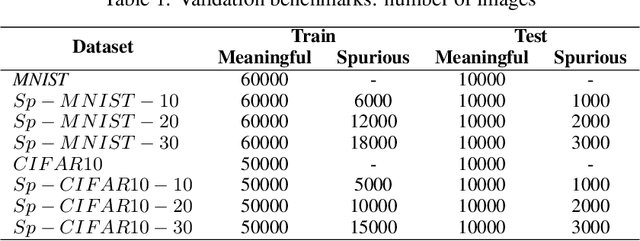
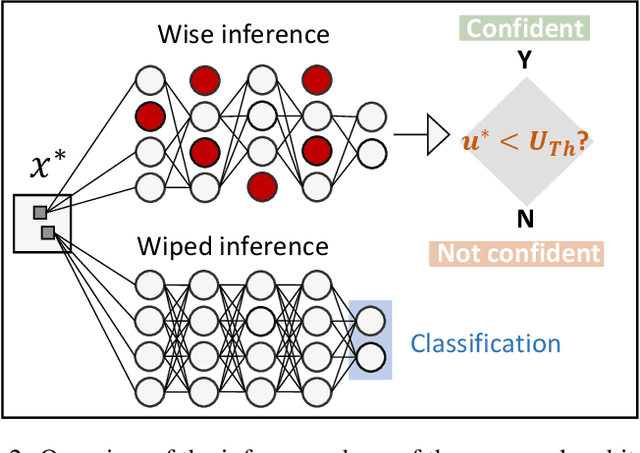
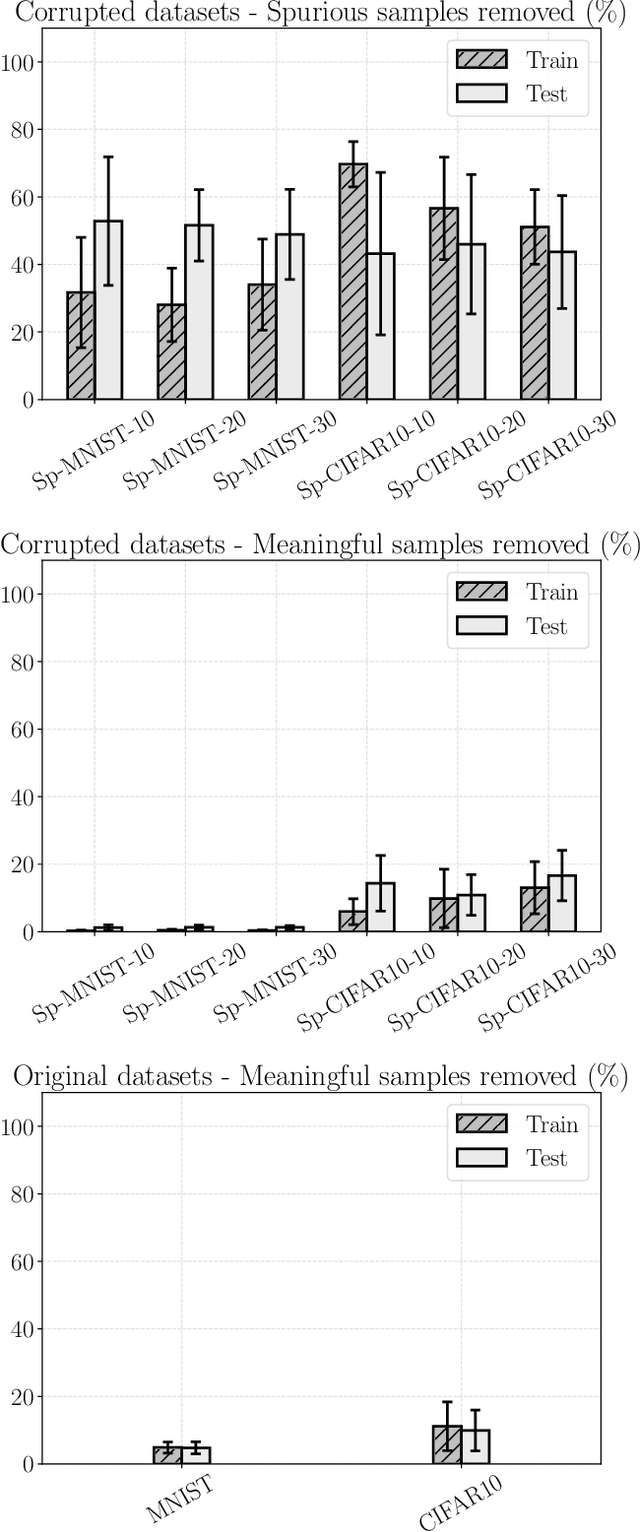
Abstract:Convolutional Neural Networks (CNNs) are supposed to be fed with only high-quality annotated datasets. Nonetheless, in many real-world scenarios, such high quality is very hard to obtain, and datasets may be affected by any sort of image degradation and mislabelling issues. This negatively impacts the performance of standard CNNs, both during the training and the inference phase. To address this issue we propose Wise2WipedNet (W2WNet), a new two-module Convolutional Neural Network, where a Wise module exploits Bayesian inference to identify and discard spurious images during the training, and a Wiped module takes care of the final classification while broadcasting information on the prediction confidence at inference time. The goodness of our solution is demonstrated on a number of public benchmarks addressing different image classification tasks, as well as on a real-world case study on histological image analysis. Overall, our experiments demonstrate that W2WNet is able to identify image degradation and mislabelling issues both at training and at inference time, with a positive impact on the final classification accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge