Dino Sejdinovic
Hyperparameter Learning via Distributional Transfer
Oct 15, 2018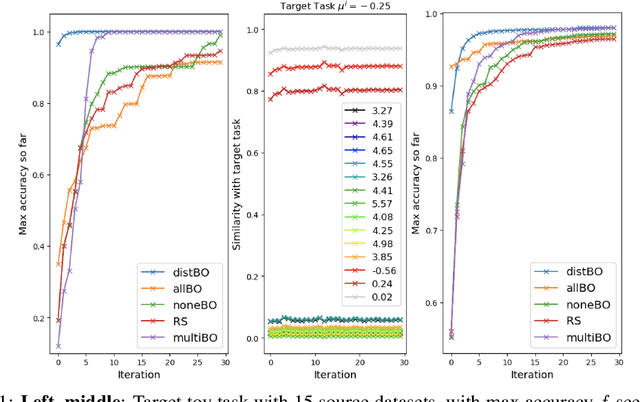
Abstract:Bayesian optimisation is a popular technique for hyperparameter learning but typically requires initial 'exploration' even in cases where potentially similar prior tasks have been solved. We propose to transfer information across tasks using kernel embeddings of distributions of training datasets used in those tasks. The resulting method has a faster convergence compared to existing baselines, in some cases requiring only a few evaluations of the target objective.
A Differentially Private Kernel Two-Sample Test
Aug 01, 2018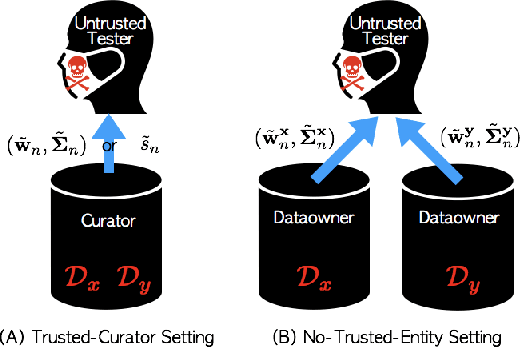
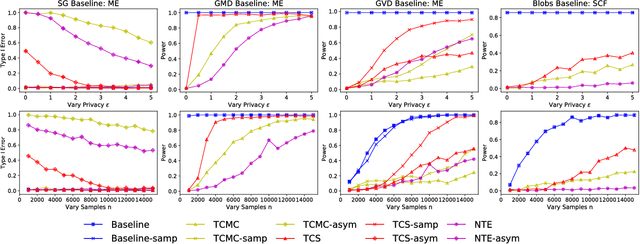
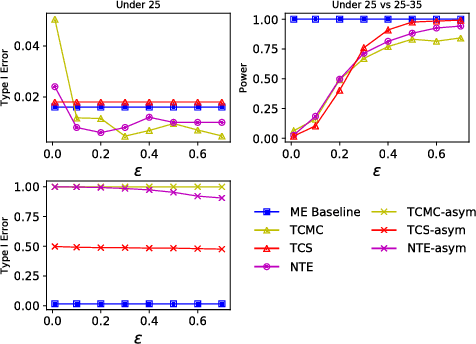
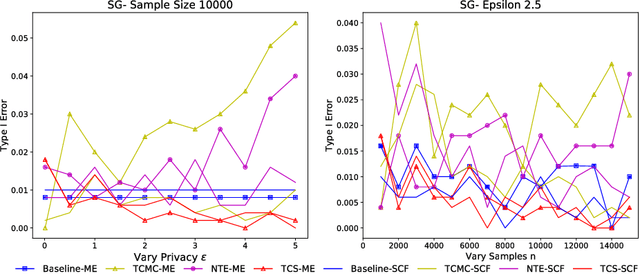
Abstract:Kernel two-sample testing is a useful statistical tool in determining whether data samples arise from different distributions without imposing any parametric assumptions on those distributions. However, raw data samples can expose sensitive information about individuals who participate in scientific studies, which makes the current tests vulnerable to privacy breaches. Hence, we design a new framework for kernel two-sample testing conforming to differential privacy constraints, in order to guarantee the privacy of subjects in the data. Unlike existing differentially private parametric tests that simply add noise to data, kernel-based testing imposes a challenge due to a complex dependence of test statistics on the raw data, as these statistics correspond to estimators of distances between representations of probability measures in Hilbert spaces. Our approach considers finite dimensional approximations to those representations. As a result, a simple chi-squared test is obtained, where a test statistic depends on a mean and covariance of empirical differences between the samples, which we perturb for a privacy guarantee. We investigate the utility of our framework in two realistic settings and conclude that our method requires only a relatively modest increase in sample size to achieve a similar level of power to the non-private tests in both settings.
Gaussian Processes and Kernel Methods: A Review on Connections and Equivalences
Jul 06, 2018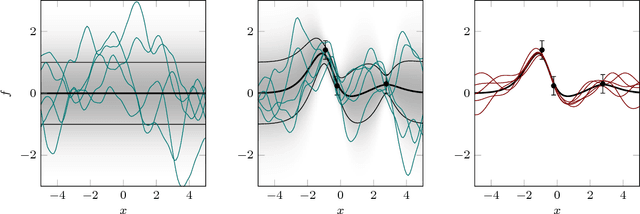
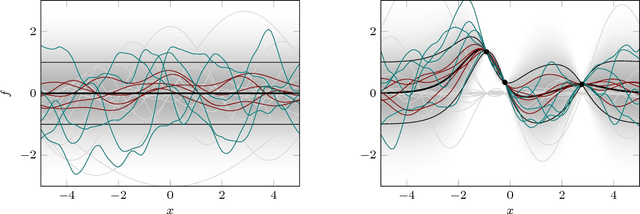
Abstract:This paper is an attempt to bridge the conceptual gaps between researchers working on the two widely used approaches based on positive definite kernels: Bayesian learning or inference using Gaussian processes on the one side, and frequentist kernel methods based on reproducing kernel Hilbert spaces on the other. It is widely known in machine learning that these two formalisms are closely related; for instance, the estimator of kernel ridge regression is identical to the posterior mean of Gaussian process regression. However, they have been studied and developed almost independently by two essentially separate communities, and this makes it difficult to seamlessly transfer results between them. Our aim is to overcome this potential difficulty. To this end, we review several old and new results and concepts from either side, and juxtapose algorithmic quantities from each framework to highlight close similarities. We also provide discussions on subtle philosophical and theoretical differences between the two approaches.
A Unified Analysis of Random Fourier Features
Jun 24, 2018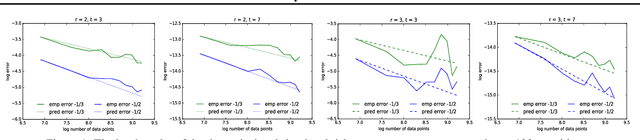
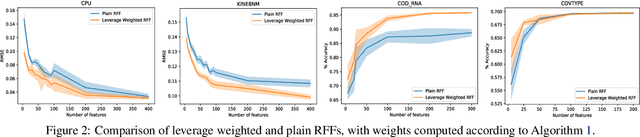
Abstract:We provide the first unified theoretical analysis of supervised learning with random Fourier features, covering different types of loss functions characteristic to kernel methods developed for this setting. More specifically, we investigate learning with squared error and Lipschitz continuous loss functions and give the sharpest expected risk convergence rates for problems in which random Fourier features are sampled either using the spectral measure corresponding to a shift-invariant kernel or the ridge leverage score function proposed in~\cite{avron2017random}. The trade-off between the number of features and the expected risk convergence rate is expressed in terms of the regularization parameter and the effective dimension of the problem. While the former can effectively capture the complexity of the target hypothesis, the latter is known for expressing the fine structure of the kernel with respect to the marginal distribution of a data generating process~\cite{caponnetto2007optimal}. In addition to our theoretical results, we propose an approximate leverage score sampler for large scale problems and show that it can be significantly more effective than the spectral measure sampler.
Hamiltonian Variational Auto-Encoder
May 29, 2018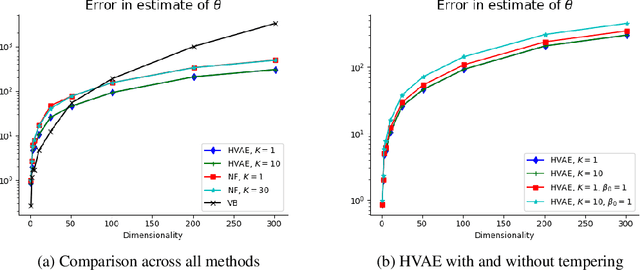
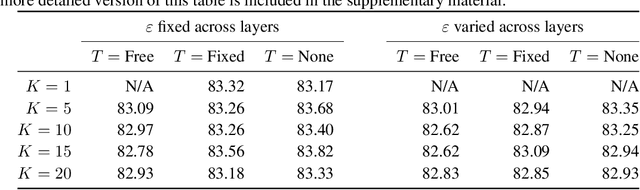
Abstract:Variational Auto-Encoders (VAEs) have become very popular techniques to perform inference and learning in latent variable models as they allow us to leverage the rich representational power of neural networks to obtain flexible approximations of the posterior of latent variables as well as tight evidence lower bounds (ELBOs). Combined with stochastic variational inference, this provides a methodology scaling to large datasets. However, for this methodology to be practically efficient, it is necessary to obtain low-variance unbiased estimators of the ELBO and its gradients with respect to the parameters of interest. While the use of Markov chain Monte Carlo (MCMC) techniques such as Hamiltonian Monte Carlo (HMC) has been previously suggested to achieve this [23, 26], the proposed methods require specifying reverse kernels which have a large impact on performance. Additionally, the resulting unbiased estimator of the ELBO for most MCMC kernels is typically not amenable to the reparameterization trick. We show here how to optimally select reverse kernels in this setting and, by building upon Hamiltonian Importance Sampling (HIS) [17], we obtain a scheme that provides low-variance unbiased estimators of the ELBO and its gradients using the reparameterization trick. This allows us to develop a Hamiltonian Variational Auto-Encoder (HVAE). This method can be reinterpreted as a target-informed normalizing flow [20] which, within our context, only requires a few evaluations of the gradient of the sampled likelihood and trivial Jacobian calculations at each iteration.
Variational Learning on Aggregate Outputs with Gaussian Processes
May 22, 2018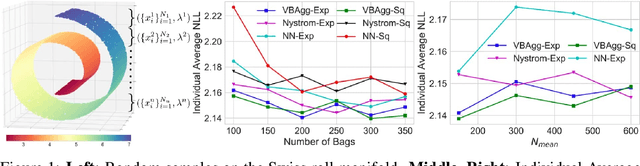
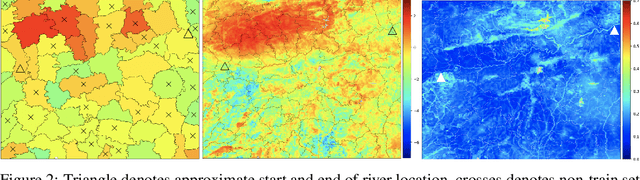
Abstract:While a typical supervised learning framework assumes that the inputs and the outputs are measured at the same levels of granularity, many applications, including global mapping of disease, only have access to outputs at a much coarser level than that of the inputs. Aggregation of outputs makes generalization to new inputs much more difficult. We consider an approach to this problem based on variational learning with a model of output aggregation and Gaussian processes, where aggregation leads to intractability of the standard evidence lower bounds. We propose new bounds and tractable approximations, leading to improved prediction accuracy and scalability to large datasets, while explicitly taking uncertainty into account. We develop a framework which extends to several types of likelihoods, including the Poisson model for aggregated count data. We apply our framework to a challenging and important problem, the fine-scale spatial modelling of malaria incidence, with over 1 million observations.
Causal Inference via Kernel Deviance Measures
Apr 12, 2018


Abstract:Discovering the causal structure among a set of variables is a fundamental problem in many areas of science. In this paper, we propose Kernel Conditional Deviance for Causal Inference (KCDC) a fully nonparametric causal discovery method based on purely observational data. From a novel interpretation of the notion of asymmetry between cause and effect, we derive a corresponding asymmetry measure using the framework of reproducing kernel Hilbert spaces. Based on this, we propose three decision rules for causal discovery. We demonstrate the wide applicability of our method across a range of diverse synthetic datasets. Furthermore, we test our method on real-world time series data and the real-world benchmark dataset Tubingen Cause-Effect Pairs where we outperform existing state-of-the-art methods.
Bayesian Approaches to Distribution Regression
Feb 22, 2018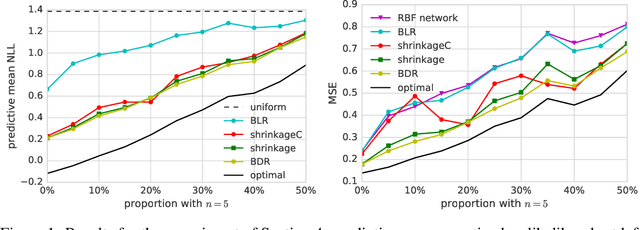
Abstract:Distribution regression has recently attracted much interest as a generic solution to the problem of supervised learning where labels are available at the group level, rather than at the individual level. Current approaches, however, do not propagate the uncertainty in observations due to sampling variability in the groups. This effectively assumes that small and large groups are estimated equally well, and should have equal weight in the final regression. We account for this uncertainty with a Bayesian distribution regression formalism, improving the robustness and performance of the model when group sizes vary. We frame our models in a neural network style, allowing for simple MAP inference using backpropagation to learn the parameters, as well as MCMC-based inference which can fully propagate uncertainty. We demonstrate our approach on illustrative toy datasets, as well as on a challenging problem of predicting age from images.
Spatial Mapping with Gaussian Processes and Nonstationary Fourier Features
Nov 15, 2017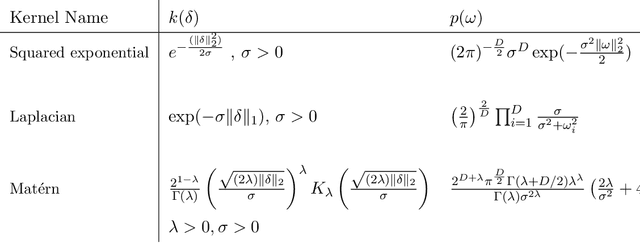
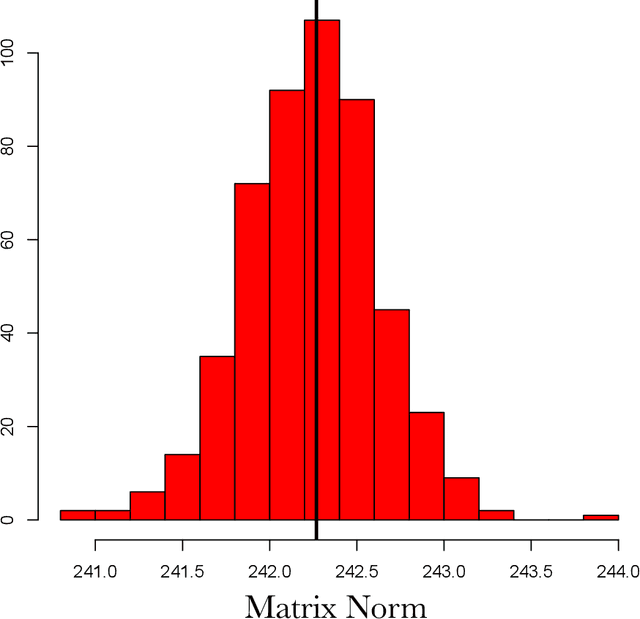

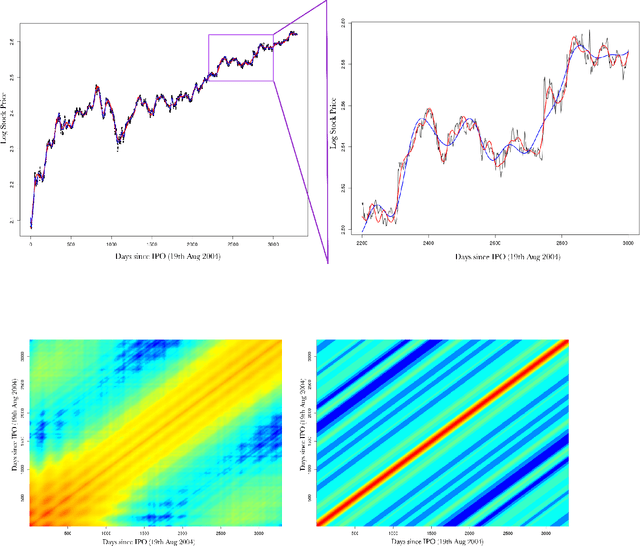
Abstract:The use of covariance kernels is ubiquitous in the field of spatial statistics. Kernels allow data to be mapped into high-dimensional feature spaces and can thus extend simple linear additive methods to nonlinear methods with higher order interactions. However, until recently, there has been a strong reliance on a limited class of stationary kernels such as the Matern or squared exponential, limiting the expressiveness of these modelling approaches. Recent machine learning research has focused on spectral representations to model arbitrary stationary kernels and introduced more general representations that include classes of nonstationary kernels. In this paper, we exploit the connections between Fourier feature representations, Gaussian processes and neural networks to generalise previous approaches and develop a simple and efficient framework to learn arbitrarily complex nonstationary kernel functions directly from the data, while taking care to avoid overfitting using state-of-the-art methods from deep learning. We highlight the very broad array of kernel classes that could be created within this framework. We apply this to a time series dataset and a remote sensing problem involving land surface temperature in Eastern Africa. We show that without increasing the computational or storage complexity, nonstationary kernels can be used to improve generalisation performance and provide more interpretable results.
Testing and Learning on Distributions with Symmetric Noise Invariance
Nov 05, 2017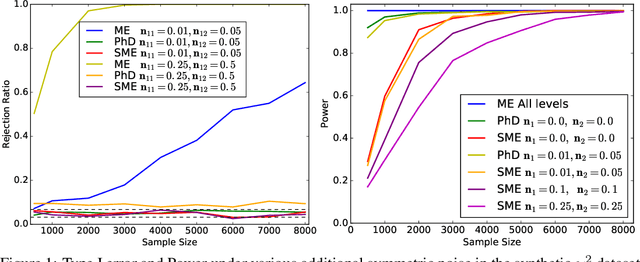
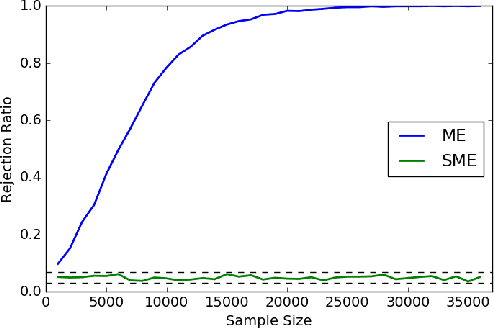
Abstract:Kernel embeddings of distributions and the Maximum Mean Discrepancy (MMD), the resulting distance between distributions, are useful tools for fully nonparametric two-sample testing and learning on distributions. However, it is rarely that all possible differences between samples are of interest -- discovered differences can be due to different types of measurement noise, data collection artefacts or other irrelevant sources of variability. We propose distances between distributions which encode invariance to additive symmetric noise, aimed at testing whether the assumed true underlying processes differ. Moreover, we construct invariant features of distributions, leading to learning algorithms robust to the impairment of the input distributions with symmetric additive noise.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge