Dustin Tran
Dima
Edward: A library for probabilistic modeling, inference, and criticism
Feb 01, 2017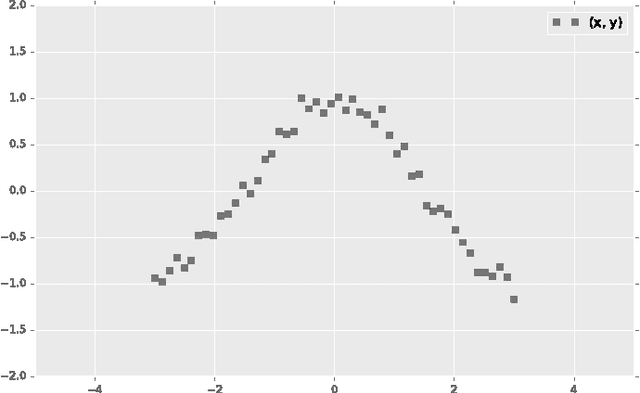
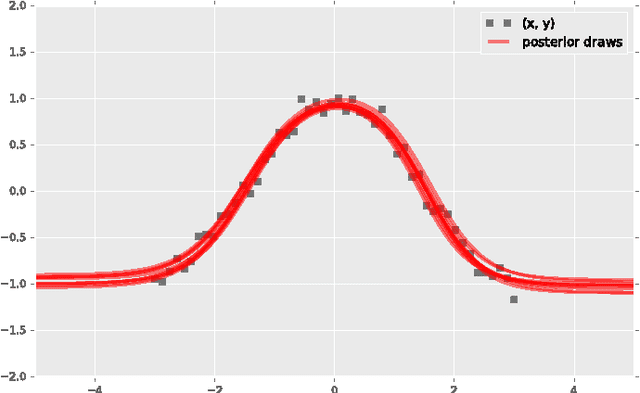
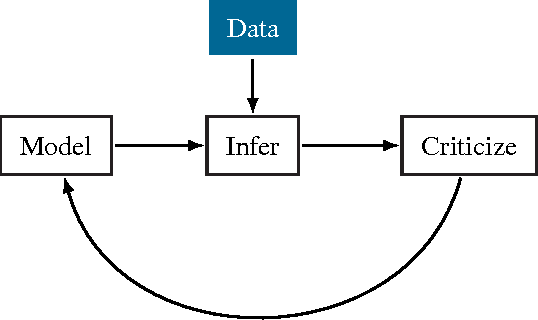
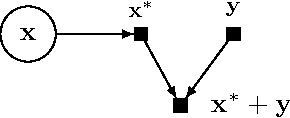
Abstract:Probabilistic modeling is a powerful approach for analyzing empirical information. We describe Edward, a library for probabilistic modeling. Edward's design reflects an iterative process pioneered by George Box: build a model of a phenomenon, make inferences about the model given data, and criticize the model's fit to the data. Edward supports a broad class of probabilistic models, efficient algorithms for inference, and many techniques for model criticism. The library builds on top of TensorFlow to support distributed training and hardware such as GPUs. Edward enables the development of complex probabilistic models and their algorithms at a massive scale.
Towards stability and optimality in stochastic gradient descent
Jun 07, 2016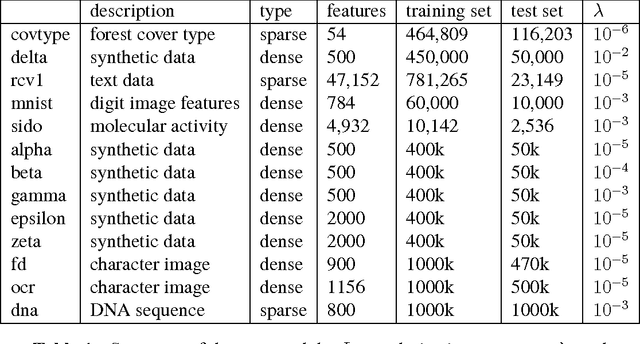
Abstract:Iterative procedures for parameter estimation based on stochastic gradient descent allow the estimation to scale to massive data sets. However, in both theory and practice, they suffer from numerical instability. Moreover, they are statistically inefficient as estimators of the true parameter value. To address these two issues, we propose a new iterative procedure termed averaged implicit SGD (AI-SGD). For statistical efficiency, AI-SGD employs averaging of the iterates, which achieves the optimal Cram\'{e}r-Rao bound under strong convexity, i.e., it is an optimal unbiased estimator of the true parameter value. For numerical stability, AI-SGD employs an implicit update at each iteration, which is related to proximal operators in optimization. In practice, AI-SGD achieves competitive performance with other state-of-the-art procedures. Furthermore, it is more stable than averaging procedures that do not employ proximal updates, and is simple to implement as it requires fewer tunable hyperparameters than procedures that do employ proximal updates.
Hierarchical Variational Models
May 30, 2016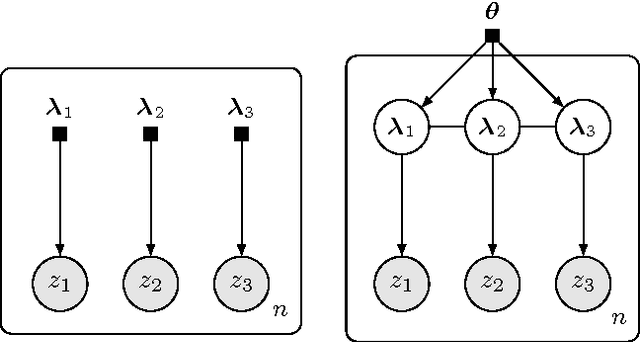
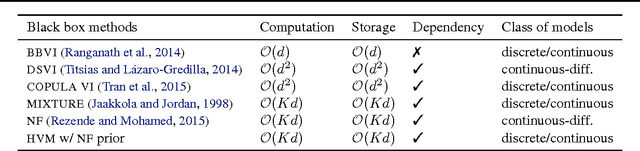

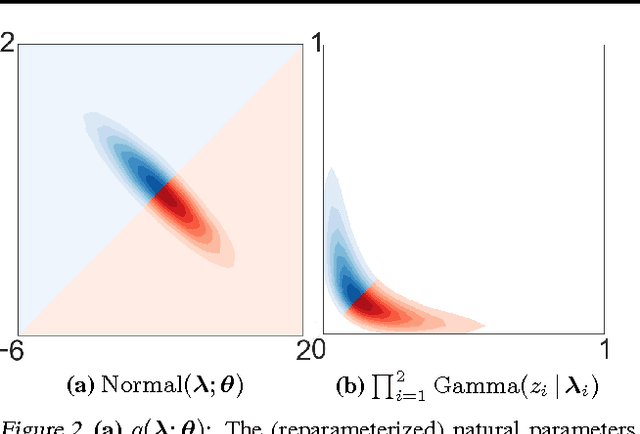
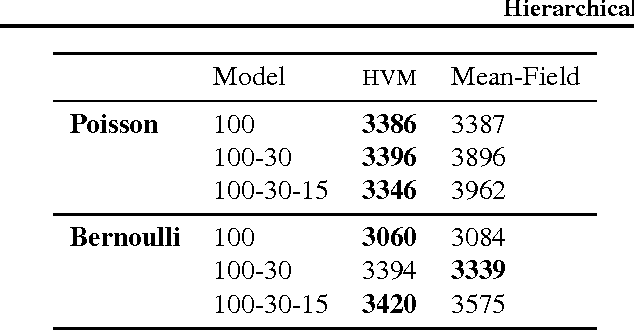
Abstract:Black box variational inference allows researchers to easily prototype and evaluate an array of models. Recent advances allow such algorithms to scale to high dimensions. However, a central question remains: How to specify an expressive variational distribution that maintains efficient computation? To address this, we develop hierarchical variational models (HVMs). HVMs augment a variational approximation with a prior on its parameters, which allows it to capture complex structure for both discrete and continuous latent variables. The algorithm we develop is black box, can be used for any HVM, and has the same computational efficiency as the original approximation. We study HVMs on a variety of deep discrete latent variable models. HVMs generalize other expressive variational distributions and maintains higher fidelity to the posterior.
The Variational Gaussian Process
Apr 17, 2016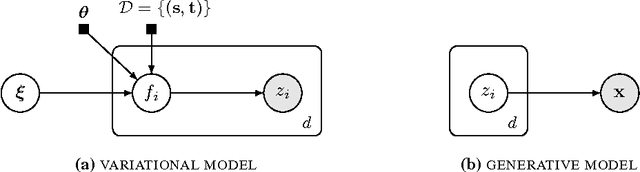
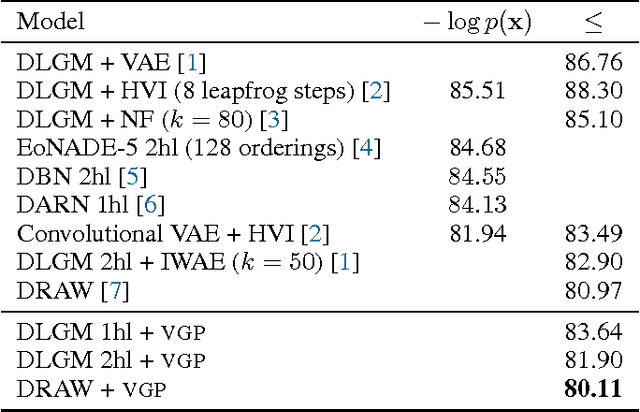

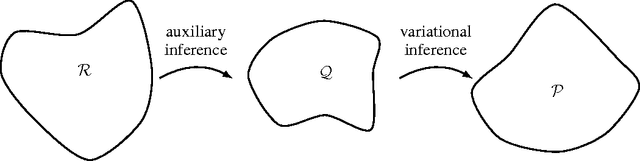
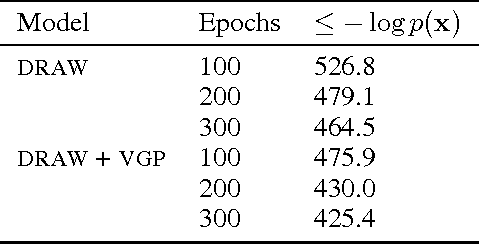
Abstract:Variational inference is a powerful tool for approximate inference, and it has been recently applied for representation learning with deep generative models. We develop the variational Gaussian process (VGP), a Bayesian nonparametric variational family, which adapts its shape to match complex posterior distributions. The VGP generates approximate posterior samples by generating latent inputs and warping them through random non-linear mappings; the distribution over random mappings is learned during inference, enabling the transformed outputs to adapt to varying complexity. We prove a universal approximation theorem for the VGP, demonstrating its representative power for learning any model. For inference we present a variational objective inspired by auto-encoders and perform black box inference over a wide class of models. The VGP achieves new state-of-the-art results for unsupervised learning, inferring models such as the deep latent Gaussian model and the recently proposed DRAW.
Spectral M-estimation with Applications to Hidden Markov Models
Mar 29, 2016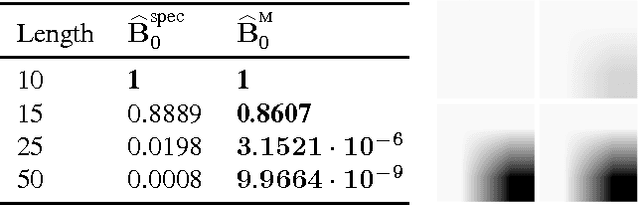

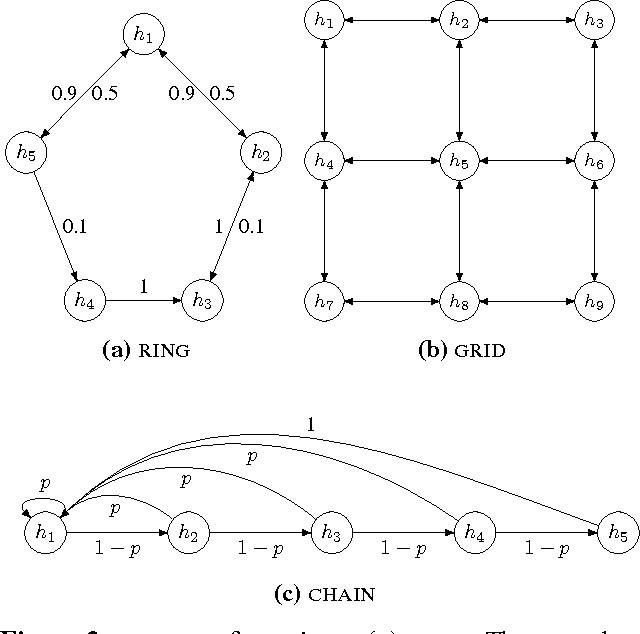

Abstract:Method of moment estimators exhibit appealing statistical properties, such as asymptotic unbiasedness, for nonconvex problems. However, they typically require a large number of samples and are extremely sensitive to model misspecification. In this paper, we apply the framework of M-estimation to develop both a generalized method of moments procedure and a principled method for regularization. Our proposed M-estimator obtains optimal sample efficiency rates (in the class of moment-based estimators) and the same well-known rates on prediction accuracy as other spectral estimators. It also makes it straightforward to incorporate regularization into the sample moment conditions. We demonstrate empirically the gains in sample efficiency from our approach on hidden Markov models.
Automatic Differentiation Variational Inference
Mar 02, 2016

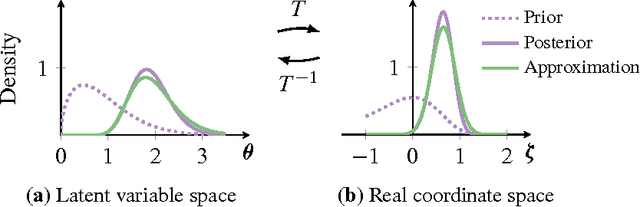

Abstract:Probabilistic modeling is iterative. A scientist posits a simple model, fits it to her data, refines it according to her analysis, and repeats. However, fitting complex models to large data is a bottleneck in this process. Deriving algorithms for new models can be both mathematically and computationally challenging, which makes it difficult to efficiently cycle through the steps. To this end, we develop automatic differentiation variational inference (ADVI). Using our method, the scientist only provides a probabilistic model and a dataset, nothing else. ADVI automatically derives an efficient variational inference algorithm, freeing the scientist to refine and explore many models. ADVI supports a broad class of models-no conjugacy assumptions are required. We study ADVI across ten different models and apply it to a dataset with millions of observations. ADVI is integrated into Stan, a probabilistic programming system; it is available for immediate use.
Copula variational inference
Oct 31, 2015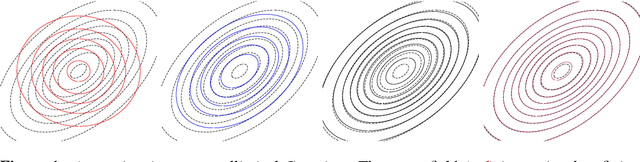
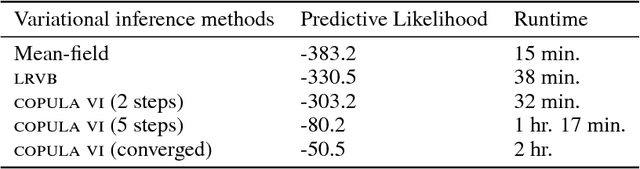
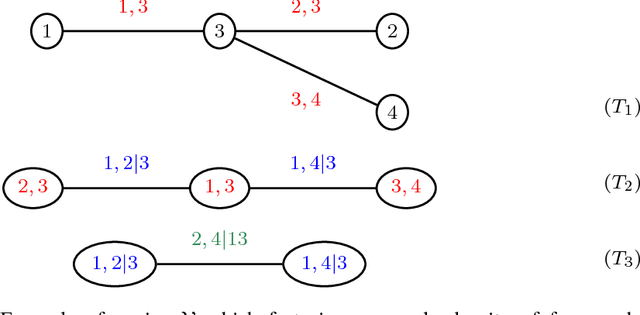
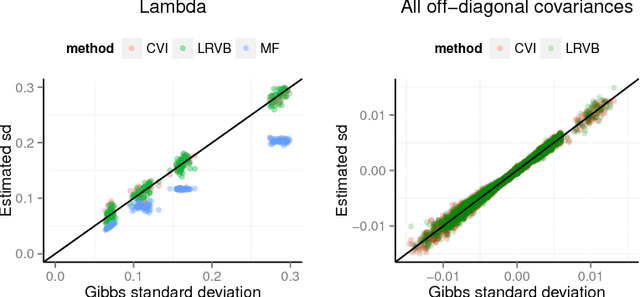
Abstract:We develop a general variational inference method that preserves dependency among the latent variables. Our method uses copulas to augment the families of distributions used in mean-field and structured approximations. Copulas model the dependency that is not captured by the original variational distribution, and thus the augmented variational family guarantees better approximations to the posterior. With stochastic optimization, inference on the augmented distribution is scalable. Furthermore, our strategy is generic: it can be applied to any inference procedure that currently uses the mean-field or structured approach. Copula variational inference has many advantages: it reduces bias; it is less sensitive to local optima; it is less sensitive to hyperparameters; and it helps characterize and interpret the dependency among the latent variables.
Stochastic gradient descent methods for estimation with large data sets
Sep 22, 2015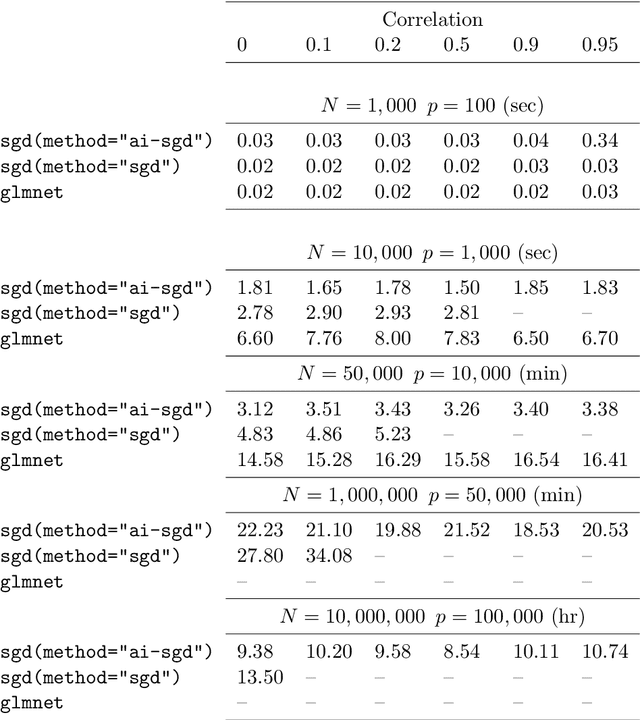
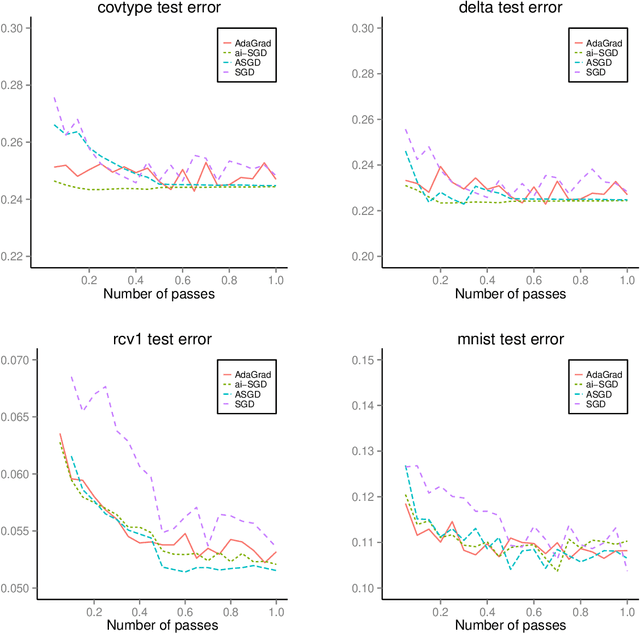
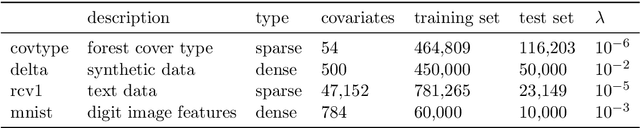
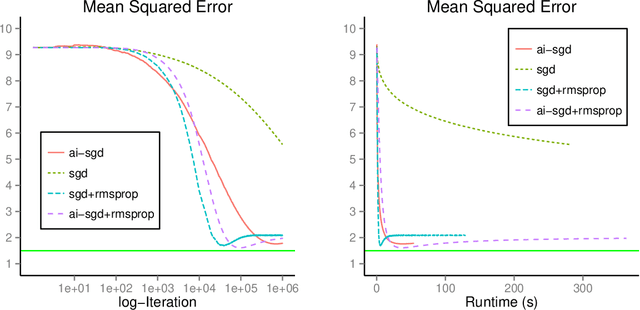
Abstract:We develop methods for parameter estimation in settings with large-scale data sets, where traditional methods are no longer tenable. Our methods rely on stochastic approximations, which are computationally efficient as they maintain one iterate as a parameter estimate, and successively update that iterate based on a single data point. When the update is based on a noisy gradient, the stochastic approximation is known as standard stochastic gradient descent, which has been fundamental in modern applications with large data sets. Additionally, our methods are numerically stable because they employ implicit updates of the iterates. Intuitively, an implicit update is a shrinked version of a standard one, where the shrinkage factor depends on the observed Fisher information at the corresponding data point. This shrinkage prevents numerical divergence of the iterates, which can be caused either by excess noise or outliers. Our sgd package in R offers the most extensive and robust implementation of stochastic gradient descent methods. We demonstrate that sgd dominates alternative software in runtime for several estimation problems with massive data sets. Our applications include the wide class of generalized linear models as well as M-estimation for robust regression.
Convex Techniques for Model Selection
Dec 02, 2014Abstract:We develop a robust convex algorithm to select the regularization parameter in model selection. In practice this would be automated in order to save practitioners time from having to tune it manually. In particular, we implement and test the convex method for $K$-fold cross validation on ridge regression, although the same concept extends to more complex models. We then compare its performance with standard methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge