Dong Wei
Three-Dimensional Segmentation of the Left Ventricle in Late Gadolinium Enhanced MR Images of Chronic Infarction Combining Long- and Short-Axis Information
May 21, 2022
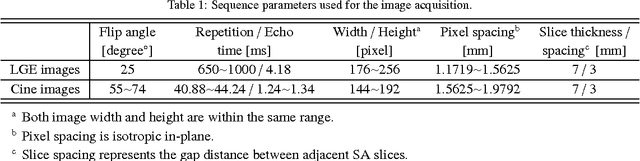
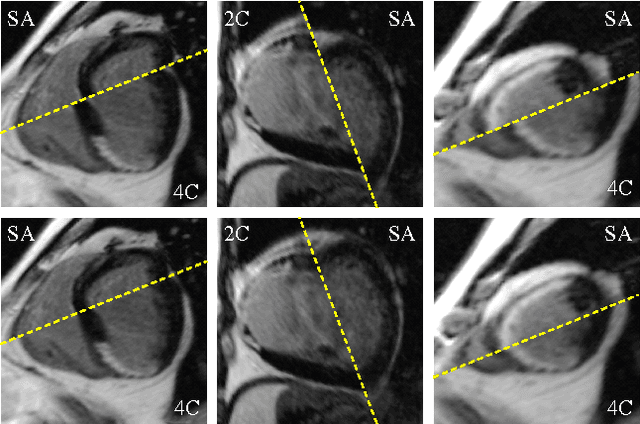
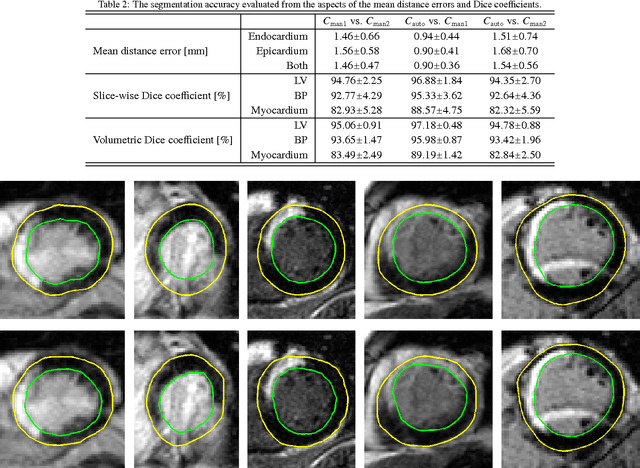
Abstract:Automatic segmentation of the left ventricle (LV) in late gadolinium enhanced (LGE) cardiac MR (CMR) images is difficult due to the intensity heterogeneity arising from accumulation of contrast agent in infarcted myocardium. In this paper, we present a comprehensive framework for automatic 3D segmentation of the LV in LGE CMR images. Given myocardial contours in cine images as a priori knowledge, the framework initially propagates the a priori segmentation from cine to LGE images via 2D translational registration. Two meshes representing respectively endocardial and epicardial surfaces are then constructed with the propagated contours. After construction, the two meshes are deformed towards the myocardial edge points detected in both short-axis and long-axis LGE images in a unified 3D coordinate system. Taking into account the intensity characteristics of the LV in LGE images, we propose a novel parametric model of the LV for consistent myocardial edge points detection regardless of pathological status of the myocardium (infarcted or healthy) and of the type of the LGE images (short-axis or long-axis). We have evaluated the proposed framework with 21 sets of real patient and 4 sets of simulated phantom data. Both distance- and region-based performance metrics confirm the observation that the framework can generate accurate and reliable results for myocardial segmentation of LGE images. We have also tested the robustness of the framework with respect to varied a priori segmentation in both practical and simulated settings. Experimental results show that the proposed framework can greatly compensate variations in the given a priori knowledge and consistently produce accurate segmentations.
Learning Shape Priors by Pairwise Comparison for Robust Semantic Segmentation
Apr 23, 2022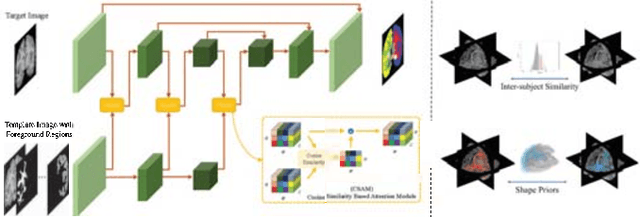
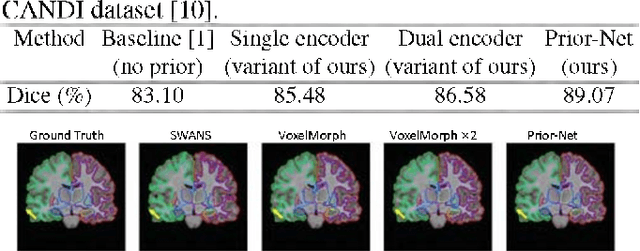
Abstract:Semantic segmentation is important in medical image analysis. Inspired by the strong ability of traditional image analysis techniques in capturing shape priors and inter-subject similarity, many deep learning (DL) models have been recently proposed to exploit such prior information and achieved robust performance. However, these two types of important prior information are usually studied separately in existing models. In this paper, we propose a novel DL model to model both type of priors within a single framework. Specifically, we introduce an extra encoder into the classic encoder-decoder structure to form a Siamese structure for the encoders, where one of them takes a target image as input (the image-encoder), and the other concatenates a template image and its foreground regions as input (the template-encoder). The template-encoder encodes the shape priors and appearance characteristics of each foreground class in the template image. A cosine similarity based attention module is proposed to fuse the information from both encoders, to utilize both types of prior information encoded by the template-encoder and model the inter-subject similarity for each foreground class. Extensive experiments on two public datasets demonstrate that our proposed method can produce superior performance to competing methods.
Domain Adaptation Meets Zero-Shot Learning: An Annotation-Efficient Approach to Multi-Modality Medical Image Segmentation
Mar 19, 2022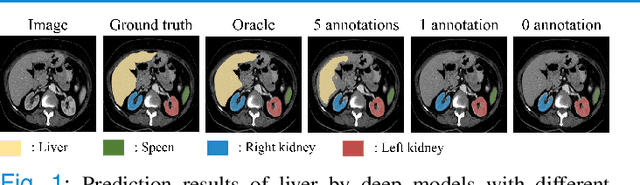
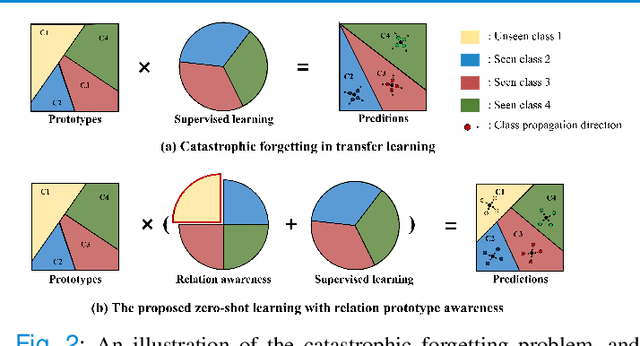
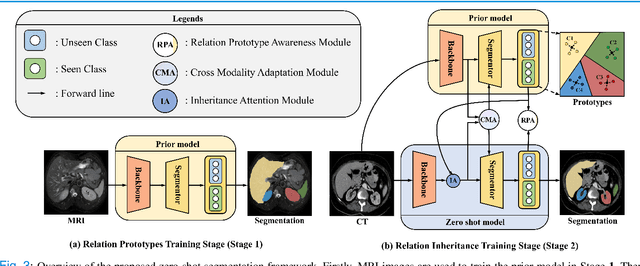
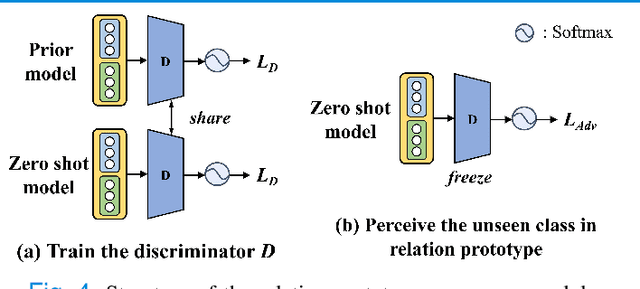
Abstract:Due to the lack of properly annotated medical data, exploring the generalization capability of the deep model is becoming a public concern. Zero-shot learning (ZSL) has emerged in recent years to equip the deep model with the ability to recognize unseen classes. However, existing studies mainly focus on natural images, which utilize linguistic models to extract auxiliary information for ZSL. It is impractical to apply the natural image ZSL solutions directly to medical images, since the medical terminology is very domain-specific, and it is not easy to acquire linguistic models for the medical terminology. In this work, we propose a new paradigm of ZSL specifically for medical images utilizing cross-modality information. We make three main contributions with the proposed paradigm. First, we extract the prior knowledge about the segmentation targets, called relation prototypes, from the prior model and then propose a cross-modality adaptation module to inherit the prototypes to the zero-shot model. Second, we propose a relation prototype awareness module to make the zero-shot model aware of information contained in the prototypes. Last but not least, we develop an inheritance attention module to recalibrate the relation prototypes to enhance the inheritance process. The proposed framework is evaluated on two public cross-modality datasets including a cardiac dataset and an abdominal dataset. Extensive experiments show that the proposed framework significantly outperforms the state of the arts.
Deep Convolutional Neural Networks for Molecular Subtyping of Gliomas Using Magnetic Resonance Imaging
Mar 10, 2022Abstract:Knowledge of molecular subtypes of gliomas can provide valuable information for tailored therapies. This study aimed to investigate the use of deep convolutional neural networks (DCNNs) for noninvasive glioma subtyping with radiological imaging data according to the new taxonomy announced by the World Health Organization in 2016. Methods: A DCNN model was developed for the prediction of the five glioma subtypes based on a hierarchical classification paradigm. This model used three parallel, weight-sharing, deep residual learning networks to process 2.5-dimensional input of trimodal MRI data, including T1-weighted, T1-weighted with contrast enhancement, and T2-weighted images. A data set comprising 1,016 real patients was collected for evaluation of the developed DCNN model. The predictive performance was evaluated via the area under the curve (AUC) from the receiver operating characteristic analysis. For comparison, the performance of a radiomics-based approach was also evaluated. Results: The AUCs of the DCNN model for the four classification tasks in the hierarchical classification paradigm were 0.89, 0.89, 0.85, and 0.66, respectively, as compared to 0.85, 0.75, 0.67, and 0.59 of the radiomics approach. Conclusion: The results showed that the developed DCNN model can predict glioma subtypes with promising performance, given sufficient, non-ill-balanced training data.
Conquering Data Variations in Resolution: A Slice-Aware Multi-Branch Decoder Network
Mar 07, 2022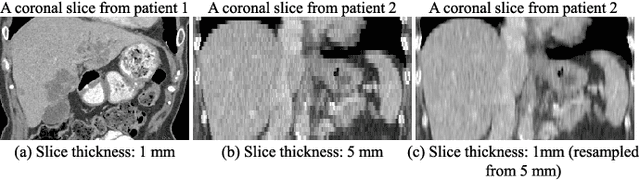
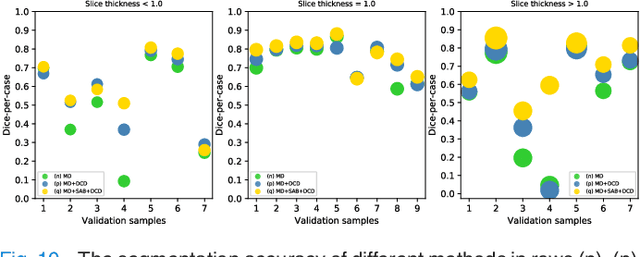
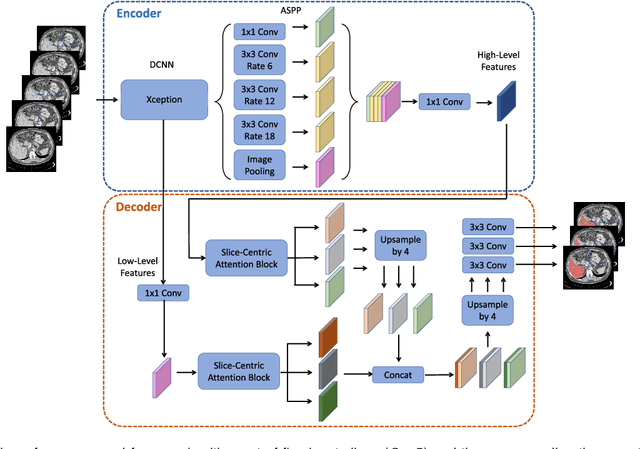
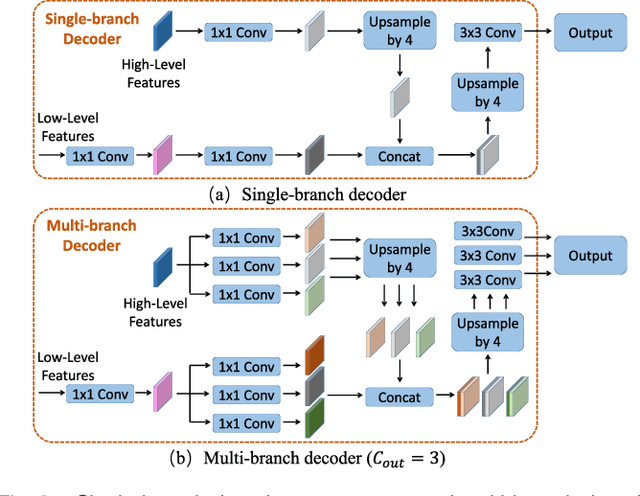
Abstract:Fully convolutional neural networks have made promising progress in joint liver and liver tumor segmentation. Instead of following the debates over 2D versus 3D networks (for example, pursuing the balance between large-scale 2D pretraining and 3D context), in this paper, we novelly identify the wide variation in the ratio between intra- and inter-slice resolutions as a crucial obstacle to the performance. To tackle the mismatch between the intra- and inter-slice information, we propose a slice-aware 2.5D network that emphasizes extracting discriminative features utilizing not only in-plane semantics but also out-of-plane coherence for each separate slice. Specifically, we present a slice-wise multi-input multi-output architecture to instantiate such a design paradigm, which contains a Multi-Branch Decoder (MD) with a Slice-centric Attention Block (SAB) for learning slice-specific features and a Densely Connected Dice (DCD) loss to regularize the inter-slice predictions to be coherent and continuous. Based on the aforementioned innovations, we achieve state-of-the-art results on the MICCAI 2017 Liver Tumor Segmentation (LiTS) dataset. Besides, we also test our model on the ISBI 2019 Segmentation of THoracic Organs at Risk (SegTHOR) dataset, and the result proves the robustness and generalizability of the proposed method in other segmentation tasks.
Simultaneous Alignment and Surface Regression Using Hybrid 2D-3D Networks for 3D Coherent Layer Segmentation of Retina OCT Images
Mar 04, 2022

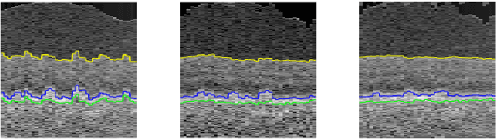
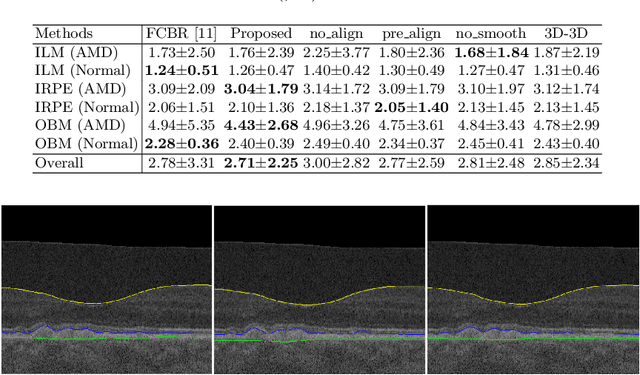
Abstract:Automated surface segmentation of retinal layer is important and challenging in analyzing optical coherence tomography (OCT). Recently, many deep learning based methods have been developed for this task and yield remarkable performance. However, due to large spatial gap and potential mismatch between the B-scans of OCT data, all of them are based on 2D segmentation of individual B-scans, which may loss the continuity information across the B-scans. In addition, 3D surface of the retina layers can provide more diagnostic information, which is crucial in quantitative image analysis. In this study, a novel framework based on hybrid 2D-3D convolutional neural networks (CNNs) is proposed to obtain continuous 3D retinal layer surfaces from OCT. The 2D features of individual B-scans are extracted by an encoder consisting of 2D convolutions. These 2D features are then used to produce the alignment displacement field and layer segmentation by two 3D decoders, which are coupled via a spatial transformer module. The entire framework is trained end-to-end. To the best of our knowledge, this is the first study that attempts 3D retinal layer segmentation in volumetric OCT images based on CNNs. Experiments on a publicly available dataset show that our framework achieves superior results to state-of-the-art 2D methods in terms of both layer segmentation accuracy and cross-B-scan 3D continuity, thus offering more clinical values than previous works.
Revisiting Experience Replay: Continual Learning by Adaptively Tuning Task-wise Relationship
Jan 06, 2022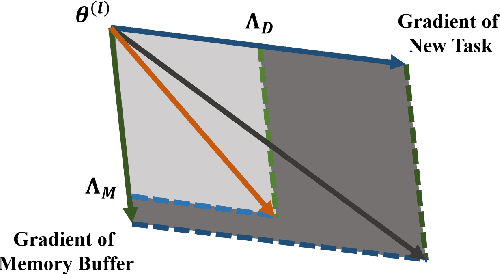
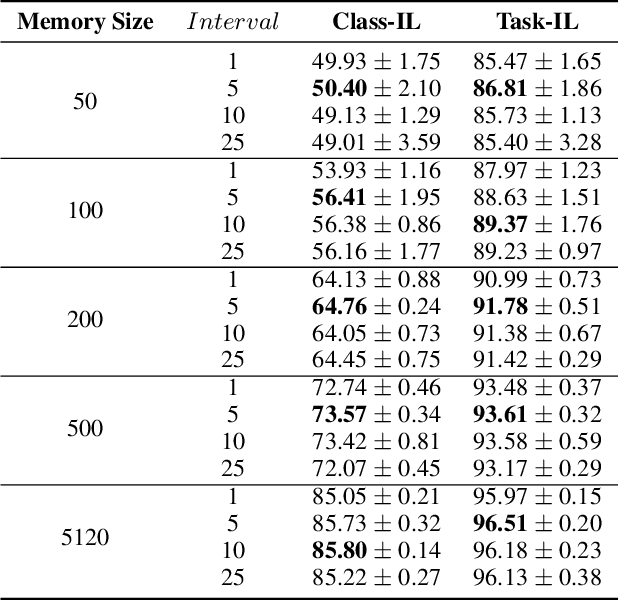
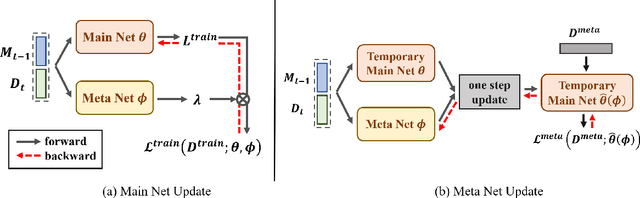
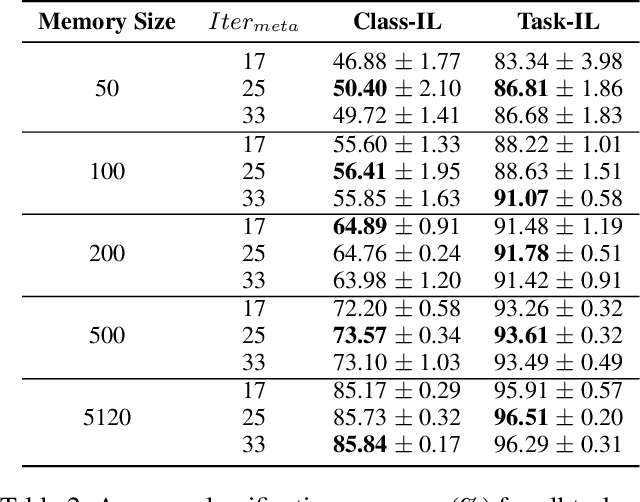
Abstract:Continual learning requires models to learn new tasks while maintaining previously learned knowledge. Various algorithms have been proposed to address this real challenge. Till now, rehearsal-based methods, such as experience replay, have achieved state-of-the-art performance. These approaches save a small part of the data of the past tasks as a memory buffer to prevent models from forgetting previously learned knowledge. However, most of them treat every new task equally, i.e., fixed the hyperparameters of the framework while learning different new tasks. Such a setting lacks the consideration of the relationship/similarity between past and new tasks. For example, the previous knowledge/features learned from dogs are more beneficial for the identification of cats (new task), compared to those learned from buses. In this regard, we propose a meta learning algorithm based on bi-level optimization to adaptively tune the relationship between the knowledge extracted from the past and new tasks. Therefore, the model can find an appropriate direction of gradient during continual learning and avoid the serious overfitting problem on memory buffer. Extensive experiments are conducted on three publicly available datasets (i.e., CIFAR-10, CIFAR-100, and Tiny ImageNet). The experimental results demonstrate that the proposed method can consistently improve the performance of all baselines.
A Unified Framework for Generalized Low-Shot Medical Image Segmentation with Scarce Data
Oct 18, 2021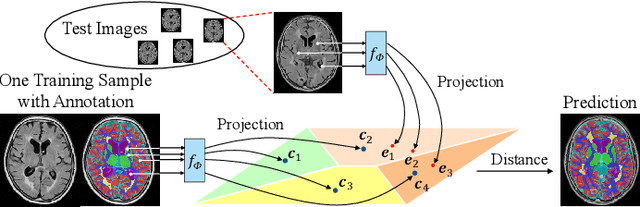
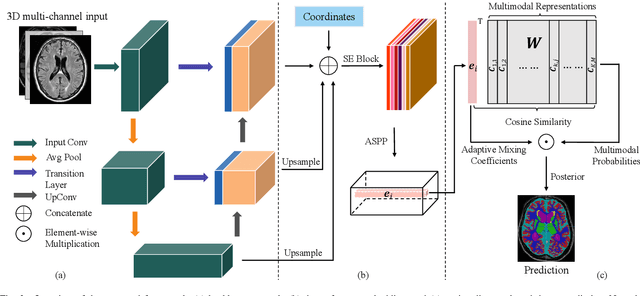
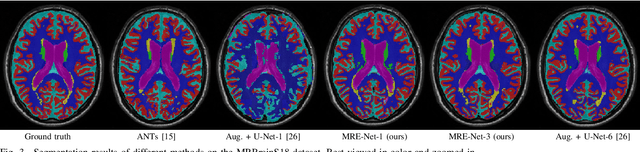
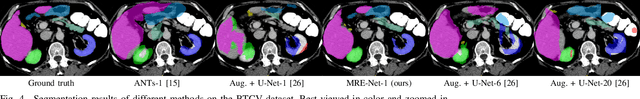
Abstract:Medical image segmentation has achieved remarkable advancements using deep neural networks (DNNs). However, DNNs often need big amounts of data and annotations for training, both of which can be difficult and costly to obtain. In this work, we propose a unified framework for generalized low-shot (one- and few-shot) medical image segmentation based on distance metric learning (DML). Unlike most existing methods which only deal with the lack of annotations while assuming abundance of data, our framework works with extreme scarcity of both, which is ideal for rare diseases. Via DML, the framework learns a multimodal mixture representation for each category, and performs dense predictions based on cosine distances between the pixels' deep embeddings and the category representations. The multimodal representations effectively utilize the inter-subject similarities and intraclass variations to overcome overfitting due to extremely limited data. In addition, we propose adaptive mixing coefficients for the multimodal mixture distributions to adaptively emphasize the modes better suited to the current input. The representations are implicitly embedded as weights of the fc layer, such that the cosine distances can be computed efficiently via forward propagation. In our experiments on brain MRI and abdominal CT datasets, the proposed framework achieves superior performances for low-shot segmentation towards standard DNN-based (3D U-Net) and classical registration-based (ANTs) methods, e.g., achieving mean Dice coefficients of 81%/69% for brain tissue/abdominal multiorgan segmentation using a single training sample, as compared to 52%/31% and 72%/35% by the U-Net and ANTs, respectively.
Unsupervised Representation Learning Meets Pseudo-Label Supervised Self-Distillation: A New Approach to Rare Disease Classification
Oct 09, 2021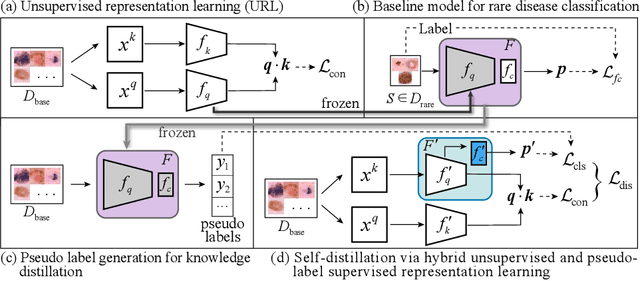
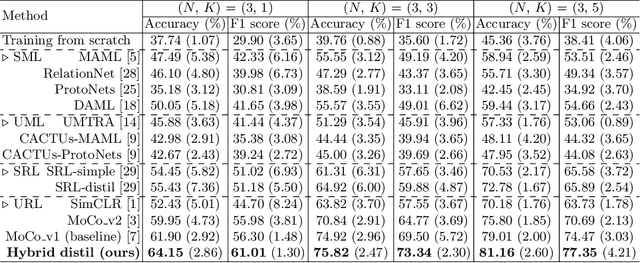
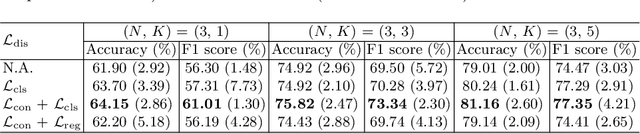
Abstract:Rare diseases are characterized by low prevalence and are often chronically debilitating or life-threatening. Imaging-based classification of rare diseases is challenging due to the severe shortage in training examples. Few-shot learning (FSL) methods tackle this challenge by extracting generalizable prior knowledge from a large base dataset of common diseases and normal controls, and transferring the knowledge to rare diseases. Yet, most existing methods require the base dataset to be labeled and do not make full use of the precious examples of the rare diseases. To this end, we propose in this work a novel hybrid approach to rare disease classification, featuring two key novelties targeted at the above drawbacks. First, we adopt the unsupervised representation learning (URL) based on self-supervising contrastive loss, whereby to eliminate the overhead in labeling the base dataset. Second, we integrate the URL with pseudo-label supervised classification for effective self-distillation of the knowledge about the rare diseases, composing a hybrid approach taking advantages of both unsupervised and (pseudo-) supervised learning on the base dataset. Experimental results on classification of rare skin lesions show that our hybrid approach substantially outperforms existing FSL methods (including those using fully supervised base dataset) for rare disease classification via effective integration of the URL and pseudo-label driven self-distillation, thus establishing a new state of the art.
Training Automatic View Planner for Cardiac MR Imaging via Self-Supervision by Spatial Relationship between Views
Sep 24, 2021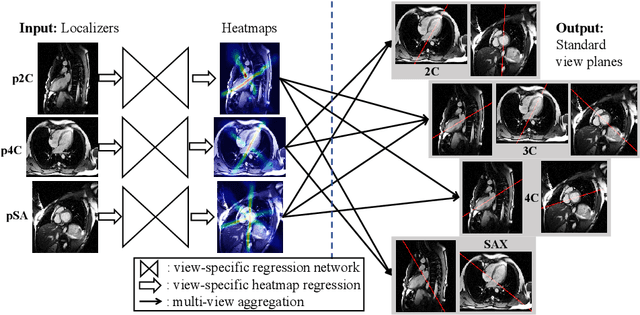
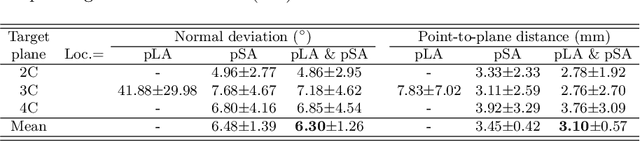
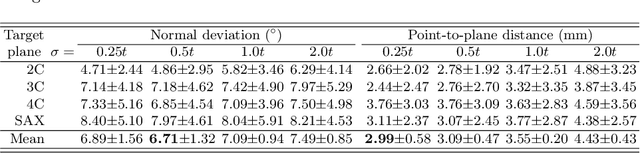
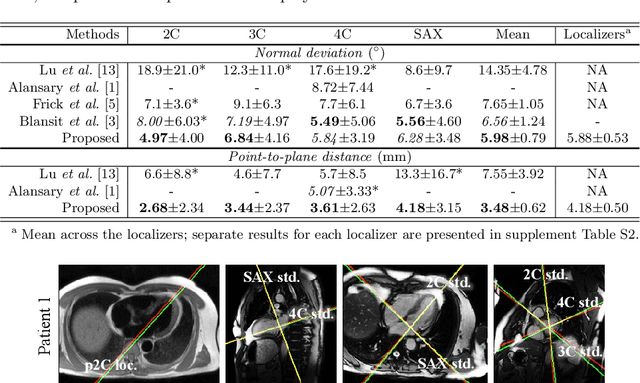
Abstract:View planning for the acquisition of cardiac magnetic resonance imaging (CMR) requires acquaintance with the cardiac anatomy and remains a challenging task in clinical practice. Existing approaches to its automation relied either on an additional volumetric image not typically acquired in clinic routine, or on laborious manual annotations of cardiac structural landmarks. This work presents a clinic-compatible and annotation-free system for automatic CMR view planning. The system mines the spatial relationship -- more specifically, locates and exploits the intersecting lines -- between the source and target views, and trains deep networks to regress heatmaps defined by these intersecting lines. As the spatial relationship is self-contained in properly stored data, e.g., in the DICOM format, the need for manual annotation is eliminated. Then, a multi-view planning strategy is proposed to aggregate information from the predicted heatmaps for all the source views of a target view, for a globally optimal prescription. The multi-view aggregation mimics the similar strategy practiced by skilled human prescribers. Experimental results on 181 clinical CMR exams show that our system achieves superior accuracy to existing approaches including conventional atlas-based and newer deep learning based ones, in prescribing four standard CMR views. The mean angle difference and point-to-plane distance evaluated against the ground truth planes are 5.98 degrees and 3.48 mm, respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge